- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Relative Stiffness Measurements of Tumour Tissues by Shear Rheology
Published: Vol 7, Iss 9, May 5, 2017 DOI: 10.21769/BioProtoc.2265 Views: 11442
Reviewed by: Ralph BottcherVikash VermaCristina Rohena

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Relative Stiffness Measurements of Cell-embedded Hydrogels by Shear Rheology in vitro
Thomas R. Cox and Chris D. Madsen
Jan 5, 2017 12460 Views

Application of Mechanical Forces on Drosophila Embryos by Manipulation of Microinjected Magnetic Particles
Arturo D’Angelo and Jérôme Solon
May 5, 2020 4409 Views
Abstract
The microenvironment of solid tumours is a critical contributor to the progression of tumours and offers a promising target for therapeutic intervention (Cox and Erler, 2011; Barker et al., 2012; Cox et al., 2016; Cox and Erler, 2016). The properties of the tumour microenvironment vary significantly from that of the original tissue in both biochemistry and biomechanics. At present, the complex interplay between the biomechanical properties of the microenvironment and tumour cell phenotype is under intense investigation. The ability to measure the biomechanical properties of tumour samples from cancer models will increase our understanding of their importance in solid tumour biology. Here we report a simple method to measure the viscoelastic properties of tumour specimens using a controlled strain rotational rheometer.
Keywords: Shear rheologyBackground
The growth of solid tumours is accompanied by pathological remodelling of the native tissue (Cox and Erler, 2011; Bonnans et al., 2014). During progression, the local tissue environment experiences physical as well as biological changes, resulting in increased tissue stiffness (elastic modulus) (Humphrey et al., 2014). Alterations in the extracellular matrix lead to the generation of new tissue properties, which activate mechano-signalling pathways within tumour cells (DuFort et al., 2011). This outside-in signalling leads to altered behaviour, cell morphology, differentiation, proliferation, migration and stemness. In preclinical animal models of cancer, these changes have been shown to drive malignant progression and metastatic spread (Erler et al., 2006; Levental et al., 2009; Bonnans et al., 2014). Thus, as a result, the targeting of matrix remodelling and in particular stiffening has received substantial attention in recent years, and several clinical trials have been initiated (Barker et al., 2012; Baker et al., 2013; Cox et al., 2013; Miller et al., 2015; Madsen et al., 2015; Cox and Erler, 2016; Kai et al., 2016).
The mechanical properties of the tumour microenvironment can readily be examined using approaches such as atomic force microscopy (AFM) and nanoindentation (Akhtar et al., 2009). These approaches provide nanometre resolution and concurrent measurement of the applied force with picoNewton resolution (Kasas and Dietler, 2008). However, AFM is not applicable to understand the elastic properties of larger 3D samples. The mechanical properties of bulk 3D tumour samples can be more accurately examined using shear rheology (Picout and Ross-Murphy, 2003). Rheology is the study of how a material deforms when forces are applied to them. Thus applying shear stress to a 3D matrix can determine the elastic modulus (stiffness) as well as viscous properties of a bulk 3D tumour tissue. In this protocol, we describe a method to measure changes on tumour stiffness by shear rheology.
Materials and Reagents
- 1,000 μl sterile pipet tip
- 1.5 ml sterile microcentrifuge tubes
- Cell strainer, 70 µm (VWR, catalog number: 734-0003 )
- Hypodermic needles, 27 G (Fine-JectR) (VWR, catalog number: 613-2012 )
- 8 mm disposable biopsy punch (KAI, catalog number: BP-80F )
- Precision syringes, 1 ml (VWR, catalog number: 613-3908 )
- Standard scalpel #11 (Fine Science Tools, catalog number: 10011-00 )
- The 4T1 wild-type cell line was obtained from F. Miller at the University of Michigan
- The SW480, early-stage colon adenocarcinoma (Duke stage B) cell line was obtained from the American Type Culture Collection (ATCC) (LGC Standards) (ATCC, catalog number: CCL-228 )
- The SW480 + LOX cell line was derived from the parent line and has been previously described (Baker et al., 2011; Baker et al., 2013)
- For the human colorectal cancer model, 8-week-old female immunodeficient MF1 nude mice were used (Envigo [formerly known as Harlan Laboratories Inc.])
- For the murine mammary carcinoma model, 8-week-old female BALB/c mice were used (Taconic Biosciences)
- DMEM (Thermo Fisher Scientific, GibcoTM, catalog number: 31966047 )
- Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10270106 )
- Penicillin-streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Sterile PBS, pH 7.2 (Thermo Fisher Scientific, GibcoTM, catalog number: 20012068 )
- Trypsin-EDTA (0.25%), phenol red (Thermo Fisher Scientific, GibcoTM, catalog number: 25200056 )
- Ethanol
- Growth medium (see Recipes)
Equipment
- Benchtop Centrifuge capable of holding 15 ml tubes
- Pipette, P1000 (Gilson)
- Cell incubator at 37 °C, 5% CO2 (HERAcell)
- Standard glass haematocytometer
- Calipers
- Stainless Steel Spatula, One End Flat, One End Bent, 6 in. in length (United Scientific Supplies, model: SSFB06 )
- Standard pattern surgical scissors blunt/blunt (Fine Science Tools, catalog number: 14000-18 )
- 8 mm sand-blasted smart-swap upper geometry, Figure 1A (arrowhead) (TA Instruments)
- 8 mm sand-blasted stepped lower geometry, Figure 1A (arrow) (TA Instruments)
- Discovery Series Hybrid rheometer (TA Instruments, model: dhr-2 )
Procedure
This protocol describes the biomechanical interrogation of tissue samples, which can be obtained from a variety of sources. Here, we utilise the 4T1/BALB/c syngeneic orthotopic model of murine mammary carcinoma (Miller and Heppner, 1979; Cox et al., 2013; Cox et al., 2015), and a subcutaneous SW480/Nude model of human colorectal cancer which has been engineered to overexpress the matrix cross-linking enzyme lysyl oxidase (LOX) (Baker et al., 2011; Baker et al., 2013). High LOX expression in primary tumours has been shown to cross-link extracellular matrix components, in particular collagens, leading to increases in tensile strength of the tissues (Levental et al., 2009; Baker et al., 2011; Baker et al., 2013). Tumour tissue can be collected from any source, including Genetically Engineered Mouse Models (GEMMS), spontaneous models of cancer, as well as orthotopic and subcutaneous models of cancer (see Note 1). Whilst we demonstrate the approach with both a human colorectal cancer model and murine breast cancer model, any solid tumour tissue, including patient material, could be used providing tissue samples meet the following criteria:
A.The tissue must be easily accessible for intact surgical resection.
B.The tissue can be processed fresh immediately (see Notes 2-4).
C.A minimum biopsy size of 8 mm in diameter and ≥ 1 mm thick can be obtained.
D.Samples can be measured immediately (see Notes 2-4).
- Measuring relative stiffness of tumour tissues
Rheological characterization was performed on all tumour samples using a TA Instruments DHR-2 controlled strain rotational rheometer fitted with an 8 mm sand-blasted parallel plate upper geometry, and an 8 mm sand-blasted lower stepped geometry (Figure 1A). Table 1 below outlines the testing parameters that we have determined to be optimal for measuring tumour tissue using a Discovery Series Hybrid rheometer (TA Instruments, model: DHR-2) under the conditions described (see Notes 5-7).
Table 1. Rheometer settingsParameter Value Temperature (°C) 21 Temperature soak time (sec) 0 Oscillation frequency (rad/sec) 1.0 Oscillation strain (%) 0.2-2.0 Data points per decade 15 Controlled strain type Continuous Oscillation [direct strain] Axial force (N) 0.03-0.05 Conditioning time (sec) 2.0 Sampling time (sec) 3.0 - Tumour implantation
- 4T1 and SW480 cell lines were maintained in DMEM/10% FBS/Pen-Strep at 37 °C and 5% CO2.
- Aspirate the growth medium from the cells and wash the cells once briefly with PBS.
- Aspirate the PBS, and add trypsin-EDTA (0.25%) enough to just cover the cells.
- Once the cells have detached, centrifuge at 300 x g for 5 min and resuspend in normal growth medium and count the cells.
- Wash the cells in PBS, then remove cell clusters by passing through a cell strainer (70 µm).
- Clean and sterilize the inoculation area of the mice with ethanol.
- Mix the cells and draw the cells into a syringe without a needle. Place a 27 G needle on the syringe and make sure to remove any air and bubbles.
- Inject 100 µl tumour cells into the anatomical location of interest (see Note 8).
- Following implantation, regular monitoring of the animals as per institutional guidelines is required, and tumour size and body weight is measured three times a week (see Note 9).
- 4T1 and SW480 cell lines were maintained in DMEM/10% FBS/Pen-Strep at 37 °C and 5% CO2.
- Experimental setup
- Startup and calibrate the rheometer according to manufacturer’s guidelines.
- Attach the stepped lower geometry to the heated peltier plate (Figure 1A) (arrow).
- Attach the 8 mm diameter upper geometry (Figure 1A) (arrowhead).
- Set the peltier temperature to the required temperature (Table 1).
- Surgically excise the tumour tissue to be measured (see Notes 2, 3, 4 and 10). Figures 1B-1D show a primary orthotopic fatpad-implanted 4T1 mammary carcinoma tumour at 3 weeks post implantation being excised. When measured externally using calipers the tumour measures approximately 520 mm3, which is sufficient to allow an 8 mm diameter tissue sample to be collected (see conditions above under Procedures).
- A ≥ 1 mm thick section is cut from the region of interest for measurement (Figures 1E and 1F). The remaining tissue not used for rheology can be used for additional analysis including RNA/DNA/protein extraction, or fixed/frozen and processed for immunohistochemistry/immunofluorescence (see Note 11).
- Using an 8 mm disposable biopsy punch, a circular biopsy is cut and extraneous material is removed (Figure 1G).
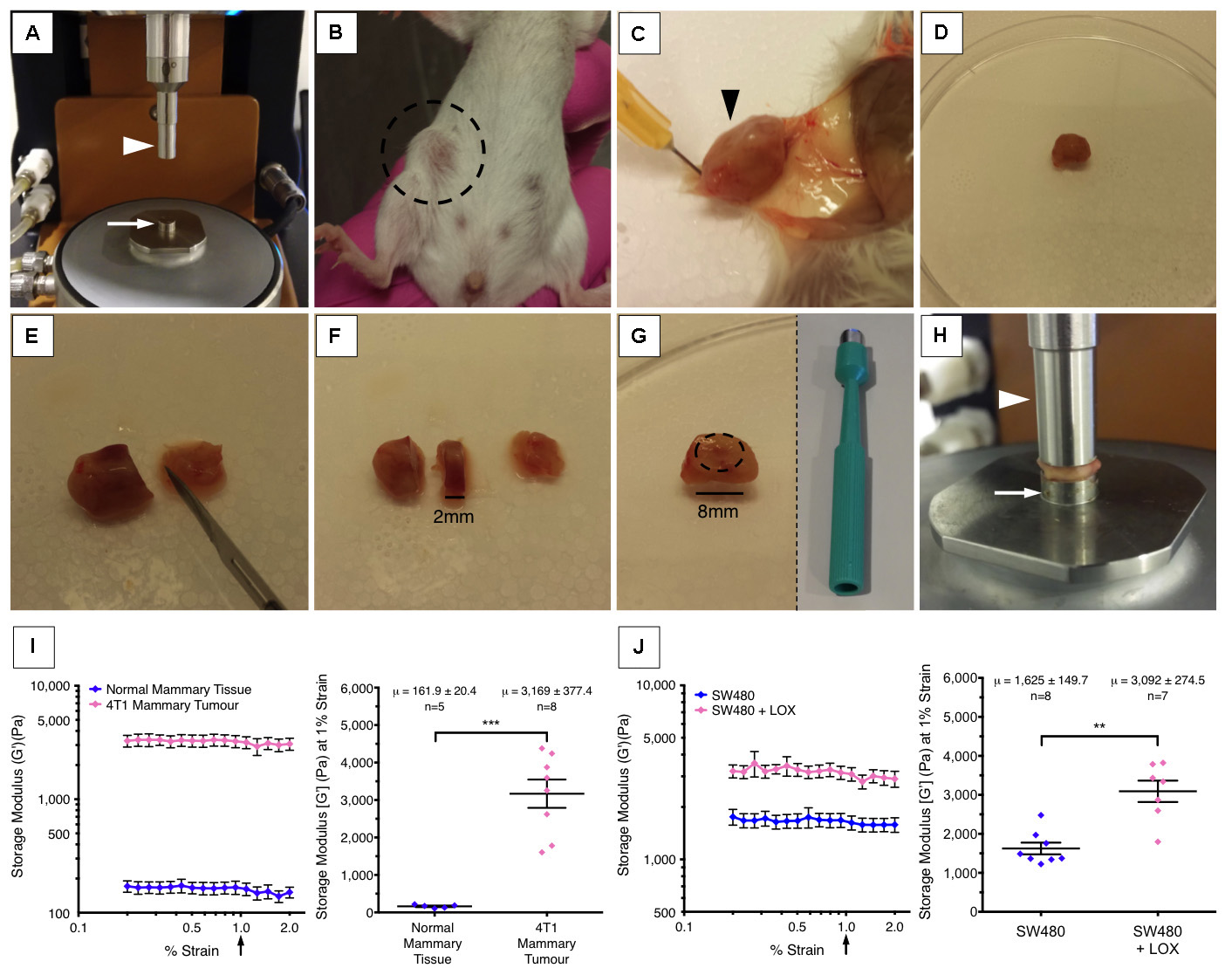
Figure 1. Rheology set-up. A. 8 mm sand-blasted lower stepped geometry (arrow), peltier plate and 8 mm sand-blasted upper geometry (arrowhead); B. Primary orthotopic 4T1 tumour (circled) at 3-week post-implantation into the mammary fatpad of a BALB/c mouse; C and D. Post-mortem surgical excision of the whole tumour (arrowhead) from the fatpad (E and F). A ≥ 1 mm section is cut using a scalpel from the region of interest in the tumour (see Notes 2-4). G. Using an 8 mm disposable biopsy punch, a circular biopsy is cut and extraneous material is removed. H. The tissue sample is then placed on the lower geometry (arrow) and the upper geometry (arrowhead) lowered so that it comes into contact with the sample. The gap between the upper geometry (arrowhead) and lower geometry (arrow) is slowly reduced until a stable axial force (see Table 1) is achieved and the measurement commences. I. Representative example of measurements taken from 4T1 mammary carcinoma tumours at 3 weeks post implantation compared to normal mammary fat pad tissue bearing no tumour. The storage modulus is measured over one decade of strain from 0.2 to 2% (left). The right-hand graph represents the storage modulus at 1% strain showing that tumour tissue is significantly stiffer than healthy control tissue. µ represents the mean storage modulus (G’) ± standard deviation. J. Representative example of how over-expression of lysyl oxidase (LOX), a potent extracellular matrix cross-linker, leads to increases in the stiffness of tumour tissue in a human colorectal cancer model. The storage modulus (G’) of tumour tissue is measured 4 weeks after implantation subcutaneously (left). The right-hand graph represents the storage modulus at 1% strain, and illustrates that over-expression of LOX in tumour cells leads to stiffening of their environment. µ represents the mean storage modulus (G’) ± standard deviation. - Zero the axial force on the rheometer.
- Carefully transfer the tumour tissue to the lower geometry (arrow) using a flat-ended spatula.
- Move the 8 mm upper geometry (arrowhead) down until it just contacts the top surface of the tissue (Figure 1H).
- Decrease the gap by 50 µm increments to increase the axial force applied to the tissue and continue until a stable axial force is reached as detailed in Table 1 (see Note 7).
- Set an oscillation strain sweep as per Table 1.
- Set an oscillation frequency as per Table 1.
- Begin measurement.
- Startup and calibrate the rheometer according to manufacturer’s guidelines.
- Analysis of the relative stiffness
- Ensure a linear viscoelastic response [storage modulus (G’)] within the strain range evaluated (Figures 1I and 1J) (see Notes 6 and 12).
- Extract the storage modulus (G’) at a matched strain between samples (example here is 1% strain) when comparing multiple tissue measurements (Figures 1I and 1J).
- If required, the elastic moduli (E) can be determined from the storage modulus (G’) using:
E = 2 x G’ (1 + υ)
where, υ = Poisson’s ratio of 0.5 for most tissues.
- Ensure a linear viscoelastic response [storage modulus (G’)] within the strain range evaluated (Figures 1I and 1J) (see Notes 6 and 12).
Data analysis
To ensure reliable data, be sure to perform at least five biological repeats within each experimental group with the appropriate controls. Extract the storage modulus (G’) at 1% strain for each repeat when comparing multiple tissue measurements (Figures 1I and 1J, right panels). Ensure a linear viscoelastic (storage modulus [G’]) response within the strain range evaluated (Figures 1I and 1J, left panels) (see Notes 5 and 6). If this is not the case, disregard the measurement. If this is a recurrent issue, frequencies and strains will need to be optimised for the specific tissue under evaluation (see Note 5).
Notes
- All animal experiments were carried out under authorisation and guidance from the Danish Inspectorate for Animal Experimentation.
- Avoid storing tissues and especially prepared biopsies for measurement in buffers such as PBS for long periods of time (in excess of 1 h) prior to measurement as this can result in the tissue swelling and breaking up affecting mechanical properties.
- Do not allow tissues or prepared biopsies to dry out as this causes shrinkage and will also affect mechanical properties.
- Store whole tissues on ice prior to preparation. Avoid storing for more than 1 h prior to measurements.
- Optimisation of measurement parameters may be required for extremely soft or extremely stiff samples. It may be necessary to determine the optimal frequency and strain parameters to ensure measurements are collected within the linear viscoelastic response of the tissue.
- The point at which a Storage modulus (G’) deviates by more than 10% from a constant (plateau) value indicates departure from linear viscoelastic behaviour.
- When applying an axial force to the samples prior to starting the measurements, ensure this is consistent across measurements. A value of 0.03-0.05 N for tumour tissues described above is sufficient.
- The number of implanted tumour cells depends on the set-up of the experiment. We implant 0.5 x 106 4T1 cells resuspended in 100 µl PBS per mouse in the 4th mammary fat pad of 8-week old BALB/c mice. We implant 1 x 106 SW480 cells resuspended in 100 µl PBS per mouse subcutaneously into the flank of 8-week old immunodeficient MF1 nude mice. In both models general anesthesia is not required in mice for implantation as approved by ethics (see Note 1). It is important to prepare a sufficient amount of cells resuspended in PBS to include the dead volume of the needle (typically 100 µl). We recommend to prepare approximately twice as much as needed. It is also important that the cells are actively growing (exponential growth) before harvesting the cells for implantation.
- Make sure to monitor the health state (body weight and respiration) of animals following tumour implantation on a regular basis. The external/superficial tumour size can be monitored using Calipers and should not exceed institutional guidelines.
- The time taken for the tumours to reach a suitable size will depend on the cell line in question and should be optimised by the researcher prior to the commencement of rheology experiments. Typically the 4T1-Balb/c syngeneic mouse model of mammary carcinoma will reach 8-10 mm in diameter within 21-24 days post implantation. The SW480/Nude subcutaneous model of colorectal cancer takes approximately 25-40 days from implantation. When collecting samples, be sure to collect via surgical resection under sterile conditions. Tissues should be minimum 8 mm in diameter and minimum 1 mm in thickness (see Figure 1). Typically a tissue thickness of 2-4 mm is ideal.
- Following rheological characterisation, tissues can be fixed with formalin or PFA and used in routine histopathological staining (such as haematoxylin and eosin staining for cells or Picrosirius Red staining for collagens). The use of these samples for immunohistochemistry, or protein/DNA/RNA extraction is cautioned against, since rheological characterisation is carried out fresh and may lead to changes in gene and protein expression during measurement.
- Tissues are typically only minimally frequency dependent within the range of 0.1 to 10 rad/sec (data not shown) and an oscillation frequency of 1 rad/sec is preferred when using the DHR2. At this frequency tumour samples show a linear viscoelastic response within the strain range evaluated (see Figures 1I and 1J).
Recipes
- Growth medium
DMEM
Fetal bovine serum (10%)
Penicillin-streptomycin (100 U/ml)
Acknowledgments
This protocol has been adapted from previous published papers (Baker et al., 2013; Cox et al., 2013; Madsen et al., 2015). TRC is supported by an NHMRC New Investigator grant, Australia. CDM is supported by the Ragnar Söderberg Foundation, BioCARE, Cancerfonden, and Åke Wiberg foundation, all Sweden. We thank Lena Wullkopf, Biotech Research Innovation Centre, University of Copenhagen for assistance and also thank Professor Janine Erler at the Biotech Research & Innovation Centre, University of Copenhagen for providing access to the rheometer.
References
- Akhtar, R., Schwarzer, N., Sherratt, M. J., Watson, R. E., Graham, H. K., Trafford, A. W., Mummery, P. M. and Derby, B. (2009). Nanoindentation of histological specimens: Mapping the elastic properties of soft tissues. J Mater Res 24(3): 638-646.
- Baker, A. M., Bird, D., Lang, G., Cox, T. R. and Erler, J. T. (2013). Lysyl oxidase enzymatic function increases stiffness to drive colorectal cancer progression through FAK. Oncogene 32(14): 1863-1868.
- Baker, A. M., Cox, T. R., Bird, D., Lang, G., Murray, G. I., Sun, X. F., Southall, S. M., Wilson, J. R. and Erler, J. T. (2011). The role of lysyl oxidase in SRC-dependent proliferation and metastasis of colorectal cancer. J Natl Cancer Inst 103(5): 407-424.
- Barker, H. E., Cox, T. R. and Erler, J. T. (2012). The rationale for targeting the LOX family in cancer. Nat Rev Cancer 12(8): 540-552.
- Bonnans, C., Chou, J. and Werb, Z. (2014). Remodelling the extracellular matrix in development and disease. Nat Rev Mol Cell Biol 15(12): 786-801.
- Cox, T. R., Bird, D., Baker, A. M., Barker, H. E., Ho, M. W., Lang, G. and Erler, J. T. (2013). LOX-mediated collagen crosslinking is responsible for fibrosis-enhanced metastasis. Cancer Res 73(6): 1721-1732.
- Cox, T. R. and Erler, J. T. (2011). Remodeling and homeostasis of the extracellular matrix: implications for fibrotic diseases and cancer. Dis Model Mech 4(2): 165-178.
- Cox, T. R. and Erler, J. T. (2016). Fibrosis and cancer: Partners in crime or opposing forces? Trends Cancer 2: 279-282.
- Cox, T. R., Gartland, A. and Erler, J. T. (2016). Lysyl oxidase, a targetable secreted molecule involved in cancer metastasis. Cancer Res 76(2): 188-192.
- Cox, T. R., Rumney, R. M., Schoof, E. M., Perryman, L., Hoye, A. M., Agrawal, A., Bird, D., Latif, N. A., Forrest, H., Evans, H. R., Huggins, I. D., Lang, G., Linding, R., Gartland, A. and Erler, J. T. (2015). The hypoxic cancer secretome induces pre-metastatic bone lesions through lysyl oxidase. Nature 522(7554): 106-110.
- DuFort, C. C., Paszek, M. J. and Weaver, V. M. (2011). Balancing forces: architectural control of mechanotransduction. Nat Rev Mol Cell Biol 12(5): 308-319.
- Erler, J. T., Bennewith, K. L., Nicolau, M., Dornhöfer, N., Kong, C., Le, Q. T., Chi, J. T., Jeffrey, S. S. and Giaccia, A. J. (2006). Lysyl oxidase is essential for hypoxia-induced metastasis. Nature 440(7088):1222-1226.
- Humphrey, J. D., Dufresne, E. R. and Schwartz, M. A. (2014). Mechanotransduction and extracellular matrix homeostasis. Nat Rev Mol Cell Biol 15(12): 802-812.
- Kai, F., Laklai, H. and Weaver, V. M. (2016). Force matters: biomechanical regulation of cell invasion and migration in disease. Trends Cell Biol 26(7): 486-497.
- Kasas, S. and Dietler, G. (2008). Probing nanomechanical properties from biomolecules to living cells. Pflugers Arch 456(1): 13-27.
- Levental, K. R., Yu, H., Kass, L., Lakins, J. N., Egeblad, M., Erler, J. T., Fong, S. F., Csiszar, K., Giaccia, A., Weninger, W., Yamauchi, M., Gasser, D. L. and Weaver, V. M. (2009). Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139(5): 891-906.
- Madsen, C. D., Pedersen, J. T., Venning, F. A., Singh, L. B., Moeendarbary, E., Charras, G., Cox, T. R., Sahai, E. and Erler, J. T. (2015). Hypoxia and loss of PHD2 inactivate stromal fibroblasts to decrease tumour stiffness and metastasis. EMBO Rep 16(10): 1394-1408.
- Miller, B. W., Morton, J. P., Pinese, M., Saturno, G., Jamieson, N. B., McGhee, E., Timpson, P., Leach, J., McGarry, L., Shanks, E., Bailey, P., Chang, D., Oien, K., Karim, S., Au, A., Steele, C., Carter, C. R., McKay, C., Anderson, K., Evans, T. R., Marais, R., Springer, C., Biankin, A., Erler, J. T. and Sansom, O. J. (2015). Targeting the LOX/hypoxia axis reverses many of the features that make pancreatic cancer deadly: inhibition of LOX abrogates metastasis and enhances drug efficacy. EMBO Mol Med 7(8): 1063-1076.
- Miller, F. R. and Heppner, G. H. (1979). Immunologic heterogeneity of tumor cell subpopulations from a single mouse mammary tumor. J Natl Cancer Inst 63(6): 1457-1463.
- Picout, D. R. and Ross-Murphy, S. B. (2003). Rheology of biopolymer solutions and gels. ScientificWorldJournal 3: 105-121.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Madsen, C. D. and Cox, T. R. (2017). Relative Stiffness Measurements of Tumour Tissues by Shear Rheology. Bio-protocol 7(9): e2265. DOI: 10.21769/BioProtoc.2265.
Category
Cancer Biology > General technique > Biomechanical assays
Cell Biology > Tissue analysis > Stiffness measurement
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









