- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Surface Inoculation and Quantification of Pseudomonas syringae Population in the Arabidopsis Leaf Apoplast
(*contributed equally to this work) Published: Vol 7, Iss 5, Mar 5, 2017 DOI: 10.21769/BioProtoc.2167 Views: 14333
Reviewed by: Jyotiska ChaudhuriAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
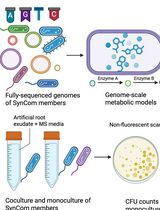
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1705 Views
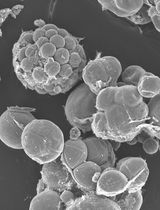
A Simple Protocol for Periodic Live Cell Observation of Flagellate Stages in the Lichen Alga Trebouxia
Enrico Boccato [...] Mauro Tretiach
Jan 20, 2026 229 Views
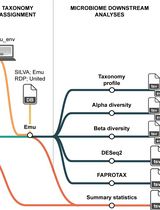
Reproducible Emu-Based Workflow for High-Fidelity Soil and Plant Microbiome Profiling on HPC Clusters
Henrique M. Dias [...] Christopher Graham
Jan 20, 2026 508 Views
Abstract
Bacterial pathogens must enter the plant tissue in order to cause a successful infection. Foliar bacterial pathogens that are not able to directly penetrate the plant epidermis rely on wounds or natural openings to internalize leaves. This protocol describes a procedure to estimate the population size of Pseudomonas syringae in the leaf apoplast after surface inoculation of Arabidopsis rosettes.
Keywords: Leaf inoculationBackground
Plant pathogenic bacteria causing foliar diseases may penetrate the leaf epidermis through wounds and natural openings such as stomata. Stomata are microscopic pores that mediate the regulation of transpiration and the exchange of gases between the plant and the atmosphere. Interestingly, we have demonstrated that bacteria can induce stomatal closure. This phenomenon is now recognized as stomatal defense, which hampers bacterial internalization into the leaf decreasing disease development (reviewed by Melotto et al., 2017, in press). Here, we describe a method adapted from Katagiri et al. (2002) and Panchal et al. (2016a and 2016b) to measure the total endophytic bacterial population of Pseudomonas syringae within Arabidopsis leaf tissue after surface inoculation. This procedure is useful to estimate bacterial penetration of leaves through stomata in a laboratory setting.
Materials and Reagents
- 3.5-inch square pots with holes (Hummert International, catalog number: 12-1300-1 )
- Soil mix, SunGro Sunshine® #1 Mix (Crop Production Services, catalog number: 1000590701 ) or equivalent
- Fine vermiculite (Growers Solution, catalog number: Vermiculite4cf )
- 48 inch x 100 ft Charcoal fiberglass screen (The Home Depot, catalog number: 3000016 )
- 15 ml and 50 ml centrifuge tubes
- Plastic domes (Hummert International, catalog number: 65-6964-1 )
- Sharpie markers, paper towels, Kimwipes, disposable gloves, rubber bands, forceps
- Square Petri dishes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 240835 )
- Micropipettes (Rainin Pipet-LiteTM)
- 1.5 ml microfuge tubes
- Plastic pestles that fit 1.5 ml microfuge tubes (SP Scienceware - Bel-Art Products - H-B Instrument, catalog number: 19923-0001 )
- Plastic trays without holes (Hummert International, catalog number: 65-6963-1 )
- Microbreathe face mask (VWR, catalog number: 10833-224 )
- Arabidopsis thaliana (L. Heyhn.) ecotype Columbia (Col-0, ABRC stock CS60000). Seeds can be stored at 4 °C and are viable for 3-4 years
- Pseudomonas syringae bacterial culture (stored in 25% glycerol at -80 °C)
- Gnatrol (Hummert International, catalog number: 01-2035-1 )
- Arabidopsis controlled release fertilizer (LEHLE SEEDS, catalog number: PM-11 )
- Appropriate antibiotic (e.g., Rifampicin)
- MgCl2 solution
- Glycerol (MP Biomedicals, catalog number: 151194 )
- Silwet L-77 (LEHLE SEEDS, catalog number: VIS-30 )
- Reagent alcohol (Sigma-Aldrich, catalog number: 793183 )
- Sterile distilled water
- Agarose (VWR, catalog number: 97062-250 )
- Tryptone (IBI Scientific, catalog number: 41116105 )
- Yeast extract (U.S.Biotech Sources, catalog number: Y01PD-500 )
- Sodium chloride (NaCl) (Fisher Scientific, catalog number: S271-500 )
- Bacteriological agar (IBI Scientific, catalog number: IB49171 )
- 0.1% agarose (see Recipes)
- Low-sodium Luria Bertani medium (see Recipes)
Equipment
- 1 ml micropipette
- Refrigerator or a cold room
- Plant growth chamber (Caron Products & Services, model: 6341-2 )
- Shaker incubator (VWR, catalog number: 12620-946 )
- Spectrophotometer (Thermo Fisher Scientific, model: Spectronic 20D+ or equivalent)
- Centrifuge (Eppendorf, model: 5810 )
- Handheld electric drill (BLACK + DECKER, catalog number: LDX120C )
- Cork borer No. 2 (Cole-Parmer, catalog number: EW-06298-98 )
- Digital hygrometer (VWR, catalog number: 35519-047 )
- Quantum meter (Apogee, catalog number: BQM )
- Vortex (BioExpress, GeneMate, catalog number: S-3200-1 )
- Autoclave
- laminar flow hood
Procedure
- Growing Arabidopsis plants
- Fill pots with the soil mix and top it off with a 2-cm layer of vermiculite (do not pack too tightly). Cover the pot with 7 x 7-inch fiberglass mesh screen (which prevents the leaves from contacting soil) and hold it in place with a rubber band.
- Place the pots in a tray and soak soil with an aqueous solution of Gnatrol (1 g L-1). Let it sit for one day.
Note: This step is used to avoid development of fungus gnats. - Mix Arabidopsis seeds in 5-10 ml of 0.1% agarose in a 15-ml centrifuge tube. Vortex the suspension for 10 sec at maximum speed to dispense the seeds evenly in agarose prior to sowing.
- Using a 1 ml micropipette, spot seeds (~5 seeds in one spot) in the four corners and the center of the pot. Label each pot with name, date, and genotype of seeds.
- Cover the tray with a plastic dome, leaving an opening of about 5-10 cm. Keep the tray in a cold room (4-8 °C) for 2 days to allow for efficient and synchronous germination (Figure 1A).

Figure 1. Dip-inoculation procedure. A. Arabidopsis seeds are sown on a mesh-covered pot and kept in a cold room (4-8 °C) for 2 days to allow for efficient and synchronous germination. B. Arabidopsis 4-5 weeks old plants are used for inoculation. C. The rosette is dipped and gently swirled in the inoculum. D. Inoculated plants are placed in a tray and incubated in an environment chamber.
Note: If a growth chamber without relative humidity control is used, cover the tray with a plastic dome leaving a 5-10 cm gap open to keep air relative humidity between 60-70%. - After two days, place the tray in growth chamber set at 22 °C, 65 ± 5% relative humidity (RH), and a 12-h photoperiod under photosynthetic active light intensity of 100 µmol m-2 sec-1.
- When seedlings are at the 2-leaf stage, thin the spots with forceps such that only one seedling is left at each spot and all 5 plants in the pot look about the same age. Add 0.2 g fertilizer per liter of soil mix as suggested by the manufacturer. Water is added to the tray as needed; generally, 1 L of water per tray 2 or 3 times per week is sufficient. Do not overwater to avoid moisture buildup, growth of algae, and fungus gnat proliferation.
- Fill pots with the soil mix and top it off with a 2-cm layer of vermiculite (do not pack too tightly). Cover the pot with 7 x 7-inch fiberglass mesh screen (which prevents the leaves from contacting soil) and hold it in place with a rubber band.
- Bacterial inoculum preparation
- From the glycerol stock, streak Pseudomonas syringae from a culture stock on low-salt Luria Bertani (LS-LB) medium supplemented with appropriate antibiotic (e.g., Pseudomonas syringae pv. tomato strain DC3000 is cultured in media containing 100 µg ml-1 of rifampicin). Incubate the plates at 28 °C for approximately 30 h. Once colonies appear on the plate, it can be stored at 4 °C for up to 10 days.
- In the morning of the day before inoculation, prepare the pre-inoculum by inoculating one colony from this fresh plate into 10 ml of LS-LB with appropriate antibiotic. Additionally, incubate 10 ml of LS-LB to use as a control for culture contamination and to set the standard (blank) for the spectrophotometer reading. Incubate the culture tubes at 28 °C and 200 rpm.
- After growing the pre-inoculum for approximately 8 h take the optical density (OD) at 600 nm using a spectrophotometer. To make the inoculum, dilute the pre-inoculum to 300 ml of fresh liquid LS-LB medium and the appropriate antibiotic to obtain an OD600 of 0.004. Use the equation V1 x C1 = V2 x C2. A 10-ml blank should also be included to calibrate the spectrophotometer.
- Incubate at 28 °C and 200 rpm until mid to late log phase is reached (OD600 of 0.8 to 1). It will take approximately 12 h.
- Collect the bacterial cells by centrifugation at 2,600 x g for 20 min at room temperature.
- Resuspend the cell pellet in sterile, distilled water to an OD600 of 0.2, which corresponds to the final inoculum concentration of 1 x 108 CFU ml-1. High inoculum concentration is required to ensure uniform infection of the leaf and avoid sampling problems and large experimental errors. Inoculum must be used immediately to avoid killing bacterial cells. Alternatively, a 1-10 mM MgCl2 solution can be used to resuspend bacterial cells and minimize possible damage associated with osmotic pressure.
- To accurately estimate the inoculum concentration, prepare a 10-time serial dilution of the inoculum and plate each dilution on square plates. See the description of serial dilution procedure in the section below (enumeration of apoplastic bacterium population).
- Add 0.02% Silwet L-77 to the inoculum to ensure that leaves are evenly covered with the inoculum; otherwise, inoculum will runoff and infection will be spotty. This step is required to guarantee reproducibility of the results.
- From the glycerol stock, streak Pseudomonas syringae from a culture stock on low-salt Luria Bertani (LS-LB) medium supplemented with appropriate antibiotic (e.g., Pseudomonas syringae pv. tomato strain DC3000 is cultured in media containing 100 µg ml-1 of rifampicin). Incubate the plates at 28 °C for approximately 30 h. Once colonies appear on the plate, it can be stored at 4 °C for up to 10 days.
- Leaf surface inoculation by dipping
- Use 4-5-week-old Arabidopsis plants with fully expanded leaves for infection (Figure 1B).
- Acclimate plants at 25 °C for approximately 12 h before inoculation. Environmental condition, other than temperature, must be the same as the used for plant growth.
- Dip-inoculate plants as shown in Figure 1C and Video 1. Swirl leaves for 5 sec to make sure all leaves are evenly coated with the inoculum. Drip excess inoculum and place the pot back in the tray (Figure 1D).
Video 1. Dip inoculation of Arabidopsis plants - Add some water in the tray and cover the tray leaving around 5-10 cm gap open such that air relative humidity of 60-65% is maintained.
Note: High relative humidity (> 70%) is not recommended as it may interfere with stomatal defense (Panchal et al., 2016a). It is important to inoculate plants at the same time of the day to ensure reproducibility across biological replicates. It is well documented that the plant immune response and stomatal movement vary with the circadian rhythm (Zhang et al., 2013). - Place the trays in an environmental chamber set at 25 °C, 60-65% RH, and a 12-h photoperiod under photosynthetic active light intensity of 100 µmol m-2 sec-1. If the environmental chamber has a humidity control, the plastic dome is not needed.
- Use 4-5-week-old Arabidopsis plants with fully expanded leaves for infection (Figure 1B).
- Enumeration of apoplastic bacterium population
- Pluck three leaves from a single plant at the petiole making sure the leaf blade is not damaged by the forceps tip. Fully expanded leaves from the second layer from rosette bottom can be used (see healthy looking plants in Figure 2A).
Note: It is highly recommended to do a time course experiment up to 3 days after inoculation. Grow all plants in the same batch for each experiment and use different plants from the same batch for each time point. The time course must include sampling right after inoculation (day 0) to obtain the initial level of total live bacterial population on the plant. For this first time point, leaves should not be sterilized as described in step D2. Do not use the younger (top) or older (bottom) leaves of the rosette, or stressed leaves with curved edges or purpling. For description of Arabidopsis growth stages, please see Boyes et al. (2001).
Figure 2. Enumeration of the bacterial population in the leaf. A. Three equal sized infected leaves are plucked and surface sterilized with 70% ethanol. B. Fours leaf disks are cut from each leaf using a cork borer. C and D. Four leaf disks diluted and 10 μl from each dilution is spotted twice (technical replicates) on square plates. Repeat B-D for each of the three leaves. - Place the leaves in 70% ethanol for 2 min. Next, put the leaves in sterile water for 1 min while rinsing it lightly from time to time in a way that the leaf blade is not damaged.
- Dry the leaves by blotting on paper towel.
- Sterilize the cork borer and forceps at every step from here on by rinsing in 70% ethanol and then in sterile water. Remove water from the cork borer by blotting on the paper towel.
- Using the cork borer number 2, which has an area of 0.125 cm2, punch out four leaf discs (total area of 0.5 cm2), two on either side of the midrib (Figure 2B).
- Use surface sterilized forceps to collect the punched-out leaf discs and put them in 100 µl sterile water in a microfuge tube (represented by the first tube in Figure 2C). Grind the tissue sample using a plastic pestle attached to a hand-held electric drill. No broken leaf pieces should be visible (see Video 2).
Video 2. Leaf tissue sampling and homogenization - Take 10 µl of this first solution and add to 90 µl of sterile water in another microfuge tube making a 1:10 dilution. Similarly, serially dilute until 10-6 for day 0 (Figure 2C) or until 10-8 for subsequent sampling time points.
- Pipet 10 µl from each tube and spot in one square of a Petri dish containing LS-LB medium with appropriate antibiotic (see Figure 2D). Each dilution should be spotted twice as technical replicates (first and second row on the plate shown in Figure 2D). (see Video 3)
- Repeat steps D5 to D8 (Figures 2B-2D) for each of the three leaves (Figure 2A).
- After plating all dilutions, let the plate air dry and incubate it at 28 °C for approximately 30 h.
Video 3. Bacterial enumeration using a serial dilution plating method
- Pluck three leaves from a single plant at the petiole making sure the leaf blade is not damaged by the forceps tip. Fully expanded leaves from the second layer from rosette bottom can be used (see healthy looking plants in Figure 2A).
Data analysis
Count single colonies forming units (CFU) from one of the dilutions (e.g., the fourth or fifth column on the plate illustrated in Figure 2D). Choose the dilution that yields a 10-100 CFU range. Estimate the bacterial population by multiplying the CFU by the dilution factor. To express the value as CFU/cm2, multiply the total CFU count by 2 as the total area of four leaf discs (Figure 2B) is 0.5 cm2. For example, if you count 25 colonies in the dilution lane of 10-5 (5th column), then the bacterial population will be 25 x 105 x 2 = 5 x 106 CFU/cm2. For each sample, there should be three biological replicates (3 leaves; Figure 2A) with 2 technical replicates (2 spots on the medium for each dilution; Figure 2D). Statistical analysis should be done by calculating the average (n = 6) and standard error using Microsoft Excel or any other statistical analysis software. Significance of the difference between two samples can be obtained by performing the Student’s t-test. Additional biological replicates must be performed by repeating the whole experiment with other plants to assess the robustness of the analysis. For scientific rigor, this experimental procedure should be repeated three times and each time should yield similar bacterial growth trends. See examples of data graphs in Panchal et al. (2016a).
Notes
- Be careful not to wound the leaves while picking them with forceps, otherwise the leaves will be squishy and hard to punch holes. Also, wounding will potentially allow 70% ethanol to enter inside the leaf tissue, killing the internal bacterial population and thus giving skewed results.
- For reducing variability in biological replicates, it is extremely crucial that plants are not stressed while growing and during the infection period. Light, temperature, and relative humidity should be kept constant. Acclimatize plants for at least 12 h in the same growth chamber where infected plants are going to be placed.
- Increasing the number of leaves sampled can reduce standard error.
- Growing bacterial cultures beyond OD600 of 1.0 does not give ideal infection results.
- Always use bacterial pathogen from fresh plate streaked from glycerol stock. Multiple sub-culturing may lead to loss of virulence.
Recipes
- 0.1% agarose
Dissolve 0.1 g agarose in 100 ml sterile distilled water by heating
Keep swirling the solution intermittently while it cools
Store and use at room temperature - Low-sodium Luria Bertani medium
10 g/L tryptone
5 g/L yeast extract
5 g/L NaCl
2.5% agar (only for solid medium)
Adjust pH to 7.0
Autoclave medium at 15 psi, 120 °C for 15 min
Allow medium to cool down to about 55 °C and add appropriate antibiotic if needed
Pour medium into plates in a laminar flow hood
Store plates in plastic bags at 4 °C to avoid medium dehydration
Acknowledgments
This work was supported by a grant from the US National Institute of Allergy and Infectious Disease (5R01AI068718) to Dr. Maeli Melotto.
References
- Boyes, D. C., Zayed, A. M., Ascenzi, R., McCaskill, A. J., Hoffman, N. E., Davis, K. R. and Görlach, J. (2001). Growth stage-based phenotypic analysis of Arabidopsis: A model for high throughput functional genomics in plants. Plant Cell 13: 1499-1510.
- Katagiri, F., Thilmony, R. and He, S. Y. (2002). The Arabidopsis thaliana-Pseudomonas syringae interaction. Arabidopsis Book 1: e0039.
- Melotto, M., Zhang, L., Oblessuc, P. R. and He, S. Y. (2017). Stomatal defense a decade later. Plant Physiol (in press).
- Panchal, S., Chitrakar, R., Thompson, B. K., Obulareddy, N., Roy, D., Hambright, W. S. and Melotto, M. (2016a). Regulation of stomatal defense by air relative humidity. Plant Physiol 172(3): 2021-2032.
- Panchal, S., Roy, D., Chitrakar, R., Price, L., Breitbach, Z. S., Armstrong, D. W. and Melotto, M. (2016b). Coronatine facilitates Pseudomonas syringae infection of Arabidopsis leaves at night. Front Plant Sci 7: 880.
- Zhang, C., Xie, Q., Anderson, R. G. Ng, G., Seitz, N. C., Peterson, T., McClung, C. R., McDowell, J. M., Kong, D., Kwak, J. M. and Lu, H. (2013). Crosstalk between the circadian clock and innate immunity in Arabidopsis. PLoS Pathogens 9(6): e1003370.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Jacob, C., Panchal, S. and Melotto, M. (2017). Surface Inoculation and Quantification of Pseudomonas syringae Population in the Arabidopsis Leaf Apoplast. Bio-protocol 7(5): e2167. DOI: 10.21769/BioProtoc.2167.
Category
Plant Science > Plant immunity > Host-microbe interactions
Microbiology > Microbe-host interactions > Bacterium
Cell Biology > Cell isolation and culture > Cell growth
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












