- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Protocol for Molecular Dynamics Simulations of Proteins
Published: Vol 6, Iss 23, Dec 5, 2016 DOI: 10.21769/BioProtoc.2051 Views: 27104
Reviewed by: Arsalan DaudiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
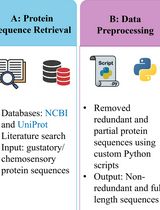
A Step-by-Step Computational Protocol for Functional Annotation and Structural Modelling of Insect Chemosensory Proteins
Rajeswari Kalepu [...] Nor Azlan Nor Muhammad
Nov 20, 2025 1709 Views
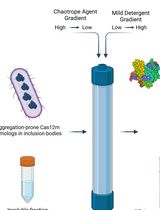
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 194 Views
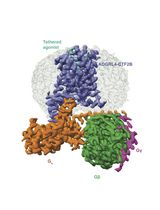
Purification of the Active-State G Protein-Coupled Receptor ADGRL4 for Cryo-Electron Microscopy Using a Modular Tag System and a Tethered mini-Gq
David M. Favara and Christopher G. Tate
Mar 5, 2026 272 Views
Abstract
Molecular dynamics (MD) simulations have become one of the most important tools in understanding the behavior of bio-molecules on nanosecond to microsecond time scales. In this protocol, we provide a general approach and standard setup protocol for MD simulations by using the Gromacs MD suite.
Keywords: Molecular dynamics simulationsBackground
While molecular dynamics (MD) simulations are increasingly getting popular in studying protein dynamics in silico, there is a strong need to correlate the results with experimental observations. It is necessary that the protein model and the chosen environment for the simulations should mimic the native environment as close as possible. In general, there are three key stages in molecular dynamics simulations: Setup, production run, and analysis of the trajectories. The setup includes the input structures, parameters, force-fields and topologies. However, the readymade setup and wrong parameters could adversely affect the outcome making a false assessment of the biological interpretation. In view of a manual setup as a better choice to setup simulations, we through this protocol, provide a general approach and standard setup protocol for MD simulations. Many online/offline tools are available to perform MD simulations. One amongst the open source tools is Gromacs (Abraham et al., 2015; Pronk et al., 2013; Van der Spoel et al.,2005), a robust and popular MD simulations suite available today which supports almost all the major force fields. In this protocol we also refer to simulations of membrane proteins previously performed in vacuum as a temporary alternative to simulations including the membrane.
Materials and Reagents
- Protein structure coordinates (http://www.rcsb.org/pdb/home/home.do)
- Appropriate Force field (A force field describes physical systems as collections of atoms kept together by interatomic force, e.g., chemical bonds, angles etc. [Meller, 2010]).
- Molecular geometry file(.gro)
- Molecular topology file(.top)
- Parameter files (.mdp) (http://manual.gromacs.org/online/mdp.html)
Equipment
Note: In general, MD simulations require parallel computing to be able to run longer simulations in a shorter period. For smaller jobs/preprocessing, a desktop workstation machine preferably running on Linux is sufficient. However, it is preferable to perform initial steps on a local machine and final md run on a computer cluster.
- A desktop PC with configuration below was used for initial setup of simulations :
- HP Pavilion Desktop Intel 4 Core(TM) i5-4570 CPU@3.2GHz each
- 16 GB memory
- 1 TB HDD
- NVIDIA GeForce GT 625
- Ubuntu V15.04
- The basic configuration of the supercomputer is as follows:
- Site: Center for Development of Advanced Computing (C-DAC)
- Cores: 30,056
- Memory: 14,144 GB
- Processor: Xeon E5-2670 8C 2.6GHz
- Operating System: CentOS
- MPI: Intel MPI
- Site: Center for Development of Advanced Computing (C-DAC)
Software
- Gromacs MD simulation suite v 5.1, (Abraham et al., 2015; Pronk et al., 2013; Van der Spoel et al., 2005)
- Rasmol (http://www.bernstein-plus-sons.com/software/rasmol/README.html) for molecular visualization
- Text editor, Gedit 3.18 (to edit the PDB file, update topology files and to edit the input run parameter files)
- 2D plotting program Grace (http://plasma-gate.weizmann.ac.il/Grace) (for visualization )
- Win SCP (to transfer files to and fro)
- SSH client/Putty (to execute the commands on a remote server from our desktop)
Procedure
A schematic representation of the workflow is shown in Figure 1. The key steps along with Gromacs commands are explained below.
Figure 1. Steps to perform MD simulations in general (Gajula et al., 2007; 2008; 2013; 2015; Abé et al., 2011)
- Obtain the protein coordinates
The basic ingredient to start MD simulations is a protein structure coordinate file in PDB format that can be downloaded from RCSB website (http://www.rcsb.org/pdb/home/home.do). RasMol tool is used for visual inspection of the protein structure and also for graphics rendering. Gedit tool is used to open and edit the text files.
Note: The Protein Databank (http://www.pdb.org/) contains thousands of high resolution protein structures that are derived by X-ray/neutron scattering and NMR methods. When the protein structure of our interest is not available, a homology model as an initial starting structure may be built by a variety of software tools like Modeller, iTassar, Discovery Studio, Pepbuild web server etc. However, once the PDB is available and downloaded, it is necessary to pre-format it depending on the presence of ligands or water molecules in the structure. It occurs that ligand coordinates are present in the PDB file and the GROMACS software doesn’t recognize the ligand. In this case, ligand chemistry needs to be explicitly defined by extracting the coordinates of the ligand and separated from main PDB file. A separated topology is constructed manually for the ligand and the information is added in the main topology file. It is also advised to remove external water molecules that are present in the PDB file.
For this study, we have prepared a protein coordinate file named ‘protein.pdb’. And, every MD simulations software follows their specific format to handle the files. First, the PDB file is converted into a GROMACS specific molecular geometry file format (.GRO) by using the pdb2gmx command as shown below. The <.GRO> file contains the molecular structure coordinates with continuous atom numbering. Moreover, pdb2gmx command also generates a very important file called topology with ‘.top’ extension. The topology file contains the molecular description such as molecular parameters, bonding, force field and charges etc. It is noticeable that the pdb2gmx program adds the missing hydrogen atoms in the structure by default.
pdb2gmx -f protein.pdb -p protein.top -o protein.gro
where,
-f flag reads the pdb file,
-p flag is used for output of the topology file,
-o flag is used to write into the Gromacs coordinate format.
Note: The pdb2gmx prompts you to select an appropriate force field for the simulations. For example, in Gromacs v 5.1, a force field ‘ffG53A7’ is recommended for the simulations of proteins with explicit solvent. - Periodic Boundary Conditions (PBC): Setup the box for simulations
Periodic Boundary Conditions were applied for the protein to be simulated to keep track of the motion of all particles and to avoid/minimize the edge effect on the surface atoms. For this purpose, a box (supercell) is defined, and that supercell is surrounded by infinitely replicated, periodic images of itself (Li, 2005). Therefore a surface particle may interact not only with particles in the same supercell but also with particles in adjacent image supercells (Li, 2005).
The command to generate a cubic box around the protein with a box edge of approximately 14A° from the proteins periphery is:
editconf -f protein.gro -o protein_editconf.gro -bt cubic -d 1.4 –c
The -c flag keeps the protein in the center of the box. The default measurement units are represented in nanometers. There are also other box types available, e.g., Triclinic, dodecahedron, rhombic dodecahedron and octahedron. While cubic is a rectangular box with all sides equal, whereas dodecahedron represents a rhombic dodecahedron, a truncated version of octahedron.
Note: An ideal value for the distance is at least 1.0Å or more for most systems. - Solvate the box
To mimic the physiological environment for a protein, the simulation system (box around the protein) need to be solvated. This step is not necessary for simulations in vacuum. However, it is an essential step for the simulations of protein both in water and/or membrane. To solvate the protein, the ‘solvate’ command needs to be used. The ‘solvate’ command adds the required number of water molecules around the protein based upon the dimensions/box type that is specified in ‘editconf’ of step 2 above.
gmx solvate -cpprotein_editconf.gro -p protein.top -o protein _water.gro
Note: The topology (.top) file is updated and now it contains the topology of water molecules in addition to protein topology.
It is necessary to neutralize the system prior to simulations. For that it is necessary to add counter ions according to the total charge of the system by using ‘genion’ command. Prior to use ‘genion’ command, a gromacs pre-processor file has to be generated which contains the atomic description + topology of the molecule being studied. - Preprocessing
The preprocessing in Gromacs is done by using ‘grompp’ command that is used to collect parameters, topology, and coordinates into a single run input file for next step, e.g., protein_b4em.tpr as shown below. It is in binary format. Once the .tpr file is generated, it is possible that the simulation can be run from any computer/cluster.
grompp -f em.mdp -c protein_water -p protein.top -o protein_b4em.tpr
where,
-f flag requires a parameter file, usually denoted as ‘.mdp’ file that can be obtained from Gromacs website (http://manual.gromacs.org/online/mdp.html).
The generated .tpr file then passed into ‘genion’ command as shown below:
genion -s protein_b4em.tpr -o protein_genion.gro -nn 3 -nq -1 -n index.ndx
Here,
The flag -s is used to input the pre-processed file generated by ‘grompp’,
-o flag is to output after neutralizing the system,
-nn for number of counter ions to be added,
-nq for the charge of the ion (in case of positive ions -np 1 should be used).
For example, in the case the system has a net charge of + 3.00. Therefore, it is needed to add three chloride ions to neutralize the overall charge. The index.ndx can be used to merge the groups and to setup up coupling groups, let’s say ‘Protein and Ligand’ as one group. - Energy minimization
In general, energy minimization is commonly used to refine low-resolution experimental structures and to remove steric hindrance that may be caused due to added hydrogen atoms. Gromacs supports different minimization algorithms: the most commonly used are steepest descent and conjugate gradient. The steepest descent algorithm is the quickest in removing the largest strains in the system but converges slowly when close to a minimum.
The ‘grompp’ command processes and prepares the input file necessary for energy minimization run.
grompp -f em -c protein_genion.gro -p protein.top -o protein_em.tpr
Once, the .tpr file is ready, submit the job for energy minimization by following command:
mdrun -s protein_em.tpr -e minim_ener.edr -o protein_em.trr -c protein_b4md.gro
Here the mdrun command is used to perform energy minimization. Mdrun reads the input file with -s flag and generates four output files after successful execution. The energy file (.edr) contains state variables temperature, pressure, etc., and much more information that can be used for further analysis of simulation data. The trajectory file is generated by specifying ‘-o’, flag and it contains coordinates and velocities. The structure file (-c) contains the final coordinates and velocities of the last step of MD simulations. A ‘.log’ file is generated by default. It is to be noted that the same mdrun command is used to perform final molecular dynamics simulations as well. However, the input parameter (.mdp) file needs to be edited accordingly.
Note: There is another step called ‘position restraining’ before we go for final MD setup. The process is the same as in step 5. This is required to allow the water molecules to adjust with the protein system. Here we restrict the positions of protein atoms and allow the water molecules to flow. Position restraint run is not necessary for simulations in vacuo. The output of this is then passed into the ‘mdrun’ command, which actually performs the simulation. - MD simulations
Now we enter into the final phase of MD simulations. The parameter file (md.mdp) needs to be adjusted accordingly. The difference between position restraint run/equilibration and production run is minimal: For production run, the position restraints and pressure coupling are turned off and simulations are carried significantly longer, say 1ns to 1ms depends on the research question and availability of resources.
grompp -f md.mdp -c protein_b4md.gro -p protein -o protein _md.tpr
mdrun -nice 4 -s protein _md.tpr -o protein _md.trr -g md_log.log -e ener_protein _md.tpr -c protein _aft_md–v
Here ‘-v’ displays the live flow of execution on your monitor. After completing a simulation with ‘mdrun’, four output files are generated same as in earlier case; these are a trajectory file (.trr) and an energy file (.edr), structure file(.gro) and log (.log) file. These files can be used for further analysis. However, it is necessary to relax the system first before making the production run.
Note: A most common issue with MD simulations run is abnormal termination of the job. The end user may tend to repeat the whole procedure. However, if the termination is due to power failure or queue limits etc., then the run can be continued from where it has stopped by using a command called ‘tpbconv’. Moreover, the same command can be used to extend the simulations further for more time period. The usage of tpbconv is similar to ‘grompp’. Instead of grompp, use ‘tpbconv’ command to produce input file required for mdrun. It is required to have the last checkpoint file ‘.cpt’ in order to extend the run.
tpbconv -s previous_md.tpr -extend (new time to be extended) -o new.tpr
The command flags are same as in steps above. Now, issue the final mdrun command as shown below:
mdrun -s new.tpr -cpiprevious_run.cpt -o protein_new_md.trr -g new_md_log.log-enew_ener -c protein _new_md–v
In general the final mdrun is performed on computer cluster or super computer. The remote servers can be connected via ssh clients like putty from windows desktop machines and the file transfers can be performed by using WinSCP tool.
Data analysis
- The MD simulation run in general generates five important output files.
- <.trr>: The full precision trajectory containing the positions, velocities and forces over time
- <.xtc>: A light weight trajectory, containing only coordinates in low precision (0.001 nm)
- <.edr>: Energy related parameters over time
- <.log>: A file containing information about the simulation
- <.gro>: A final structure file
- After successful completion of the final production run it is necessary to analyze the trajectories. In general, the trajectory analysis is done in three phases. First, to perform some standard checks to assess the quality of the simulation runs. The system should have attained equilibrium during the simulations to be able to consider for further analysis of the trajectory. This is often checked by looking at quantities such as the temperature vs. time; total energy vs. time etc. by using the GRACE tool. If the results from these analyses are satisfactory, then some basic calculations were measured before actually trying to address the actual research question. One of the most fundamental properties to determine is whether the protein is close to the experimental structure during and after simulations. This is done by calculating the root-mean-square deviation (RMSD) of the backbone atoms with respect to the X-ray structure. In the final phase, the results from different simulations can be combined and analyzed further. Please refer to (Gajula et al., 2013a; 2013b; Kumar et al., 2013; Gajula et al., 2015; Kumar et al., 2016) for detailed MD simulations data analysis procedure. It is also reported that the computer simulations data is comparable to experimental data for the better understanding of the structure and dynamics of proteins (Gajula et al., 2008; Gajula et al., 2013a; 2013b; Gajula et al., 2015).
Notes
The most important input files manually prepared for the simulations setup are the ‘parameter files’ with ‘.mdp’ extension. For the energy minimization step, usually we denote it with name em.mdp; and position restraining ‘pr.mdp’ and for final production run we may name it as ‘md.mdp’. The standard and optimum average values to be used are provided below. However, it is necessary and at the discretion of the user to prepare these files based on the research question and after careful review of literature.
em.mdp
; em.mdp - used as input into grompp to generate em.tpr
integrator = steep ; Algorithm (steep = steepest descent minimization)
emtol = 1000.0 ; Stop minimization when the maximum force < 1000.0 kJ/mol/nm
emstep = 0.01 ; Energy step size
nsteps = 50000 ; Maximum number of (minimization) steps to perform
; Parameters describing how to find the neighbors of each atom and how to calculate the interactions
nstlist = 1 ; Frequency to update the neighbor list and long range forces
cutoff-scheme = Verlet
ns_type = grid ; Method to determine neighbor list (simple, grid)
coulombtype = PME ; Treatment of long range electrostatic interactions
rcoulomb = 1.0 ; Short-range electrostatic cut-off
rvdw = 1.0 ; Short-range Van der Waals cut-off
pbc = xyz ; Periodic Boundary Conditions (yes/no)
md.mdp
title = md1n ; MD simulation job name
; Run parameters
integrator = md ; leap-frog integrator
nsteps = 500000 ; 2 * 500000 = 1000 ps (1 ns)
dt = 0.002 ; 2 fs
; Output control
Nstxout = 5000 ; save coordinates every 10.0 ps
Nstvout = 5000 ; save velocities every 10.0 ps
Nstenergy = 5000 ; save energies every 10.0 ps
Nstlog = 5000 ; update log file every 10.0 ps
nstxout-compressed= 5000 ; save compressed coordinates every 10.0 ps
; nstxout-compressed replaces nstxtcout
compressed-x-grps= System; replaces xtc-grps
; Bond parameters
Continuation = yes ; Restarting after NPT
constraint_algorithm= lincs ; holonomic constraints
constraints = all-bonds ; all bonds (even heavy atom-H bonds) constrained
lincs_iter = 1 ; accuracy of LINCS
lincs_order = 4 ; also related to accuracy
; Neighborsearching
cutoff-scheme= Verlet
ns_type = grid ; search neighboring grid cells
nstlist = 10 ; 20 fs, largely irrelevant with Verlet scheme
rcoulomb = 1.0 ; short-range electrostatic cutoff (in nm)
rvdw = 1.0 ; short-range van der Waals cutoff (in nm)
; Electrostatics
coulombtype = PME ; Particle Mesh Ewald for long-range electrostatics
pme_order = 4 ; cubic interpolation
fourierspacing= 0.16 ; grid spacing for FFT
; Temperature coupling is on
Tcoupl = V-rescale ; modified Berendsen thermostat
tc-grps = Protein Non-Protein; two coupling groups - more accurate
tau_t = 0.1 0.1 ; time constant, in ps
ref_t = 300 300 ; reference temperature, one for each group, in K
; Pressure coupling is on
Pcoupl = Parrinello-Rahman; Pressure coupling on in NPT
Pcoupltype = isotropic ; uniform scaling of box vectors
tau_p = 2.0; time constant, in ps
ref_p = 1.0; reference pressure, in bar
compressibility= 4.5e-5 ; isothermal compressibility of water, bar^-1
; Periodic boundary conditions
pbc = xyz ; 3-D PBC
; Dispersion correction
DispCorr = EnerPres ; account for cut-off vdW scheme
; Velocity generation
gen_vel = no ; Velocity generation is off
Acknowledgments
This protocol was adapted from the previously published studies by MNV Prasad Gajula (2007; 2008; 2013) by using Gromacs and it was performed by MNV Prasad Gajula (2011; 2013; 2015; 2016). This work was supported by SERB, Department of Science and Technology, India through Ramanujan Fellowship SR/S2/RJN-22/2011. Our sincere thanks to Prof. Dr. H.-J. Steinhoff and Dr. Prashanth Suravajhala for critical reviewing and kindly editing the manuscript. Sincere thanks to Dr. C. Cheralu, the director of Institute of Biotechnology, PJTSAU for his extended support. We highly acknowledge ‘C-DAC, India’ for providing the computational facilities.
References
- Abraham, M.J., Murtola, T., Schulz, R., Páll, S., Smith, J.C., Hess, B. and Lindahl, E. (2015). GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1(2):19-25.
- Abé, C., Dietrich, F., Gajula, P., Benz, M., Vogel, K. P., van Gastel, M., Illenberger, S., Ziegler, W. H. and Steinhoff, H. J. (2011). Monomeric and dimeric conformation of the vinculin tail five-helix bundle in solution studied by EPR spectroscopy. Biophys J 101(7): 1772-1780.
- Borovykh, I. V., Ceola, S., Gajula, P., Gast, P., Steinhoff, H. J. and Huber, M. (2006). Distance between a native cofactor and a spin label in the reaction centre of Rhodobacter sphaeroides by a two-frequency pulsed electron paramagnetic resonance method and molecular dynamics simulations. J Magn Reson 180(2): 178-185.
- Gajula, M.P. (2008). Computer simulation meets experiment: Molecular dynamics simulations of spin labeled proteins. PhD Thesis. Osnabrueck, urn:nbn:de:gbv: 700-2008041631.
- Gajula, M. P., Borovykh, I. V., Beier, C., Shkuropatova, T., Gast, P. and Steinhoff, H. J. (2007). Spin-labeled photosynthetic reaction centers from Rhodobacter sphaeroides studied by electron paramagnetic resonance spectroscopy and molecular dynamics simulations. Appl Magn Reson 31(1-2): 167-178.
- Gajula, M. P., Milikisyants, S., Steinhoff, H. J. and Huber, M. (2007). A short note on orientation selection in the DEER experiments on a native cofactor and a spin label in the reaction center of Rhodobacter sphaeroides. Appl Magn Reson31(1-2): 99-104.
- Gajula, M. P., Soni, G., Babu, G., Rai, A. and Bharadvaja, N. (2013a). Molecular interaction studies of shrimp antiviral protein, PmAV with WSSV RING finger domain in silico. J Appl Bioinform Comput Biol 2:10-17.
- Gajula, M. P., Steinhoff, H. J., Kumar, A., Siddiq, E. A., Polumetla, A. K. and Lendzian, F. (2015). Displacement of the tyrosyl radical in RNR enzyme: A sophisticated computational approach to analyze experimental data. BICOB: 211-219.
- Gajula, M. P., Vogel, K. P., Rai, A., Dietrich, F. and Steinhoff, H. J. (2013b). How far in-silico computing meets real experiments. A study on the structure and dynamics of spin labeled vinculin tail protein by molecular dynamics simulations and EPR spectroscopy. BMC genomics 14(2): 1.
- Li, J. (2005). Basic molecular dynamics. In Lu, G. (Ed.). Handbook of Materials Modeling. Springer Netherlands, pp: 565-588.
- Kumar, A., Kumar, S., Kumar, U., Suravajhala, P. and Gajula, M. P. (2016). Functional and structural insights into novel DREB1A transcription factors in common wheat (Triticum aestivum L.): A molecular modeling approach. Comput Biol Chem 64: 217-226.
- Kumar, A., Mishra, D. C., Rai, A., Sharma, M. and Gajula, M. P. (2013). In silico analysis of protein-protein interaction between resistance and virulence protein during leaf rust disease in wheat (Triticum aestivum L.). World Res J Pept Protein 2(1): 52-58.
- Meller, J. (2010). Molecular Dynamics. eLS 5.
- Pronk, S., Pall, S., Schulz, R., Larsson, P., Bjelkmar, P., Apostolov, R., Shirts, M. R., Smith, J. C., Kasson, P. M., van der Spoel, D., Hess, B. and Lindahl, E. (2013). GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29(7): 845-854.
- Van Der Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E. and Berendsen, H. J. (2005). GROMACS: fast, flexible, and free. J Comput Chem 26(16): 1701-1718.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Gajula, M. P., Kumar, A. and Ijaq, J. (2016). Protocol for Molecular Dynamics Simulations of Proteins. Bio-protocol 6(23): e2051. DOI: 10.21769/BioProtoc.2051.
Category
Systems Biology > Interactome > Protein-protein interaction
Biochemistry > Protein > Structure
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













