- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measuring Oxygen Consumption Rate in Caenorhabditis elegans
Published: Vol 6, Iss 23, Dec 5, 2016 DOI: 10.21769/BioProtoc.2049 Views: 11292
Reviewed by: Jyotiska ChaudhuriJian ChenAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
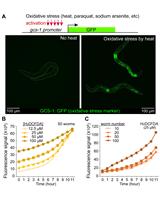
Measurement of Intracellular ROS in Caenorhabditis elegans Using 2’,7’-Dichlorodihydrofluorescein Diacetate
Dong Suk Yoon [...] Dong Seok Cha
Mar 20, 2018 16728 Views
Oxygen Consumption Measurements in Caenorhabditis elegans Using the Seahorse XF24
Suraiya Haroon and Marc Vermulst
Jul 5, 2019 6754 Views

Fluorescent Polysome Profiling in Caenorhabditis elegans
Dan Shaffer and Jarod A Rollins
Sep 5, 2020 7660 Views
Abstract
The rate of oxygen consumption is a vital marker indicating cellular function during lifetime under normal or metabolically challenged conditions. It is used broadly to study mitochondrial function (Artal-Sanz and Tavernarakis, 2009; Palikaras et al., 2015; Ryu et al., 2016) or investigate factors mediating the switch from oxidative phosphorylation to aerobic glycolysis (Chen et al., 2015; Vander Heiden et al., 2009). In this protocol, we describe a method for the determination of oxygen consumption rates in the nematode Caenorhabditis elegans.
Keywords: AgeingBackground
Recent evidence underlines mitochondrial function as a potential contributor in the maintenance of organismal homeostasis and viability (Vafai and Mootha, 2012). Cellular oxygen consumption is highly recognized as a fundamental indicator of mitochondrial function, reflecting reactive oxygen species (ROS) production and metabolic activity of the cell. Therefore, several methods have been developed to measure oxygen consumption rates in cells or entire organisms (Dranka et al., 2011; Li and Graham, 2012; Luz et al., 2015; Perry et al., 2013). These approaches provided insight into the pivotal roles of mitochondria in disease progression and pathogenesis (Scheibye-Knudsen et al., 2015). In this protocol, we describe a method for the determination of oxygen consumption rates in the nematode C. elegans by using a Clark-type polarographic oxygen sensor electrode (Hansatech, King’s Lynn, England).
Materials and Reagents
- Consumables
- Commercial cigarette paper
- Polytetrafluorethylene (PTFE) membrane (provided by Hansatech, King’s Lynn, England)
- 15 ml tube (SARSTEDT, catalog number: 62.554.016 )
- 1.5 ml tube (Sigma-Aldrich, catalog number: Z606340 )
- Paper towel
- Greiner Petri dishes (60 x 15 mm) (Greiner Bio One, catalog number: 628161 )
- Maintenance kit (Hansatech, King’s Lynn, England)
- Commercial cigarette paper
Biologicalreagents
- C. elegans strains (wild type [N2] and dct-1[tm376])
- Escherichia coli OP50 strain (obtained from the Caenorhabditis Genetics Center)
- C. elegans strains (wild type [N2] and dct-1[tm376])
- Chemical reagents
- Distilled water
- Nitrogen gas provided in a tank
- PierceTM BCA Protein Assay Kit (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 23225 )
- Potassium dihydrogen phosphate (KH2PO4) (EMD Millipore, catalog number: 1048731000 )
- K2HPO4
- Sodium chloride (NaCl) (EMD Millipore, catalog number: 1064041000 )
- BactoTM peptone (BD, BactoTM, catalog number: 211677 )
- Streptomycin (Sigma-Aldrich, catalog number: S-6501 )
- Agar (Sigma-Aldrich, catalog number: 05040 )
- Cholesterol stock solution (SERVA Electrophoresis, catalog number: 17101.01 )
- Calcium chloride (CaCl2) (Sigma-Aldrich, catalog number: C-5080 )
- Magnesium sulfate (MgSO4) (Sigma-Aldrich, catalog number: M-7506 )
- Nystatin stock solution (Sigma-Aldrich, catalog number: N-3503 )
- Na2HPO4 (EMD Millipore, catalog number: 1065860500 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P-5405 )
- Potassium chloride (KCl) buffer (see Recipes)
- Nematode growth medium (NGM) agar plates (see Recipes)
- M9 buffer (sterile; see Recipes)
- Phosphate buffer (sterile; see Recipes)
- Distilled water
Equipment
- Dissecting stereomicroscope (Olympus, model: SMZ645 )
- Incubators for stable temperature (AQUA®LYTIC incubator 20 °C)
- DW1/AD Clark-type polarographic oxygen sensor (Hansatech Instruments, model: Oxygraph Plus System )
- Water bath at 20 °C
- Tabletop centrifuge (Eppendorf, model: 5424 )
- Sonicator (Sonics & Material, model: VC 130PB )
Software
- Oxygraph Plus software (Hansatech, King’s Lynn, England)
- Microsoft Office 2011 Excel
Procedure
- Growth and synchronization of nematode population
- Use a dissecting stereomicroscope to select L4 larvae of each strain. Place 4-6 L4 larvae on a freshly Escherichia coli (OP50) seeded NGM plate (Figures 1A-1C). Use at least three plates for each nematode strain.
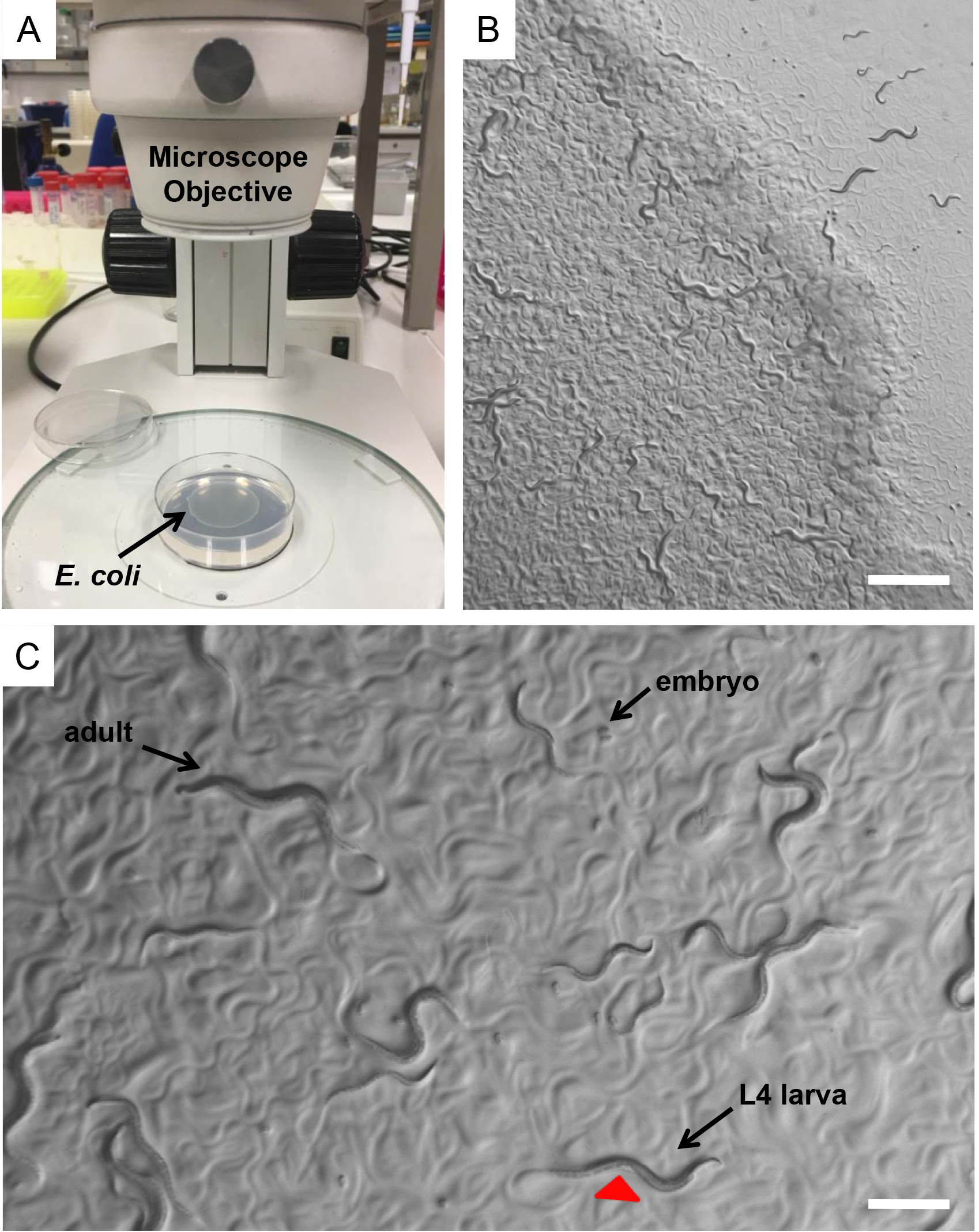
Figure 1. Caenorhabditis elegans in the laboratory. A. E. coli (OP50) seeded NGM plates on the base of a dissecting stereomicroscope. Bacterial lawn is visible on the surface of the agar. B. A mix population of nematodes observed through the dissecting stereomicroscope. Nematodes leave tracks on the plate surface indicating their movements on the bacterial lawn. Scale bar = 1 mm. C. During development worms increase in size throughout four larval stages (L1, L2, L3 and L4). L4 hermaphrodite nematodes can be distinguished by the developing vulva (red arrowhead), which is a clear half circle in the center of the ventral side. Scale bar = 0.5 mm. - Incubate the worms at the standard temperature of 20 °C.
- Four days later the plates contain mixed nematode population (Figure 1B).
- Synchronize nematodes by picking L4 larvae of each strain under a dissecting stereomicroscope and transfer them onto separate plates.
- Add 20-25 L4 larvae per plate. For each experimental condition, use at least five plates.
- Use a dissecting stereomicroscope to select L4 larvae of each strain. Place 4-6 L4 larvae on a freshly Escherichia coli (OP50) seeded NGM plate (Figures 1A-1C). Use at least three plates for each nematode strain.
- Set up the electrode disc and the electrode chamber
- Read the manual to assemble properly the electrode chamber according to the manufacturers’ instructions.
- Cut a 1.5 cm2 paper spacer (cigarette paper) and a similar size piece of polytetrafluorethylene (PTFE) membrane (Figure 2A; Video 1).

Figure 2. Oxygen consumption rate in C. elegans. A. Spare parts of oxygraph apparatus: 1. Oxygraph electrode control unit, 2. Oxygraph chamber (top plate, water jacket, borosilicate glass reaction vessel and base ring, 3. Polytetrafluorethylene (PTFE) membrane, 4. Commercial cigarette papers, 5. Membrane applicator, 6. Electrode disc, 7. Small and large O-rings, 8. Standard plunger assembly, 9. Oxygraph Plus software, 10. Electrode maintenance kit (cotton buds and rapid Hansatech polishing paste). B. Slopes depict oxygen consumption rate upon measurements of wild type (2, 4, 6) and dct-1(tm376) (1, 3, 5) animals. C. Oxygen consumption rates normalized to total protein content. DCT-1 depleted animals display increased oxygen consumption levels (*** P < 0.001; unpaired t-test). Error bars denote SEM.Video 1. Preparation of electrode disc and chamber during oxygen consumption measurement - Place a small droplet of potassium chloride (KCl) buffer on the top of the dome of the electrode disc (Video 1).
- Place the cigarette paper on the center of the dome and cover it with the similar size piece of PTFE membrane (Figure 2A; Video 1).
- Place the small O-ring over the dome and then place the large O-ring around (Figure 2A; Video 1). If the large O-ring is not placed, when the disc is installed in the electrode chamber, the measurements will be affected by ambient air due to impaired sealing (see Note 1; Video 1).
- Once the electrode disc has been successfully prepared, check the response of the disc prior to any experiment. Connect the electrode disc to the control unit.
- Open the software and start recording. A few minutes later the signal will be stabilized and would be around 2,000 mV in air.
- Test the electrode disc by breathing on the electrode. A steep drop of the signal should be observed due to decreased oxygen levels. Then, the signal should be returned to the original levels as the ambient oxygen equilibrates around the electrode disc.
- If the signal does not respond, it may be caused by an inadequate electrode preparation. Then, disassemble the electrode disc, clean the electrode and repeat the procedure described above (steps B1-B8).
- Read the manual to assemble properly the electrode chamber according to the manufacturers’ instructions.
- Liquid phase calibration of the electrode disc
- Install the prepared electrode disc into the electrode chamber (Figure 2A, Video 1).
- Place and connect the electrode chamber on to the rear of the control unit (Figure 2A).
- Place 2 ml sterilized distilled water into the reaction vessel of the chamber.
- Water should be equilibrated to the assay temperature before calibration process. Set up the water bath at desired temperature and connect it with the electrode chamber to maintain stable temperature during experiments (see Note 2).
- Open Oxygraph Plus software and set up the appropriate temperature and atmospheric pressure.
- Turn on stirrer to provide smooth stirring of the sample avoiding bubbles generation, which could cause noisy signals.
- Wait until a plateau of the signal has been reached.
- Establish zero oxygen conditions. Bubble nitrogen gas into the reaction vessel of the chamber to get rid all the oxygen out of the sample. Wait until the signal has reached a plateau.
- Add 1 ml M9 buffer into the reaction vessel of the chamber and wait until system equilibration.
- Install the prepared electrode disc into the electrode chamber (Figure 2A, Video 1).
- Assess oxygen consumption rate of the samples
- Enable stirring throughout the duration of the experimental process.
- Wash off the NGM plates using M9 buffer and collect nematodes in a 15 ml tube.
- Let the animals to settle with gravity for few minutes. Use adult worms. Remove most of the supernatant, which contains bacteria, eggs and larvae (L1, L2, L3 and L4). Repeat this step two more times. After the last wash, reduce the volume to ~1.5 ml.
- Remove M9 buffer from the reaction vessel of the chamber (see Note 3).
- Add 1 ml nematode suspension into the reaction vessel of the chamber.
- Monitor oxygen consumption for 5-10 min.
- Prepare the next sample. Wash off other plates containing different strains or treated animals. Repeat steps D3-D5.
- Recover the animals thoroughly from the reaction vessel and place them in a 1.5 ml tube (see Note 3).
- Place the tube on ice. The tubes should be kept on ice until protein determination.
- Measure the next sample. Repeat steps D5-D9 (see Note 4).
- Save the recorded data from Oxygraph Plus software (Figure 2B).
- Collect the samples from ice and proceed to worm lysis and protein determination.
- Insert sonicator tip into sample and sonicate each sample using ten (10) pulses at 70% power each time. Immediately place the samples on ice (see Note 5).
- Spin down samples at 14,800 x g for 10 min at 4 °C.
- Transfer supernatants (solubilized proteins) into new 1.5 ml tubes and measure protein concentration using a standard protein kit, such as PierceTM BCA Protein Assay Kit, following the instructions of the manufacturer (Estimated protein concentration range: 0.05-0.9 μg).
- Open and review the data (obtained during step D11) in Microsoft Office 2011 Excel (Figure 2B).
- Divide the rate of the negative slope (oxygen consumption; obtained during step D6) with the total protein amount (nmol/min/ml/mg) in Microsoft Office 2011 Excel software (Figure 2C).
- Subject the data to further and more advanced statistical analysis (Figure 2C).
- Enable stirring throughout the duration of the experimental process.
- Electrode maintenance and storage
- Clean the electrode disc after use.
- Use the small cotton bud and the polishing paste from the maintenance kit.
- Grip gently and remove the brown/black deposition of silver chloride and oxidized salt (KCl) that normally generated on silver electrode (see Note 6). Repeat polishing until all brown/black deposits on the electrode surface are removed.
- Gently polish the center of the electrode dome avoiding scratching.
- Rinse the electrode disc with distilled water to remove all traces of the polishing paste (see Note 7).
- Dry the electrode disc by using a paper towel.
- Store the electrode disc in an air-tight vessel.
- Clean the electrode disc after use.
Data analysis
- For each strain or condition, use at least 100 animals to obtain more accurate results.
- For each experiment, the same number of nematodes should be examined for each strain and condition.
- Each assay should be repeated at least three (3) times.
- Use the Student’s t-test with a significance cut-off level of P < 0.05 for comparisons between two groups.
- Use the one-factor (ANOVA) variance analysis and correct by the post hoc Bonferroni test for multiple comparisons.
Notes
- Do not over-tighten the large O-ring by screwing too strong because it could cause drifting signals during measurements.
- Oxygen electrode discs are very sensitive to temperature fluctuations affecting oxygen levels. Maintain experimental temperature stable throughout the duration of the experiment. Equilibrate the electrode disc to the assay temperature before taking place any calibration and/or measurement.
- Be careful not to destroy the membrane of the electrode disc during buffer removal.
- Perform three independent measurements per strain or condition.
- During sonication, samples should be kept cold to avoid protein degradation. Incubate samples constantly on ice during sonication.
- The electrode disc should never be left to dry out with KCl in place. Crystallization of KCl remnants could oxidized and destroy the electrode. Maintaining the disc to a high standard is extremely important.
- Do not wet the electrical connector of the disc during maintenance process.
Recipes
- Phosphate buffer (1 M)
- For 1 L, dissolve 102.2 g KH2PO4 and 57.06 g K2HPO4 in distilled water and fill up to 1 L. This is a 1 M solution, pH 6.0
- Autoclave and store at room temperature
- Nematode growth medium (NGM) agar plates
- Mix 3 g NaCl, 2.5 g BactoTM peptone, 0.2 g streptomycin, 17 g agar and add 900 ml distilled water. Autoclave
- Let cool to 55-60 °C
- Add 1 ml cholesterol stock solution, 1 ml 1 M CaCl2, 1 ml 1 M MgSO4 (sterile), 1 ml nystatin stock solution, 25 ml sterile 1 M phosphate buffer, pH 6.0, and distilled sterile water up to 1 L
- Pour about 8 ml medium per Petri dish and leave to solidify
- Keep the plates at 4 °C until used
- M9 buffer
- Dissolve 3 g KH2PO4, 6 g Na2HPO4, 5 g NaCl in 1 L distilled water. Autoclave
- Let cool and add 1 ml 1 M MgSO4 (sterile)
- Store M9 buffer at 4 °C
- Potassium chloride (KCl) buffer
- Dissolve 17.5 g KCl in 100 ml distilled water
- Store KCl buffer at 4 °C
Acknowledgments
This work was funded by grants from the European Research Council (ERC), the European Commission 7th Framework Programme and Bodossaki Foundation Postdoctoral Research Fellowship. The protocol has been adapted from Palikaras et al. (2015) Nature 521, 525-528.
References
- Artal-Sanz, M. and Tavernarakis, N. (2009). Prohibitin couples diapause signalling to mitochondrial metabolism during ageing in C. elegans. Nature 461(7265): 793-797.
- Chen, Z., Wang, Z., Guo, W., Zhang, Z., Zhao, F., Zhao, Y., Jia, D., Ding, J., Wang, H., Yao, M. and He, X. (2015). TRIM35 Interacts with pyruvate kinase isoform M2 to suppress the Warburg effect and tumorigenicity in hepatocellular carcinoma. Oncogene 34(30): 3946-3956.
- Dranka, B. P., Benavides, G. A., Diers, A. R., Giordano, S., Zelickson, B. R., Reily, C., Zou, L., Chatham, J. C., Hill, B. G., Zhang, J., Landar, A. and Darley-Usmar, V. M. (2011). Assessing bioenergetic function in response to oxidative stress by metabolic profiling. Free Radic Biol Med 51(9): 1621-1635.
- Li, Z., and Graham, B. H. (2012). Measurement of mitochondrial oxygen consumption using a Clark electrode. Methods Mol Biol 837: 63-72.
- Luz, A. L., Rooney, J. P., Kubik, L. L., Gonzalez, C. P., Song, D. H. and Meyer, J. N. (2015). Mitochondrial morphology and fundamental parameters of the mitochondrial respiratory chain are altered in Caenorhabditis elegans strains deficient in mitochondrial dynamics and homeostasis processes. PLoS One 10(6); e0130940.
- Palikaras, K., Lionaki, E. and Tavernarakis, N. (2015). Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521(7553): 525-528.
- Perry, C. G., Kane, D. A., Lanza, I. R. and Neufer, P. D. (2013). Methods for assessing mitochondrial function in diabetes. Diabetes 62(4): 1041-1053.
- Ryu, D., Mouchiroud, L., Andreux, P. A., Katsyuba, E., Moullan, N., Nicolet-Dit-Felix, A. A., Williams, E. G., Jha, P., Lo Sasso, G., Huzard, D., Aebischer, P., Sandi, C., Rinsch, C. and Auwerx, J. (2016). Urolithin A induces mitophagy and prolongs lifespan in C. elegans and increases muscle function in rodents. Nat Med 22(8): 879-888.
- Scheibye-Knudsen, M., Fang, E. F., Croteau, D. L., Wilson, D. M., 3rd, and Bohr, V. A. (2015). Protecting the mitochondrial powerhouse. Trends in cell biology 25(3): 158-170.
- Vafai, S. B. and Mootha, V. K. (2012). Mitochondrial disorders as windows into an ancient organelle. Nature 491(7424): 374-383.
- Vander Heiden, M. G., Cantley, L. C. and Thompson, C. B. (2009). Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324(5930): 1029-1033.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Palikaras, K. and Tavernarakis, N. (2016). Measuring Oxygen Consumption Rate in Caenorhabditis elegans. Bio-protocol 6(23): e2049. DOI: 10.21769/BioProtoc.2049.
Category
Developmental Biology > Cell growth and fate > Ageing
Biochemistry > Other compound > Oxygen
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












