- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cell-to-cell DNA Transfer among Thermus Species
Published: Vol 6, Iss 22, Nov 20, 2016 DOI: 10.21769/BioProtoc.2006 Views: 10191
Reviewed by: Daan C. SwartsMaria SinetovaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

A Fast and Easy Method to Study Ralstonia solanacearum Virulence upon Transient Gene Expression or Gene Silencing in Nicotiana benthamiana Leaves
Wenjia Yu and Alberto P. Macho
Aug 5, 2021 4722 Views
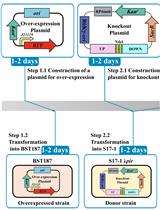
A New Tool for the Flexible Genetic Manipulation of Geobacillus kaustophilus
Ryotaro Amatsu [...] Ken-ichi Yoshida
Sep 5, 2022 1929 Views
Efficient Genetic Transformation and Suicide Plasmid-mediated Genome Editing System for Non-model Microorganism Erwinia persicina
Tingfeng Cheng [...] Lei Zhao
Mar 20, 2024 2472 Views
Abstract
The ability to transfer DNA via direct cell-to-cell contact-dependent process similar to conjugation has been described in Thermus thermophilus (Tth). Here, we detail the mating experiment protocol involving the lateral transfer of thermostable antibiotic resistance markers (i.e., kanamycin: KmR; hygromycin: HygR) between Thermus cells, enabling the selection and quantification of the transfer frequencies. Briefly, liquid cultures of both mates are mixed and laid onto a nitrocellulose filter on a TB plate. After incubation at 60 °C, filters are resuspended upon selective plating. The contribution of DNA uptake by transformation is abolished by the addition of DNase I to the mix. This protocol can be used for the transfer of large DNA fragments (> 10 kb) to Thermus species.
Background
Conjugation, as the chief mechanism for horizontal gene transfer for most bacteria, is a highly specialized process by which DNA is transferred between two cells which are in direct contact (Lederberg and Tatum, 1946). Classical conjugation involves the unidirectional transfer of a DNA molecule, generally plasmid-encoded, from a donor to a recipient cell, which remains passive. However, alternative models have been described for a wide variety of bacteria. In the laboratory, conjugation experiments are fruitful for marker-exchange mutagenesis, among others. Over other methods of genetic transfer, conjugation is advantageous in terms of minimal disruption of the bacterial envelope and the feasibility to transfer large DNA fragments, including large chromosomal regions (> 10 kb).
A conjugation-like process among T. thermophilus cells was reported a decade ago, where chromosomal markers were transferred following a high frequency of recombinants Hfr-like process, as evidenced by interrupted mating assays employing liquid mixes of different strains (Ramírez-Arcos et al., 1998). Recently, the model proposed for Thermus thermophilus describes the transfer as a two-step bidirectional process where a functional competence apparatus is required in the recipient cell but not in the donor, bestowing this phenomenon the name of ‘transjugation’ (Blesa et al., 2015). Compared to liquid mating assays aforementioned, transfer frequencies obtained with the protocol described here are higher and more robust. Higher reproducibility of the assays is reached with this protocol compared to mating tests in liquid. Besides, validation of the conjugative transfer is ensured by the addition of DNase.
Materials and Reagents
- Sterile plastic Petri dish plates (standard size; 100 x 15 mm)
- Sterile 1.5 ml microcentrifuge tubes (STARSTEDT, catalog number: 72.690.001 )
- Sterile tips for micropippettes
- Nitrocellulose filters, 0.22 μm (EMD Millipore, catalog number: GSWP02500 )
- Donor and recipient Thermus strains, containing selectable markers (i.e., Thermus sp geneA::kat and Thermus sp geneB::hyg)*
- Tryptone (Conda, catalog number: 1612 )
- Yeast extract (Conda, catalog number: 1702 )
- Sodium chloride (NaCl) (EMD Millipore, catalog number: 1.06404.1000 )
- Agar (grade A) (Conda, catalog number: 1800 )
- Thermus water (carbonate-rich mineral water)
- Appropriate thermostable selectable markers (i.e., kanamycin: KmR; Hygromycin: HygR)**
- Kanamycin sulphate (Sigma-Aldrich, catalog number: K1377-5G )
- Hygromycin B (Sigma-Aldrich, catalog number: BH7772-1G )
- DNase I, grade II from bovine pancreas (1 mg/ml) and its buffer solution (Roche Diagnostics, catalog number: 10104159001 )
- TB liquid medium (see Recipes)
- TB agar plates (see Recipes)
Notes:
- *Several Thermus thermophilus strains such as HB27, NAR1 and HB8, have been successfully employed in mating assays (Ramírez-Arcos et al., 1998; César et al., 2011; Blesa et al., 2015). All parental Thermus strains can be acquired at the DSMZ ollection.
- **Vectors containing thermostable selectable markers are available at RIKEN repository
Equipment
- 50 ml Erlenmeyer flask
- Rotary shaker incubator reaching 70 °C (150 rpm) (Thermo Scientific, Thermo ScientificTM, catalog number: SHKE 420HP )
- Glass plating beads or a spreader
- Autoclave device (CertoClav, model: EL 18L )
- Tweezers (EMD Millipore, catalog number: XX6200006P )
- Vortex Minishaker MS2 (IKA, model: MS2 )
- Spectrophotometer (Hitachi, model: U-2000 )
- Pipette P2, 0.2-2 μl (Gilson, Pipetman classicTM, catalog number: F144801 )
- Pipette P20, 2-20 μl (Gilson, Pipetman classicTM, catalog number: F123600 )
- Pipette P100, 20-100 μl (Gilson, Pipetman classicTM, catalog number: F123615 )
- Pipette P200, 50-200 μl (Gilson, Pipetman classicTM, catalog number: F123601 )
- 70 °C heater (Gemini, model: Memmert UE400 )
- Bench-top microcentrifuge with rotor for 1.5 ml microcentrifuge tubes (Eppendorf, model: Mini Spin® plus )
- Humid chambers
Notes: - Tupperware devices are preferentially used. Humidified plastic bags are also valid.
- Addition of a paper towel, soaked on Thermus water and laid on the bottom of the chamber would enhance humidity maintenance. Ventilation should be avoided in order to conserve humidity. An example is provided in the photographs beneath (Figure 1).
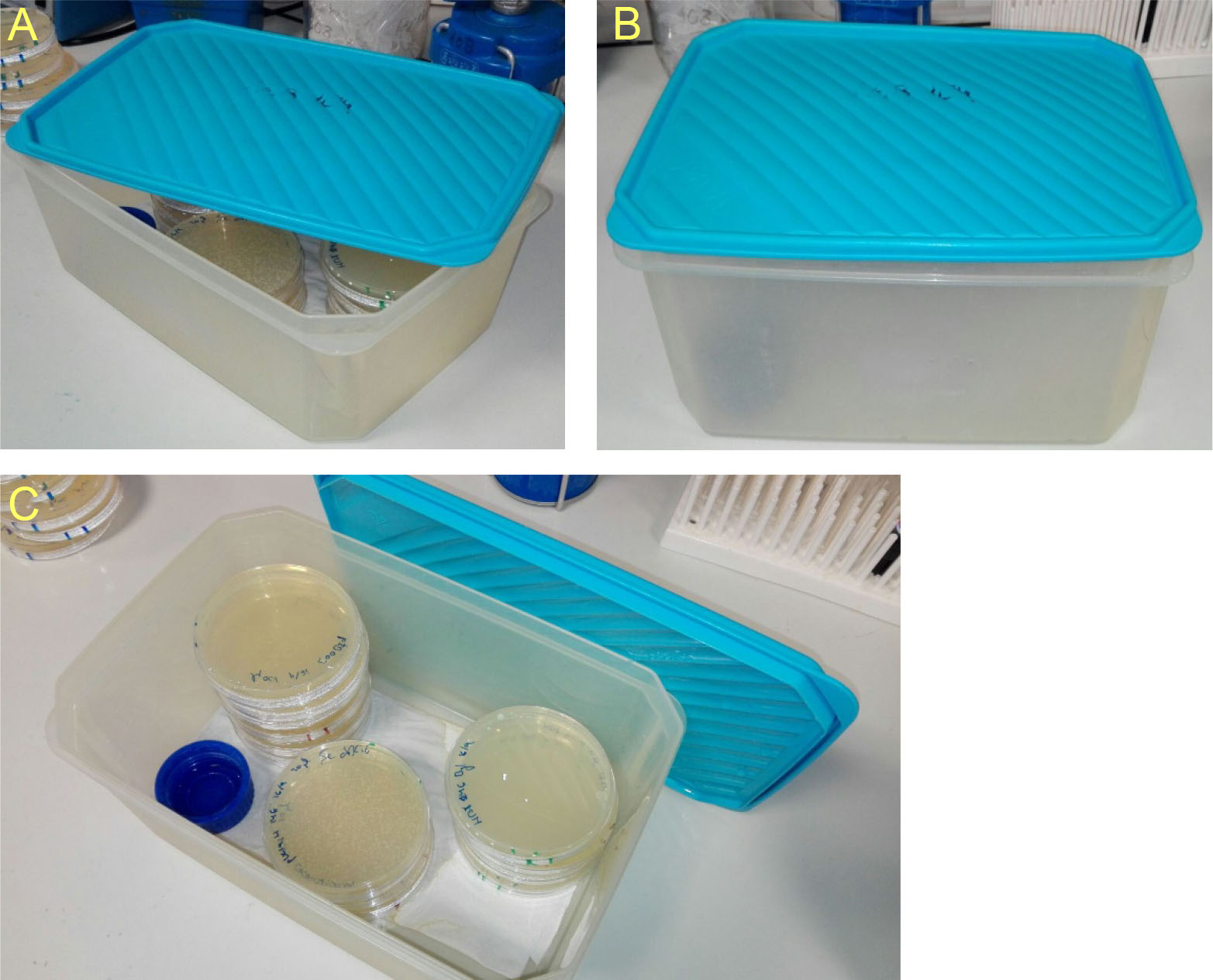

Figure 1. Humid chambers. Descriptive photographs of the humid chambers used to incubate Thermus plates. Place a piece of paper soaked with Thermus water and a bottle cap filled with the same water on the bottom of the chambers to maintain high humidity.
Procedure
A schematic summary is also provided underneath (Figure 2).
- Culture preparation
- Use 50 μl of each saturated T. thermophilus liquid growth culture to inoculate 10 ml TB medium containing the appropriate antibiotics (KmR [30 μg/ml] or HygR [100 μg/ml]) or inoculate the culture from the glycerol stock or from a frozen pellet.
- Incubate overnight the culture at 65 °C in a shaker incubator at 150 rpm.
Note: If cultures have not grown optimally overnight, it is highly recommended to delay the assay. - Mating mix
- Take 100 μl of each saturated overnight grown cells (OD600 ~1.5-2), place them on sterile microcentrifuge tubes and centrifuged them separately for 4 min at 1,500 x g at room temperature (~24 °C).
Note: In general, 1:1 cells ratio (both saturated cultures; OD600 ~1.5-2) is employed unless specific tests are accomplished or one of the mates has grown in lesser amount. - Discard the supernatant and resuspend each pellet in 6 μl of warm TB medium (pre-heated at 60 °C) and supplement each of them with 0.6 units of DNase I (~0.8 μl; Roche Diagnostics) and its buffer (~1.5 μl of buffer 10x).
Note: Room temperature TB medium can also be employed for resuspension. - Mix both mates and gently vortex for 5 sec.
Note: The purpose for vortexing is just to enhance the mixing. It should not be vigorous nor prolonged to avoid harnessing the cells. - Lay the cell mix on a sterile nitrocellulose filter which had been placed on a non-selective pre-warmed TB agar plate.
Note: A pre-warmed plate facilitates the attachment of the cells to the filter. - Mating transfer
- Place the TB agar plate with the cell-containing nitrocellulose filter on a heater at 60 °C for 4 h.
Note: The plate does not need to be incubated inside a humid chamber. - Mating interruption
- Take the agar plate out from the heater and pick the filter containing the mating strains with tweezers.
- Soak the filter in 1 ml of sterile TB medium in a microcentrifuge tube at room temperature.
Note: Pre-heated medium (~60 °C) is recommended as it fastens the detachment of the cells from the filters. - Vortex vigorously each tube to detach the cells from the filter.
Note: It is preferable to perform several vortex times (e.g., 4 times, max 30-40 sec) than one prolonged one (larger than 1.5 min) to minimize the disturbance upon the cells. - Plating
- Prepare serial dilutions of the cell mix in TB medium in sterile microcentrifuge tubes.
- Plate 100 μl onto selective TB agar plates using glass beads or a spreader, one with both antibiotics (KmR and HygR) and also plate 100 μl of the mix on its appropriate dilution and selective marker (KmR or HygR).
Note: Appropriate dilutions should be plated, for instance, 10-6 for each parental strain. Despite differences among mating efficiencies and thus, transconjugants arousal, in general, transconjugants should be plated 10-1 and 10-3 dilutions, which are common in standard matings. - Incubate the TB agar plates at 60 °C for 48 h inside humid chambers.
Note: Humidity preservation inside the humid chambers is essential for proper growth of colonies. - Transfer frequencies’ calculation
- Count the CFUs on each selective plates.
- Express the frequency of transfer as the CFUs grown in both markers (KmR and HygR) divided by the CFUs of the recipient mate (for instance, HygR).
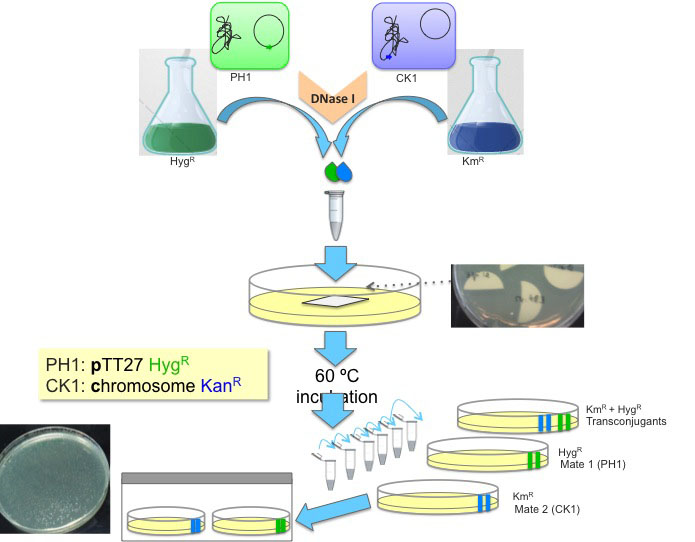
Figure 2. Schematic representation of the Thermus transjugation procedure. Thermus derivatives PH1, a hygromycin resistant strain (in green, H-termed) harbouring the hyg cassette at the megaplasmid (P-termed) is mated to CK1, a kanamycin resistant strain (in blue, K-termed) harbouring the kat cassette in the chromosome (C-termed). Saturated cultures of each of these strains are mixed in equal proportion (100 μl each) and incubated together in the presence of DNase I and then diluted for plating in different selective plates.
This transjugation protocol was optimized for Thermus thermophilus HB27, which shows the highest rates of cell-to-cell transfer. Less efficient strains should be incubated for longer periods or larger culture volumes employed in order to achieve reproducible data and maximize the transjugants yields.
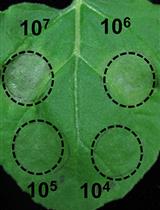
Qualitative assay of transjugation can be also assayed by laying 107 Thermus cells (for instance, 10 μl of saturated culture of CK1) on top of a pre-dried double selective agar plate (KmR and HygR) and topped with 107 cells of the counterpart strain (for instance, 10 μl of saturated culture of PH1). One unit of DNase I is added to the spot so as to prevent transformation (Figure 3). The plate is incubated in humid chambers, like aforementioned.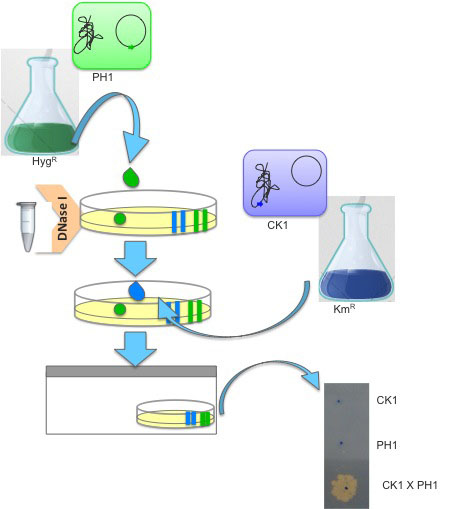
Figure 3. Schematic representation of the qualitative assay of Thermus transjugation procedure. A double selective Thermus agar plate (KmR and HygR) is amended with a 10 μl drop of saturated cultures of Thermus derivatives PH1, a hygromycin resistant strain (in green, H-termed) harbouring the hyg cassette at the megaplasmid (P-termed) which is then topped with an equal volume of a saturated culture of CK1, a kanamycin resistant strain (in blue, K-termed) harbouring the kat cassette in the chromosome (C-termed). Both cultures are incubated together in the presence of DNase I and incubated at 60 °C in humid chambers.
Data analysis
- Transfer frequencies are calculated as aforementioned, dividing the number of CFU transconjugants grown in double selective plates (KmR and HygR) by the CFUs grown in single selective plate (for instance, HygR). Data can be analysed through a set of statistical procedures, as described in Blesa et al. (2015). In general, to examine differences among frequencies of transfers, Student’s t-test and Mann-Whitney U-tests may be used. Transfer efficiencies among strains tagged alike can be compared using these tests too. Besides, Wilcoxon-tests may be used when comparing sets of transfer frequencies of different strains to assess whether the transfer frequency mean differs. When the purpose is to compare the ease for transfer a marker placed in different regions of the genome, one-way ANOVA assays will fit as differences among and within various loci transfer frequencies can be addressed using these tests. Simple linear regression test can be implemented for modelling the relationship between loci and frequency of transfer. Post-hoc Tukey and Bonferroni tests should be applied when convenient.
- At least three independent experiments should be performed for reliable and reproducible data.
Notes
- Transjugants should be re-streaked twice on selective plates and further grown in liquid under each particular selective condition so as to stabilize the integration of the markers.
- DNAse I is compulsory to defeat transformation transfer.
- Matings can also be performed on liquid TB in presence of DNase I. Standard experiments involving 100 μl of each mating pair (unless otherwise required) are centrifuged independently for 4 min at 1,500 x g and resuspended in 50 μl of TB medium mixed in 1.5-ml microcentrifuge tubes amended with 5 units of DNase I, centrifuged for 4 min at 1,500 x g and resuspended in 100 μl of TB medium, supplementing the mix with 5 units of DNase I and incubated for 5 h at 60 °C under shaking (180 rpm). After incubation, appropriate dilutions are plated on selective plates, as described above.
- For other markers or properties, appropriate selection should be applied. For example, in transjugation of the large denitrification island, the ordinary protocol may be adapted to anoxic conditions for selection. Further adaptations for the transfer of the denitrification island can be reviewed in Álvarez et al. (2014). Briefly, overnight cultures grown at 70 °C are harvested and washed together in liquid TB medium and supplemented with 5 units of DNase I. Then, 60 μl of the mix is laid onto 0.22 μm nitrocellulose filter and incubated at 60 °C for 24 h. Cells are then detached, resuspended in 1 ml of TB medium and added to 10 ml TB medium in anaerobiosis tubes, amended with potassium nitrate (40 mM) and the appropriate antibiotics for selection. Tubes are incubated for 48 h at 70 °C without agitation. Two re-inoculation steps under the same conditions are required prior to plating. Individual colonies should be streaked and tested for denitrifying respiration and presence of denitrifying genes should be confirmed by polymerase chain reaction (PCR).
- Erlenmeyer flasks do not require any special sealing. Standard lids are enough to prevent evaporation and preserve sterile conditions.
Recipes
- TB liquid medium
0.8 % tryptone (w/v)
0.4 % yeast extract (w/v)
50 mM NaCl, pH 7.5
Thermus water (carbonated-rich mineral water)*
pH may be adjusted by the addition of NaOH - TB agar plates
0.8% tryptone (w/v)
0.4% yeast extract (w/v)
50 mM NaCl, pH 7.5
1.8% agar (grade A) (w/v)
Thermus water (carbonated-rich mineral water)*
*Note: The analytical composition of the spring mineral water employed, per mg/ml, is listed beneath (Table 1). Other mineral waters easily purchasable such as Evian water or a laboratory solution resembling this recipe, can be employed.
Table 1. Mineral composition of the spring water from Jaraba Spring in Zaragoza (Spain)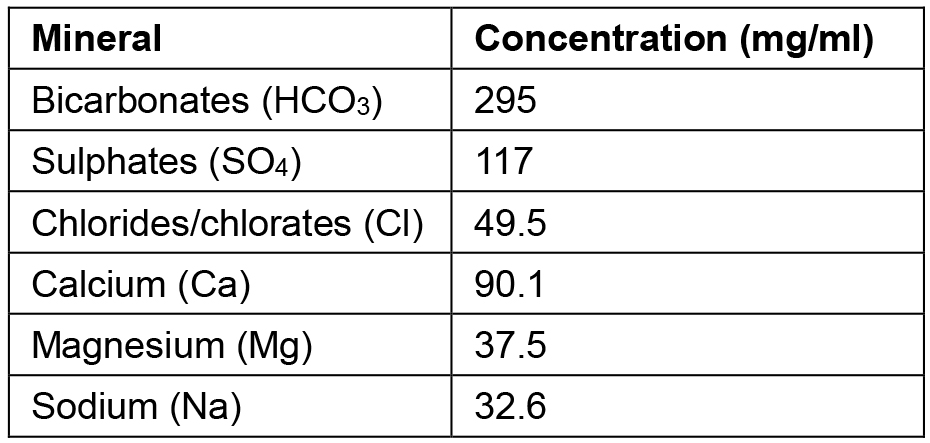
Acknowledgments
This work has been funded by Grants BIO2013-44963-R, and FP7-PEOPLE-2012-IAPP No. 324439. A. Blesa held an FPI contract from the Spanish Ministry of Economy and Competitivity.
References
- Alvarez, L., Bricio, C., Blesa, A., Hidalgo, A. and Berenguer, J. (2014). Transferable denitrification capability of Thermus thermophilus. Appl Environ Microbiol 80(1): 19-28.
- Blesa, A., Cesar, C. E., Averhoff, B. and Berenguer, J. (2015). Noncanonical cell-to-cell DNA transfer in Thermus spp. is insensitive to argonaute-mediated interference. J Bacteriol 197(1): 138-146.
- César, C. E., Alvarez, L., Bricio, C., van Heerden, E., Littauer, D. and Berenguer, J. (2011). Unconventional lateral gene transfer in extreme thermophilic bacteria. Int Microbiol 14(4): 187-199.
- Lederberg, J. and Tatum, E. L. (1946). Gene recombination in Escherichia coli. Nature 158(4016): 558.
- Ramirez-Arcos, S., Fernandez-Herrero, L. A., Marín, I. and Berenguer, J. (1998). Anaerobic growth, a property horizontally transferred by an Hfr-like mechanism among extreme thermophiles. 180(12): 3137-3143.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Blesa, A. and Berenguer, J. (2016). Cell-to-cell DNA Transfer among Thermus Species. Bio-protocol 6(22): e2006. DOI: 10.21769/BioProtoc.2006.
Category
Microbiology > Microbial genetics > Transformation
Molecular Biology > DNA > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










