- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Phagocytosis Assay to Measure Uptake of Necroptotic Cancer Cells by BMDCs
Published: Vol 6, Iss 21, Nov 5, 2016 DOI: 10.21769/BioProtoc.1997 Views: 13382
Reviewed by: Xi FengJalaj GuptaRuth A. Franklin

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
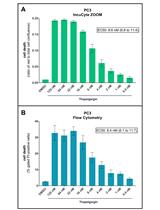
An Image-based Assay for High-throughput Analysis of Cell Proliferation and Cell Death of Adherent Cells
Paula Szalai and Nikolai Engedal
May 5, 2018 11046 Views

Evaluation of Anticancer activity of Silver Nanoparticles on the A549 Human Lung Carcinoma Cell Lines through Alamar Blue Assay
Nikita Sharma [...] Surendra Nimesh
Jan 5, 2019 14751 Views
Real-time IncuCyte® Assay for the Dynamic Assessment of Live and Dead Cells in 2D Cultures
Arlene K. Gidda [...] Sharon M. Gorski
Feb 5, 2025 2949 Views
Abstract
This protocol is a flow cytometry-based method to measure the phagocytosis efficiency of necroptotic target cells by bone marrow-derived dendritic cells (BMDCs) in vitro (Aaes et al., 2016). The method is a slightly modified and updated version of the protocols used in previously published papers (Krysko et al., 2006; Brouckaert et al., 2004). In brief, the target cells are labeled with a CellTrackerTM dye before they are induced to undergo cell death. After a co-culture period of 2 h with BMDCs, the cells are immunostained with a dendritic cell marker and dead cell marker, and the phagocytic efficiency is quantified using a flow cytometer. This protocol can readily be used for target cells undergoing cell death modalities other than necroptosis.
Background
Studying the phagocytic uptake of necroptotic cells by BMDCs in vitro, is a preliminary step in the examination of immunogenic cell death models (Obeid et al., 2007). Efficient uptake will allow the phagocyte to cross-present antigens to leukocytes and thereby create an immune reaction towards the dead target cells. In this protocol we make use of a CellTrackerTM dye. This type of dye may be toxic in certain concentrations, which may vary depending on which cell type is used. Thus, we recommend users to first find the optimal concentration of the dye for the target cell in use. Optimally, the CellTrackerTM dye itself should not induce any cell death, but should label the target cells so that they become easily separable from the CD11c-positive BMDCs.
Materials and Reagents
- Delta treated 10 cm Petri dishes (Thermo Fisher Scientific, NuncTM, catalog number: 153066 )
- 15 ml polystyrene centrifuge tubes (Corning, Falcon®, catalog number: 352095 )
- 6-well suspension plates (SARSTEDT, catalog number: 83.3920.500 )
- 96 V well, 2.0 ml polypropylene plate (Greiner Bio One, MASTERBLOCK®, catalog number: 780285 )
- 5 ml round-bottom polystyrene tubes (Corning, Falcon®, catalog number: 352054 )
- Bone marrow-derived dendritic cells (BMDCs) isolated from BALB/c WT mice (see Procedure)
- CT26 Necroptosis-inducible cells (DD_RIPK3) (Aaes et al., 2016)
- Dulbecco’s modified Eagle medium (DMEM), high glucose (Thermo Fisher Scientific, GibcoTM, catalog number: 41965062 )
- Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10270106 )
- Sodium pyruvate (Sigma-Aldrich, catalog number: S8636 )
- L-glutamine (Lonza, catalog number: BE17-605F )
- Roswell park memorial institute (RPMI) 1640 medium (Thermo Fisher Scientific, GibcoTM, catalog number: 52400025 )
- 2-mercaptoethanol (50 mM) (Thermo Fisher Scientific, GibcoTM, catalog number: 31350010 )
- Murine GM-CSF (VIB Protein Service Facility, UGent-VIB Inflammation Research Center)
- ACK lysing buffer (Lonza, catalog number: 10-548E )
- Dulbecco’s phosphate-buffered saline (DPBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 14190094 )
- CellTrackerTM green CMFDA dye (Thermo Fisher Scientific, Molecular ProbesTM, catalog number: C7025 )
- Doxycycline (Sigma-Aldrich, catalog number: D9891 )
- B/B homodimerizer (Takara Bio, catalog number: 635059 )
- Purified rat anti-mouse CD16/CD32 (Mouse BD Fc BlockTM) (BD, PharmingenTM, catalog number: 553142 )
- APC hamster anti-mouse CD11c (BD, PharmingenTM, catalog number: 550261 )
- SYTOX® blue nucleic acid stain (Thermo Fisher Scientific, Molecular ProbesTM, catalog number: S11348 )
- BD FACSuite CS&T Research Beads Kit (BD, FACSuiteTM, catalog number: 650621 )
- DMEM CT26 culture medium (see Recipes)
- RPMI BMDC/Co-culture medium (see Recipes)
- FACS buffer (see Recipes)
- Staining solution (see Recipes)
Equipment
- Table top centrifuge
- BD FACSVerseTM flow cytometer (BD, model: BD FACSVERSE )
Software
- BD FACSuiteTM software (BD)
- FlowJo software version 10.0.8 or newer (FlowJo)
Procedure
Bone marrow cells were isolated by crushing the femurs and tibias of 7-week-old BALB/c WT mice. Red blood cells were lysed with ACK lysing buffer, and the cells were differentiated into dendritic cells for eight days using RPMI culture medium – 400,000 cells per 2 ml per 6-well. On day three, 2 ml of fresh culture medium was added, and on day six the medium was replaced with fresh culture medium.
- Cell death induction in target, CT26 DD_RIPK3, cells
- Seed 1 x 106 CT26 DD_RIPK3 cells in 10 ml DMEM culture medium per 10 cm Petri dish.
- Wait 5 h for the cells to attach.
- Wash the Petri dishes with serum-free DMEM.
- Add 10 ml of serum-free DMEM + 1 µM CellTrackerTM green CMFDA to each plate.
- Incubate for 30 min at 37 °C.
- Remove the medium, and wash the plate with RPMI culture medium.
- Add 10 ml RPMI culture medium to each Petri dish, incubate for 2 h at 37 °C.
- Induce cell death by adding 1 µg/ml doxycycline + 10 nM B/B homodimerizer.
- Incubate for another 18 h at 37 °C.
- Seed 1 x 106 CT26 DD_RIPK3 cells in 10 ml DMEM culture medium per 10 cm Petri dish.
- Co-culture with BMDCs
- Collect the BMDCs and CT26 target cells into separate 15 ml Falcon® tubes
- Live CT26 cells are collected by adding EDTA (2 ml per 75 cm2), incubating at 37 °C for 2 min, then collecting in RPMI culture medium (8 ml per 75 cm2). Centrifuge at 300 x g for 5 min.
- Dead CT26 cells are collected by pipetting only the cells in solution. Centrifuge at 400 x g for 5 min.
- Count the cells and resuspend them in RPMI culture medium.
- Seed the cells in 6-well suspension plates.
- Seed 400,000 BMDCs + 400,000 CT26 cells (ratio 1:1) in a total volume of 4 ml per well.
- Incubate for 2 h at 37 °C.
- Include wells with cells for single stain and compensation controls:
- CT26 cells Non-stained target cell control
- CT26 cells – cell death-induced SYTOX® blue single stain control
- CT26 cells CellTrackerTM green single stain control
- BMDCs Non-stained BMDC control
- BMDCs CD11c-APC single stain control
- Immunostaining
- Transfer the cells to a deep 96-well plate.
- Spin down at 400 x g for 5 min at 4 °C.
- Discard the supernatant by flicking the plate, and resuspend the pellet in 200 µl FACS buffer.
- Spin down at 400 x g for 5 min at 4 °C.
- Resuspend the samples in 200 µl of staining solution (or in single stain solution) with a final antibody concentration (APC-CD11c) of 0.1 µg per 8 x 105 cells.
- Incubate for 30 min at 4 °C covered from light.
- Wash the samples and spin down at 400 x g for 5 min at 4 °C.
- Resuspend in FACS buffer + 1.25 µM SYTOX® blue (or in FACS buffer only for the single stained samples).
- Transfer to 5 ml Falcon® round-bottom tubes.
- Keep the samples on ice (for maximum 1 h) until acquisition on the flow cytometer.
- On the flow cytometer use FACSuiteTM to acquire minimum 50-100,000 events per sample.
- Transfer the cells to a deep 96-well plate.
- Gating strategy in FlowJo
True uptake of CMFDA-labeled dead cell material by BMDCs is determined using a gating strategy that allows analysis of only single cells and is determined as CD11c+ CMFDA+ double-positive cells expressed as a percentage of all CD11c+ cells. See Figure 1 and Table 1 for the gating strategy and compensation matrix used in FlowJo.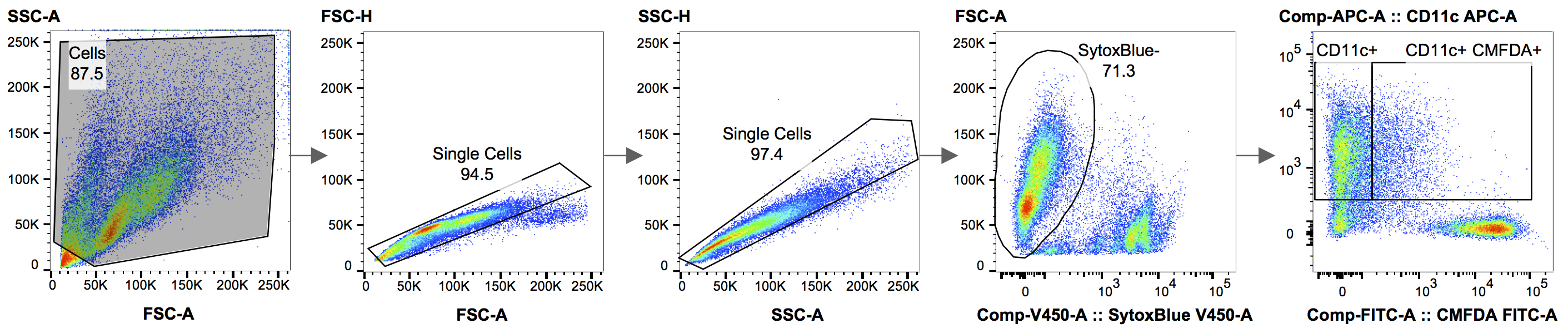
Figure 1. Gating strategy used for the analysis in FlowJo. Figure 1 shows the gating strategy used, when analyzing the results in FlowJo. Using the forward and side scatter area (FSC-A vs. SSC-A), the debris is excluded from the analysis. In the following two steps, doublet cells are removed, by drawing a gate around the single cells, which roughly gather along the diagonal of FSC-A vs. FSC-H, followed by SSC-A vs. SSC-H. A Live/Dead staining, in this case SYTOX® blue, is used to eliminate dead cells. In the final panel, the live, SYTOX® blue-negative, cells can now be analyzed for phagocytosis by plotting Cell Tracker Green CMFDA (target cells) vs. CD11c-APC (BMDCs).
Table 1. Compensation matrix generated in FlowJo for the analysis of the phagocytic BMDCs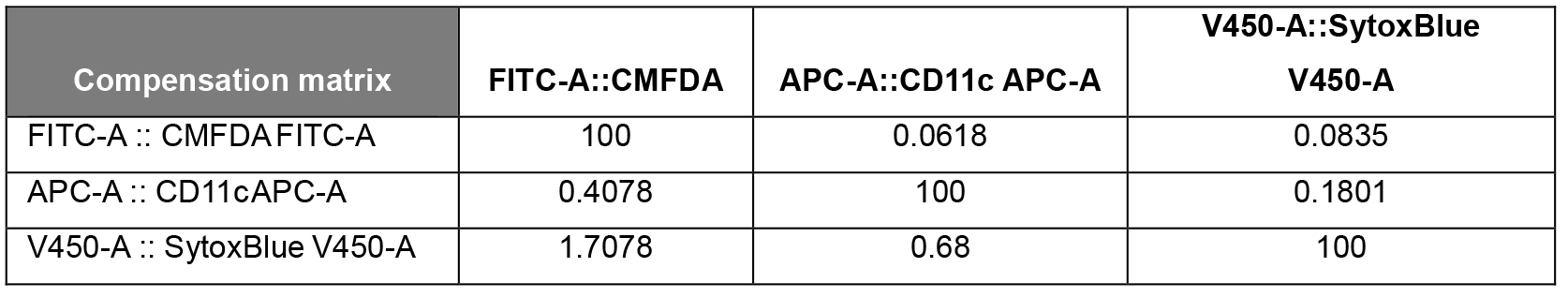
Data analysis
- Software
- BD FACSuiteTM software (BD)
- FlowJo software version 10.0.8 or newer (FlowJo)
- The experiment should be repeated a minimum of three times, each time including technical triplicates of each sample. The statistical difference between BMDC uptake of live cells versus that of necroptotic cells can be analyzed with a Mann-Whitney non-parametric t-test.
- The phagocytic BMDCs (see gating strategy in the legend of Figure 1) are calculated as the percentage of double-positive CMFDA+ CD11c+ cells (BMDCs that have engulfed target cells) out of all CD11c+ cells (the total BMDC population).
Notes
To verify the efficiency of the cell death induction in the target cells, it is advised to run a cell death analysis on the flow cytometer in parallel with the phagocytosis analysis. Cell death analysis should include the nucleic acid staining combined with an Annexin V fluorescent probe (Aaes et al., 2016). Tet-On induction for 18 h with doxycycline results in > 85% cell death.
Recipes
- DMEM CT26 culture medium
Note: Store at 4 °C; heat to 37 °C before use.
DMEM
10% FBS
1.3% Na-pyruvate
1.4 mM L-glutamine - RPMI BMDC/Co-culture medium
Note: Store at 4 °C; heat to 37 °C before use.
RPMI
5% FBS
1.3% Na-pyruvate
1.4 mM L-glutamine
50 µM 2-Me
20 ng/ml mGM-CSF - FACS buffer
Note: Store at 4 °C.
DPBS
0.5% FBS - Staining solution
Note: Prepare just before use, keep on ice.
FACS buffer
Fc BlockTM diluted 1/250
APC Hamster anti-mouse CD11c diluted 1/350
Acknowledgments
Peter Vandenabeele’s research group is part of the Cancer Research Institute Ghent (CRIG). Funding is supported by Interuniversity Attraction Poles (IAP 7/32), Research Foundation Flanders (FWO), Methusalem grants, Ghent University grants and the Belgian Foundation Against Cancer.
The authors declare no conflicts of interest.
References
- Aaes, T. L., Kaczmarek, A., Delvaeye, T., De Craene, B., De Koker, S., Heyndrickx, L., Delrue, I., Taminau, J., Wiernicki, B., De Groote, P., Garg, A. D., Leybaert, L., Grooten, J., Bertrand, M. J., Agostinis, P., Berx, G., Declercq, W., Vandenabeele, P. and Krysko, D. V. (2016). Vaccination with necroptotic cancer cells induces efficient anti-tumor immunity. Cell Rep 15(2): 274-287.
- Brouckaert, G., Kalai, M., Krysko, D. V., Saelens, X., Vercammen, D., Ndlovu, M. N., Haegeman, G., D'Herde, K. and Vandenabeele, P. (2004). Phagocytosis of necrotic cells by macrophages is phosphatidylserine dependent and does not induce inflammatory cytokine production. Mol Biol Cell 15(3): 1089-1100.
- Krysko, D. V., Denecker, G., Festjens, N., Gabriels, S., Parthoens, E., D'Herde, K. and Vandenabeele, P. (2006). Macrophages use different internalization mechanisms to clear apoptotic and necrotic cells. Cell Death Differ 13(12): 2011-2022.
- Obeid, M., Tesniere, A., Ghiringhelli, F., Fimia, G. M., Apetoh, L., Perfettini, J. L., Castedo, M., Mignot, G., Panaretakis, T., Casares, N., Metivier, D., Larochette, N., van Endert, P., Ciccosanti, F., Piacentini, M., Zitvogel, L. and Kroemer, G. (2007). Calreticulin exposure dictates the immunogenicity of cancer cell death. Nat Med 13(1): 54-61.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Aaes, T. L., Krysko, D. V. and Vandenabeele, P. (2016). Phagocytosis Assay to Measure Uptake of Necroptotic Cancer Cells by BMDCs. Bio-protocol 6(21): e1997. DOI: 10.21769/BioProtoc.1997.
Category
Cancer Biology > Cell death > Cell biology assays > Cell viability
Immunology > Immune cell function > Dendritic cell
Cell Biology > Cell viability > Cell death
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










