- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
The Chick Embryo Chorioallantoic Membrane as an in vivo Model to Study Metastasis
Published: Vol 6, Iss 20, Oct 20, 2016 DOI: 10.21769/BioProtoc.1962 Views: 22756
Reviewed by: HongLok LungAnita UmeshAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
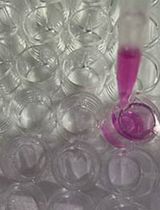
An Experimental Protocol for the Boyden Chamber Invasion Assay With Absorbance Readout
Kathleen C. Brown [...] Piyali Dasgupta
Aug 5, 2024 2760 Views
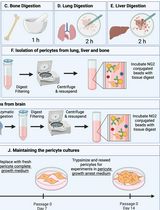
Isolation and Culture of Primary Pericytes from Mouse
Tamara McErlain [...] Meera Murgai
Apr 20, 2025 2985 Views
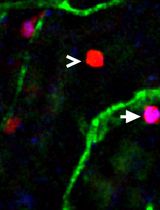
Utilizing EdU to Track Leukocyte Recruitment to the Brain
Zoie K. Lipfert [...] David P. Sullivan
Dec 5, 2025 1559 Views
Abstract
Metastasis is a complex process that includes several steps: neoplastic progression, angiogenesis, cell migration and invasion, intravasation into nearby blood vessels, survival in the circulatory system, extravasation followed by homing into distant tissues, the formation of micrometastases, and finally the growth into macroscopic secondary tumors. This complexity makes metastases difficult to investigate and quantify in animal models. The chick embryo is a unique in vivo model that overcomes many limitations for studying the metastatic process, due to the accessibility of the chorioallantoic membrane (CAM), a well-vascularized extra-embryonic tissue located under the eggshell, that is receptive to the xenografting of mammalian tumor cells, including human. Since the chick embryo is naturally immunodeficient at this stage, the CAM can support the engraftment of tumor cells, and their growth therein can faithfully recapitulate most of the characteristics of the carcinogenic process including: growth, invasion, angiogenesis and colonization of distant tissues (Deryugina and Quigley, 2008; Zijlstra et al., 2002). The CAM sustains rapid tumor formation within 5-7 days after cancer cell grafting. This feature provides a unique experimental model for a rapid study of the intravasation and colonization steps of the metastatic cascade. Furthermore, using quantitative PCR to detect species-specific sequences, such as Alu, the chick embryo CAM model can be used to monitor and quantify the presence of the xenografted, ectopic tumor cells in distant tissues. Thus, the chick embryo model has proved a valuable tool for cancer research, in particular for the investigation of molecules and pathways involved in cancer metastasis and to analyze the response of metastatic cancer to potential therapies (Herrero et al., 2015; Casar et al., 2014). In this respect, the use of the rapid and quantitative spontaneous metastasis chick embryo model can provide an alternative approach to conventional mouse model systems for screening anti-cancer agents.
Keywords: Metastasis modelMaterials and Reagents
- 20 G needles (BD, PrecisionGlideTM, catalog number: 305175 )
- 30 G needles (BD, PrecisionGlideTM, catalog number: 305128 )
- MicroAmp® optical 96 well PCR plate (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: N8010560 )
- MicroAmp® optical adhesive film (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: 4311971 )
- Cotton tipped applicators, cotton swab, Iodine liquid (Thermo Fisher Scientific)
- Laboratory tape 1/2" x 500" (VWR, catalog number: 470144-262 )
- Fertilized chicken eggs (Gilbert farm, Tarragona, Spain)
- A375 melanoma cell line (ATCC, catalog number: CRL-1619 )
- SKMEL2 (ATCC, catalog number: HTB68 )
- RKO colorectal cancer cell line (ATCC, catalog number: CRL-2577 )
- HCT116 (ATCC, catalog number: CCL-247 )
- DEL 22379 (Vichem Chemie, Budapest)
- Trypsin 0.05% with EDTA (1 mM), liquid (Thermo Fisher Scientific, GibcoTM, catalog number: 25300-054 )
- Phosphate-buffered saline (PBS) (1x, pH 7.4), liquid (Thermo Fisher Scientific, GibcoTM, catalog number: 10010023 )
- Penicillin-streptomycin (10,000 U/ml) (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Dulbecco’s modified Eagle medium (DMEM) (Thermo Fisher Scientific, catalog number: 41965062 )
- Fetal bovine serum (Thermo Fisher Scientific, GibcoTM, catalog number: 10270-106 )
- QIA amp genomic DNA purification kit (QIAGEN, catalog number: 158906 ; 158910 ; 158914 )
- Primers (HPLC purification, IDT DNA technologies)
Alu (human) sense: 5’ ACGCCTGTAATCCCAGGACTT 3’
Alu (human) antisense: 5’ TCGCCCAGGCTGGCTGGGTGCA 3’
Chicken GAPDH sense: 5’ GAGGAAAGGTCGCCTGGTGGATCG 3’
Chicken GAPDH antisense: 5’ GGTGAGGACAAGCAGTGAGGA ACG 3’ - SYBR® green mix real time PCR (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: 4472908 )
Equipment
- Incubator 37 °C, 60% humidity (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 51028117 )
- Rotating eggs trays (automatic eggs turner) (GQF, catalog number: 1611 )
- Tugon tube (Drifton, model: Tygon LMT )
- Egg candler (Lyon, model: 950-170 )
- Microsurgical kits, sterile forceps, push pin, dissection scissors, needle nose forceps (VWR, IntegraTM Miltex®, catalog number: 95042-542 )
- Dremel 100 rotary tool (Dremel, model: 100N/7 )
- Dremel cut off wheels number 36 (Dremel)
- Hemocytometer, Neubauer chamber (EMD Millipore)
- 2-20 µl pipette (Eppendorf, Eppendorf Research®, catalog number: 3120000038 )
- 20-200 µl pipette (Eppendorf, Eppendorf Research®, catalog number: 3120000054 )
- Automatic pipette (Eppendorf, Eppendorf Easypet® 3, catalog number: 4430000018 )
- Real time PCR instrument (Thermo Fisher Scientific, Applied BiosystemsTM, model: StepOne plus RT PCR )
Software
- Graph Pad Prism software
Procedure
A schematized summary of the spontaneous metastasis assay is illustrated in Figure 1.
Figure 1. The chick embryo for spontaneous metastasis model. After 10 days of incubation the CAM is dropping and tumor cells (green) are applied to it. After allowing xenografted tumor cells to grow, the tumor and chicken tissues are harvested on day 5-7 for Alu PCR.
- Preparing the eggs for xenografting tumor cells
- Freshly fertilized chicken eggs are incubated on their side in a rotating incubator at 37 °C and 60% humidity for 10 days. The eggs are rotated three times per hour (Figures 2.1 and 2.2)
- On day 10 the eggs are placed on their side on an egg rack (Figure 2.3). Use a tube lamp or other suitable light source to candle the eggs by shining the light at the blunt end of the egg where the air sack is located. The embryo must be located near the bottom of the egg and the air sack on its right. Localize and mark using a pencil the allantoic vein that is located at the top of the eggshell, right where several blood vessels cross (Figure 2.4).
- Clean the area, including and around the mark, using a cotton swab soaked with iodine (Figure 2.5).
- Drill a small hole through the eggshell into the air sack using a 30-gauge syringe needle (Figures 2.6 and 3A).
- Make another hole near the allantoic vein, which penetrates the shell membrane but not the CAM, using a Dremel rotary tool kit (Figures 2.7 and 3B). The CAM is attached to the inner surface of the shell, so care should be exercised at this point. Use a 20-gauge syringe needle with a small hook on the end to make a third very small hole in the eggshell membrane (Figure 3C, Note 1).
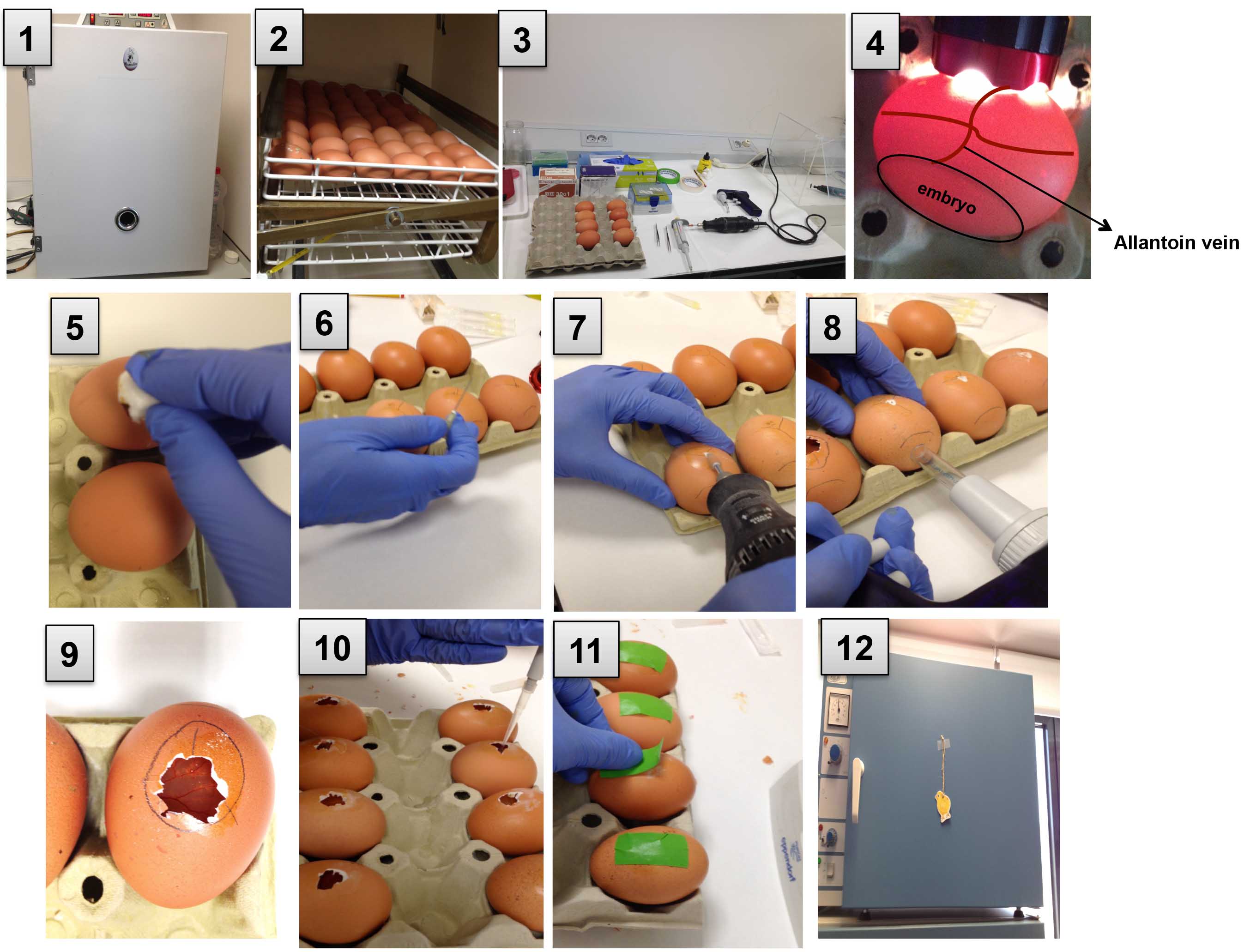

Figure 2. A serial depiction of the model. 1 and 2. Incubate the eggs; 3. Prepare a clean area and set all the materials; 4. Mark the location of the allantoic vein; 5. Apply iodine using a cotton swab; 6 and 7. Drill holes in the eggshell; 8. Drop the CAM; 9. Cut an opening for tumor cell grafting; 10. Apply the tumor cells and wait for 5 min; 11. Seal the hole using laboratory tape; 12. Incubate the eggs. - To suction, create with an automatic pipette aid fitted with a piece of Tygon tubing placed against the hole in the airsac. To separate the CAM from the shell and let it drop, apply a mild vacuum to the hole over the air sack so the blood vessel drops down, away from the eggshell and attaches to the embryo (Figures 2.8 and 3D).
- Use a cut off wheel (Dremel) to cut a square window (1 cm2) approximately 0.5 cm away from the branch point in the vein to expose the underlying CAM. A pair of needle-nose forceps can be used to remove a small section of eggshell in the square window and expose the CAM (Figures 2.9 and 3E).
- Once the CAM is dropped, seal the hole located near the chorioallantoic vein using laboratory tape.
- Then place the eggs in an egg rack into a stationary incubator at 37 °C and 60% humidity in anticipation for grafting the tumor cells.

Figure 3. A six panel diagram illustrating the appearance of the chick embryo metastasis model in cross section at key steps. A. After 10 days of incubation, the allantoic vein is positioned against the top of the egg. Drill one hole in the eggshell into the air sack using a 30 G syringe needle. B. Make another hole using a Dremel rotary tool kit adjacent to the attachment point for the allantoic vein. C. Make a third small hole in the eggshell membrane using a 20 G syringe needle. D. Apply a mild vacuum into the hole over the air sack to evacuate the air and drop the CAM. E. Cut a square window (1 cm2) using a Dremel cut off wheel and remove a small section of eggshell to expose the CAM. F. Graft the tumor cells onto the CAM. - Preparing tumor cells for grafting
- Detach tumor cells from their culture dishes using trypsin/EDTA and wash with phosphate buffered saline (PBS) twice in order to remove any residual media.
- Cells are counted using a Neubauer chamber and resuspended in serum-free DMEM at 40 million cells/ml depending on the cell line used (Note 2).
- Grafting the tumor cells onto the CAM
- Using a 20-200 μl pipette place 25 μl of the cell suspension onto a small area of the CAM (Figures 2.10 and 3F). The optimal number of cells should be determined empirically, but can range from 0.4 x 105 to 2 x 106 depending on the growth and ‘invasiveness’ characteristics of the tumor cell line used. Different tumors and cell suspensions derived from tumors have been implanted on the CAM (Tables 1 and 2). Tumor cell lines grow and intravasate to the vasculature with different efficiencies. For instance, suspensions of A375 melanoma (1 x 106), SKMEL2 melanoma (1 x 106), RKO colorectal carcinoma (2 x 106), HCT116 (0.5 x 106), U87 glioblastoma (0.2 x 106), PC3 prostate carcinoma (1 x 106), Hep3 epidermoid carcinoma (0.4 x 106), HT-1080 fibrosarcom (0.5 x 106) cell lines; have been grafted on the CAM of embryonated chicken eggs. All cell lines tested formed 4-8 mm size tumors, which recapitulated hallmarks of corresponding human specimens. Depending on the tissue origin, number of tumor cells and their proliferation capacity, primary CAM tumors can reach up to 500-600 mg in 6-7 days after cell inoculation.
- Seal the window in the egg tightly with laboratory tape and leave the eggs in a position with the embryos standing upright for 5-10 min, in order to allow the cells to settle (Figure 2.11).
- After 10 min, return the eggs to the stationary incubator (Figure 2.12).
- Allow cells to grow for 5-7 days depending on the nature of the tumor cell line used. This time should be enough for a macroscopic tumor to be visible.
- The CAM model allow experimental studies of potential anti-tumorigenic and anti-metastatic compounds. Tumors can be treated topically on the upper CAM with potential anti-cancer drugs. For instance, to evaluate the effects of DEL22379 in A375 and RKO tumor growth we inoculated tumor cells on the CAM and incubate the embryos for 4 days at 37 °C and 60% humidity. Then, we added 10 µM DEL22379 (prepare freshly, soluble in DMSO) topically on the CAM and treatment was refreshed every 12 h. Two days later tumors were harvested.
Table 1. Different tumor samples implanted onto the CAM
Table 2 .Tumor cells tested onto the CAM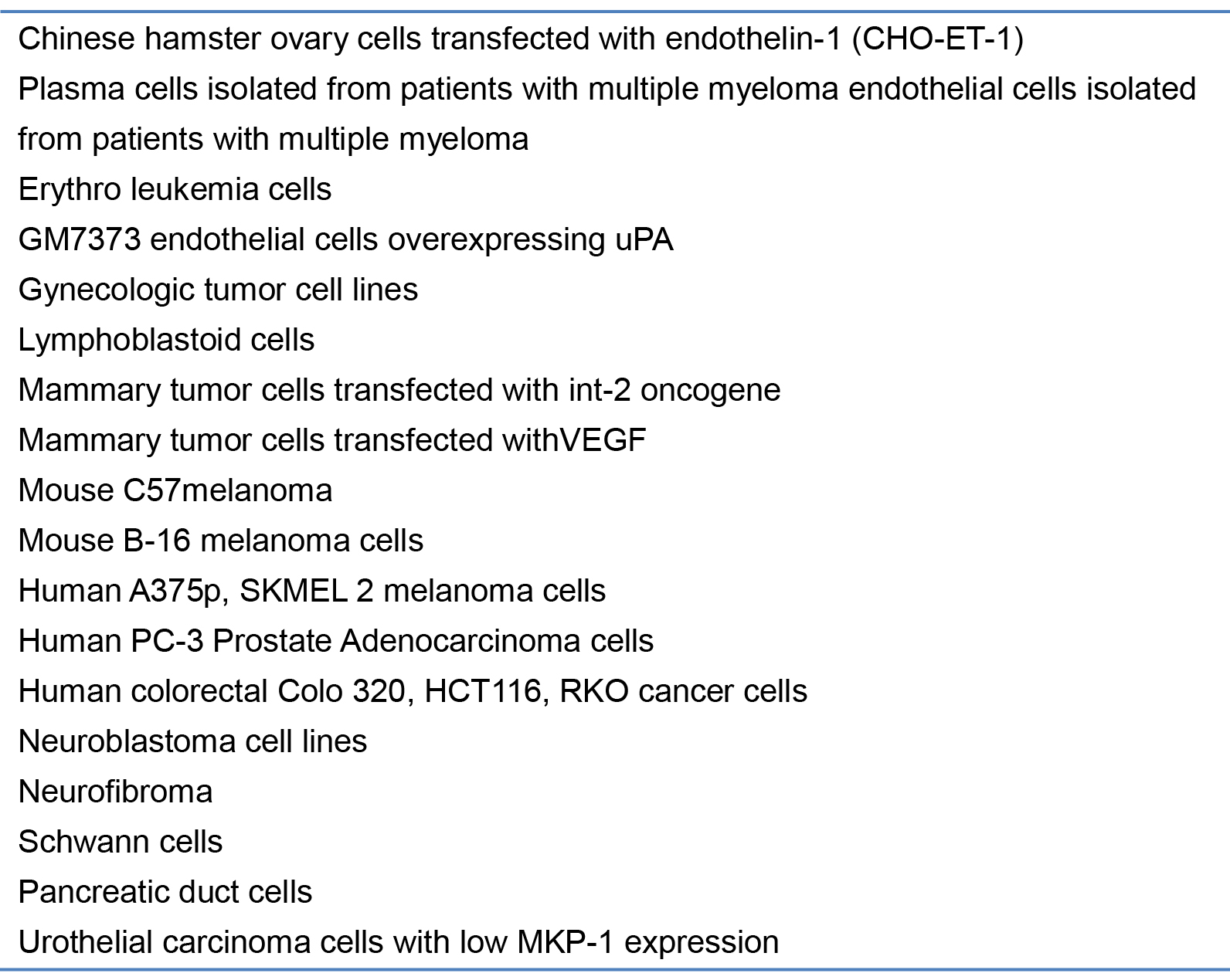
We describe the different steps for graft the tumor cells onto the CAM using the spontaneous metastasis model in Video 1.Video 1. The chick chorioallantoic membrane as an in vivo xenograft model to study tumor growth and metastasis - Harvesting tumors and chick embryo tissues
- Perform dissections in an area free of contaminating DNA (Figure 4.1). To detect metastatic invasion this assay quantifies the presence of ectopic DNA in chicken organs using a highly sensitive PCR approach. Therefore, it is very important to prevent contamination with exogenous DNA.
- In order not to have cross-contamination of your samples, use three separated sets of surgery tools. One set for cutting the egg, one set for harvesting the primary tumor and one set for removing the internal tissues.
- Prepare three wash containers for sequential rinsing of the tools between each animal and change their liquid between different experimental groups. Sequentially these washes are: distilled water, 70% ethanol and 1x PBS.
- Get the eggs from the stationary incubator.
- Open a new window removing some of the eggshell such that the tumors become visible. At this point, you can take some pictures of the appearance of the macroscopic tumor (Figures 4-6).
- Resect the primary tumor from the CAM and weigh the tumor.
- Remove the chick embryo from the eggshell by cutting the shell radially into equal halves.
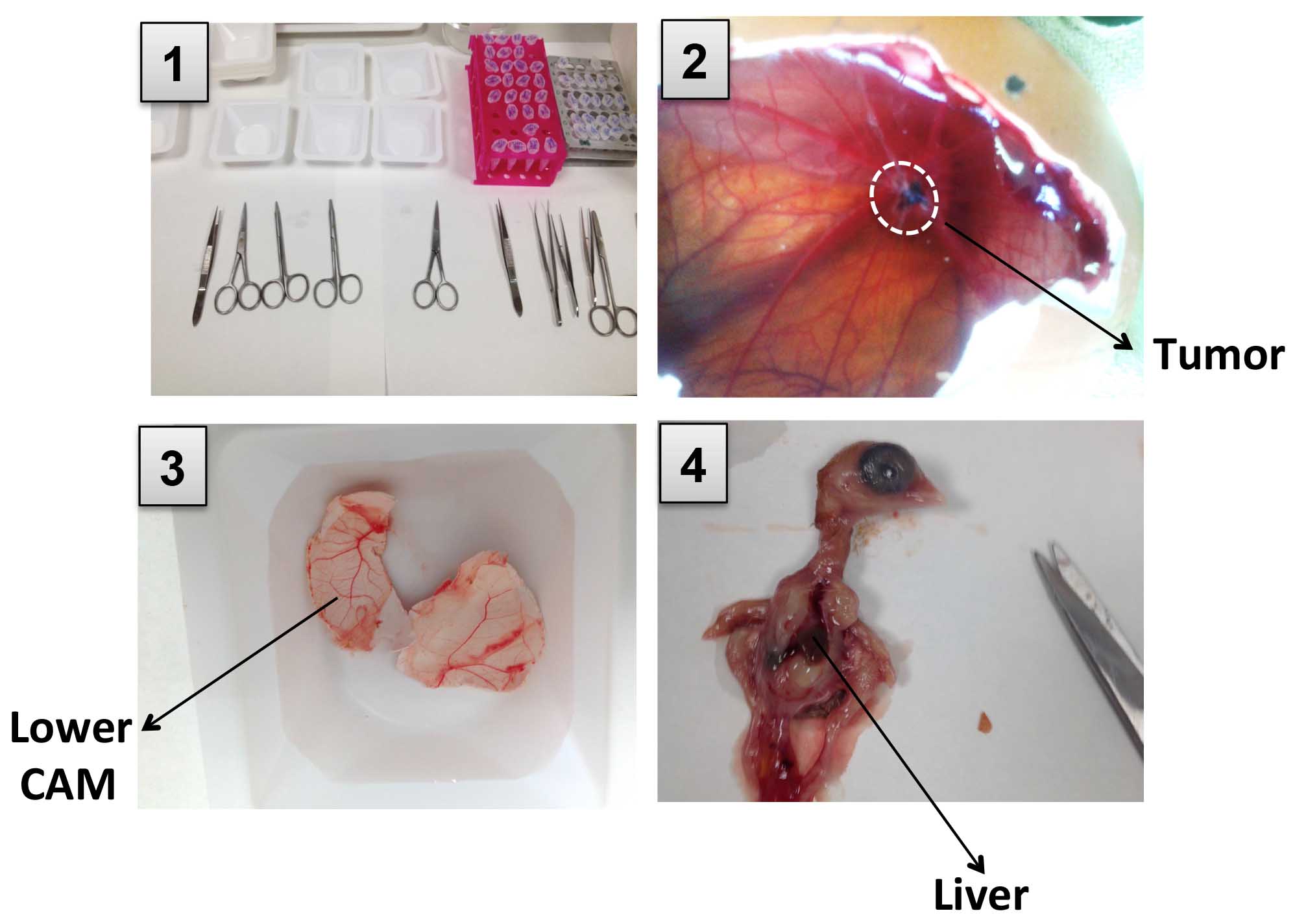
Figure 4. Pictures showing tumor and chicken tissues harvest. 1. Perform dissection in a clean area. 2. Open a new window removing some of the eggshell and resect the primary tumor from the CAM. 3. Cut the eggshell radially into equal halves and harvest lower CAM. 4. Open the chick embryo cutting through the sternum and collect a piece of liver. - Transfer the embryo to a clean weight boat. The animal must be dissected using a clean set of tools.
- Open the chick embryo cutting through the sternum. Once the embryo is open, collect a piece of the liver. To harvest the lung, cut the rib cage and separate from the breast.
- Harvested samples can be processed immediately for DNA isolation or stored at -80 °C for later extraction of DNA, if necessary.
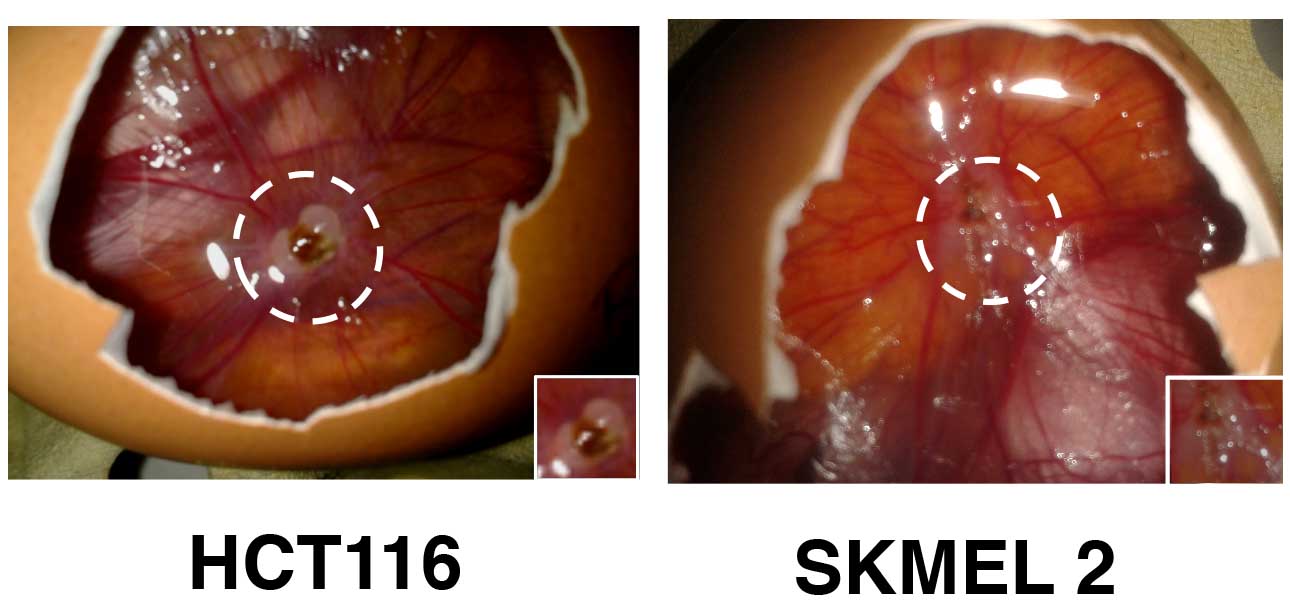
Figure 5. Different tumors have been implanted on the CAM. Tumors were formed using 0.5 x 106 HCT116 colorectal cancer cells or 1 x 106 SKMEL2 melanoma cells during 5 and 7 days respectively. - Genomic DNA isolation
- To extract genomic DNA from tissues, we use the DNA kit from Qiagen. Purified DNA following manufactuer's instructions. DNA can be used immediately or stored at -20 °C. Dilute the DNA 1:50 or 1:25 to get 30 ng/µl in nuclease free water before using it for the PCR reaction.
- To extract genomic DNA from tissues, we use the DNA kit from Qiagen. Purified DNA following manufactuer's instructions. DNA can be used immediately or stored at -20 °C. Dilute the DNA 1:50 or 1:25 to get 30 ng/µl in nuclease free water before using it for the PCR reaction.
- PCR analysis
- To detect human tumor cell DNA in the chick tissues we use quantitative PCR for human Alu sequences using Alu-specific primers and SYBR green mix amplification kit. Prepare the reaction mix in a final volume of 10 μl, with 0.4 μM of each primer. Optimize the amount of template DNA empirically, we usually use 30 ng. Use chicken GAPDH primers as an internal control. The PCR is run under the following conditions: 95 °C for 2 min, 40 cycles at 95 °C for 30 sec, 63 °C for 30 sec, 72 °C for 30 sec.
- Generate a standard curve using a dilution series of human DNA (102, 103, 104) from the original tumor cells. Use this standard curve to quantify human tumor cells in a chick embryonic metastasis assay.
- Measure Ct values in triplicate and against the standard curve calculate the number of tumor cells in each CAM and liver sample.
- Invaded cells are detected using either 30 ng of 50 ng of total CAM DNA. Use water as a negative control.
- To evaluate results and statistical significance of control and experimental groups, use Graph Pad Prism software and Student t-test or ANOVA analysis (Figure 6).
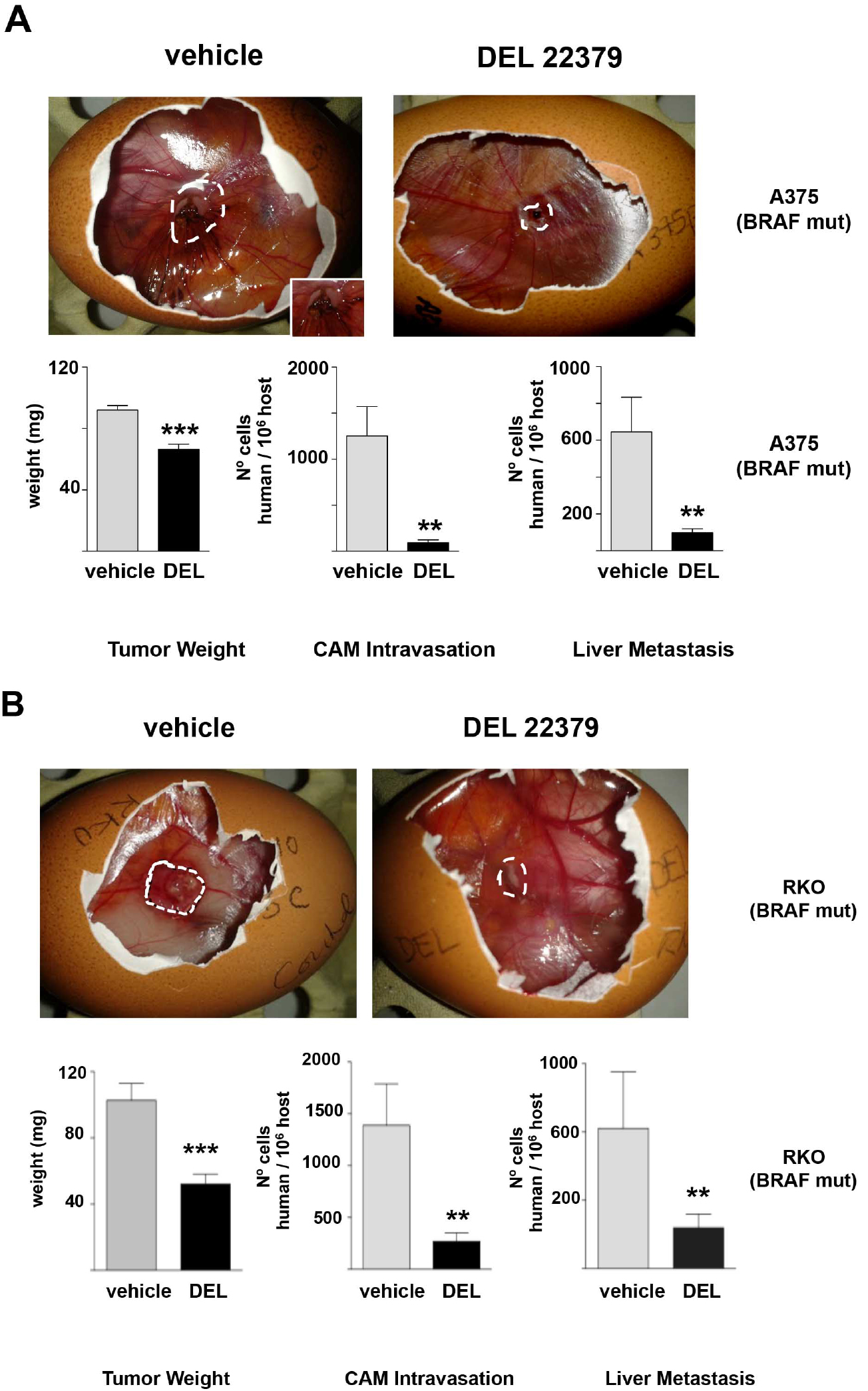
Figure 6. DEL 22379 inhibits tumor growth and metastasis of cancer cells. Tumors were formed using 1 x 106 A375p melanoma cells (A) or 2 x 106 RKO colorectal cancer cells (B). Cells were applied to the CAM of day-10 embryos. Four days after applying the tumor cells half of the embryos were treated with DMSO and the other half with DEL22379 (10 µM). Two days later tumors were harvested to weigh. CAM and chick organs (lung) were harvested from embryos bearing tumors and subjected to quantitative Alu PCR. Conversely, quantitative PCR of chGAPDH was used as an internal control to confirm the presence of equivalent quantities of host genomic DNA. Data shows average ± SEM from 3 independent experiments. **P < 0.05; ***P < 0.005, by unpaired Student t-test (Herrero et al., 2015).
Notes
- A small hook on the end of the 20 G syringe needle is necessary to make the third hole in the eggshell membrane. Try to do it very softly and carefully not to tear the underlying CAM.
- Place the tube on ice during the experiment to avoid the formation of clusters of tumor cells.
Acknowledgments
This protocol was adapted from the previously published studies, Zijlstra et al. (2002), Deryugina et al. (2008), Casar et al. (2014) and it was used in Herrero et al. (2015). We are grateful to Dr. Elena Deryugina and Dr. James Quigley for teaching us the techniques and advise. We thanks to Dr. Marian Ros and Dr. María Felix for providing equipment, technical assistance, and advise. PC lab was supported by grant SAF-2015-63638-R from the Spanish Ministry of Economy-Fondos FEDER and by the Red Temática de Investigación Cooperativa en Cáncer (RTICC) (RD/12/0036/0033), Spanish Ministry of Health. BC was supported by Fundación Francisco Cobos - CSIC.
References
- Casar, B., Rimann, I., Kato, H., Shattil, S. J., Quigley, J. P. and Deryugina, E. I. (2014). In vivo cleaved CDCP1 promotes early tumor dissemination via complexing with activated β1 integrin and induction of FAK/PI3K/Akt motility signaling. Oncogene 33(2): 255-268.
- Deryugina, E. I. and Quigley, J. P. (2008). Chick embryo chorioallantoic membrane model systems to study and visualize human tumor cell metastasis. Histochem Cell Biol 130(6): 1119-1130.
- Herrero, A., Pinto, A., Colon-Bolea, P., Casar, B., Jones, M., Agudo-Ibanez, L., Vidal, R., Tenbaum, S. P., Nuciforo, P., Valdizan, E. M., Horvath, Z., Orfi, L., Pineda-Lucena, A., Bony, E., Keri, G., Rivas, G., Pazos, A., Gozalbes, R., Palmer, H. G., Hurlstone, A. and Crespo, P. (2015). Small molecule inhibition of ERK dimerization prevents tumorigenesis by RAS-ERK pathway oncogenes. Cancer Cell 28(2): 170-182.
- Zijlstra, A., Mellor, R., Panzarella, G., Aimes, R. T., Hooper, J. D., Marchenko, N. D. and Quigley, J. P. (2002). A quantitative analysis of rate-limiting steps in the metastatic cascade using human-specific real-time polymerase chain reaction. Cancer Res 62(23): 7083-7092.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Crespo, P. and Casar, B. (2016). The Chick Embryo Chorioallantoic Membrane as an in vivo Model to Study Metastasis. Bio-protocol 6(20): e1962. DOI: 10.21769/BioProtoc.1962.
Category
Cancer Biology > Invasion & metastasis > Cell biology assays
Cell Biology > Cell movement > Cell migration
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












