- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
EST-SSR Analysis and Cross-species Transferability Study in Lavandula
Published: Vol 6, Iss 15, Aug 5, 2016 DOI: 10.21769/BioProtoc.1891 Views: 10286
Reviewed by: Samik BhattacharyaYi ZhangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
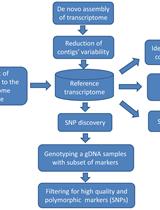
Experimental Pipeline for SNP and SSR Discovery and Genotyping Analysis of Mango (Mangifera indica L.)
Michal Sharabi-Schwager [...] Ron Ophir
Aug 20, 2016 11585 Views
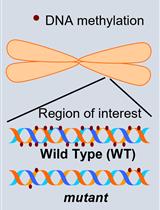
Investigation of Transposon DNA Methylation and Copy Number Variation in Plants Using Southern Hybridisation
Vivek Hari Sundar G. and P. V. Shivaprasad
Jun 5, 2022 3434 Views
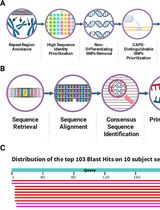
CAPS-Based SNP Genotyping for Nitrogen-Response Phenotypes in Maize Hybrids
Jannis Jacobs [...] Peter K. Lundquist
Dec 20, 2025 588 Views
Abstract
The genus Lavandula comprises of several economically important lavender species that are mainly cultivated worldwide for essential oil production. Identification of lavender species and their cultivars has been a huge bottleneck in lavender industries due to lack of appropriate identification mechanisms. Recent advances in modern technologies would help to address these identification issues through development of potential molecular markers, including simple sequence repeats (SSRs). SSRs can be developed from specific species, and can be potentially used for related species, which lack the source sequences to develop species-specific SSRs. Here, we describe the guidelines and steps of identifying and analyzing SSRs from expressed sequence tag (EST) sequences of lavender species. We also detail the validation procedures of selected EST-SSRs in distinguishing source (donor) species as well as related species.
Keywords: EST-SSRMaterials and Reagents
- 1.5 ml centrifuge tube
- 1-200 µl volume pipette tips
- Leaf tissue (Okanagan Lavender and Herb Farm and The Greenery Garden Center, Kelowna, BC, Canada)
- DNA sequences (expressed sequence tag) (EST)
- Liquid N2 (Praxair)
- Genomic DNA Mini kit (Plant) GP100 (Geneaid Biotech Ltd., catalog number: GP100 )
- 10x PCR buffer (NEB, catalog number: M0320S )
- Taq polymerase (NEB, catalog number: M0320S )
- dNTPs (Omega Bio-Tek, catalog number: TQAC135 )
- MgCl2 solution (NEB, catalog number: M0320S )
- Bovine serum albumin (BSA) (NEB, catalog number: B9000S )
- Custom-synthesized primers (Thermo Fisher Scientific, InvitrogenTM)
- SYBR® Safe DNA gel staining (Thermo Fisher Scientific, InvitrogenTM, catalog number: S33102 )
- 10x DNA loading dye (Ward’s Science, catalog number: 389115 )
- Acrylamide 40% solution (Acrylamide: Bis-Acrylamide 19:1) (Thermo Fisher Scientific, catalog number: BP1406-1 )
- 50 bp DNA ladder (NEB, catalog number: N3236S )
- 1 kb DNA ladder (FroggaBio, catalog number: DM010-R500 )
- Nuclease-free water (Thermo Fisher Scientific, catalog number: BP5611 )
- Ammonium persulfate (APS) (Thermo Fisher Scientific, catalog number: BP179-100 )
- NNN'N'-tetramethyl-ethylenediamine (TEMED) (Sigma-Aldrich, catalog number: T9281 )
- Tris Base (Thermo Fisher Scientific, catalog number: BP152-1 )
- Glacial acetic acid (Thermo Fisher Scientific, catalog number: 351272-212 )
- Na2EDTA (VWR International, catalog number: CA71007-124 )
- Boric acid (Thermo Fisher Scientific, catalog number: B168-1 )
- PCR reaction mix (see Recipes)
- 0.5 M EDTA (see Recipes)
- 50x TAE buffer (see Recipes)
- 1x TAE buffer (see Recipes)
- 5x TBE buffer (see Recipes)
- 0.5x TBE working solution (see Recipes)
- 10% Ammonium persulfate (APS) (v/w) (see Recipes)
- 1 M Tris (see Recipes)
- 10 mM Tris buffer, pH 8.0 (see Recipes)
- 1% agarose gel (Thermo Fisher Scientific, catalog number: BP160-500 ) (see Recipes)
- 6% polyacrylamide gel (see Recipes)
Equipment
- Desktop computer (Dell, model: precision T3610 Tower Workstation )
- Mortar and pastel (VWR International, Porcelain)
- Balance (Shimadzu Corporation, model: ELB300 )
- Freezer (-20 °C) (Frigidaire, model: FFFH20F2QW )
- Thermocycler (Thermo Fisher Scientific, ABI Applied BiosystemTM, model: Veriti® 96-Well Thermal Cycler )
- Horizontal gel electrophoresis (Thermo Fisher Scientific, OwlTM EasyCastTM, model: B2 Mini )
- Vertical gel electrophoresis (Bio-Rad Laboratories, model: Mini-protein Tetra system )
- Microcentrifuge (Eppendorf, model: 5417C )
- Microwave oven (Danby Designer)
- Water bath (Thermo Fisher Scientific, IsotempTM Digital-Control, model: 210 )
- Gel Imager (Mandel Scientific, model: Kodak Gel Logic 440 )
- Sterilmatic Autoclave (Thomas Scientific, model: STME-L )
- Spectrophotometer (Thermo Scientific, NanoDropTM, model: 2000 )
Software
- BatchPrimer 3 (http://probes.pw.usda.gov/cgi-bin/batchprimer3/batchprimer3.cgi) (You et al., 2008)
- SSR mining server (https://www.rosaceae.org/bio/content?title=&url=/cgi-bin/gdr/gdr_ssr) (Jung et al., 2008)
- Perl script MIcroSAtelitte (MISA) identification tool (http://pgrc.ipk-gatersleben.de/misa/)
Note: optional software, but not used in our study - Blast2go online platform (https://www.blast2go.com/) (Conesa et al., 2005)
- OligoAnalyzer 3.1 (https://www.idtdna.com/calc/analyzer)
Procedure
- SSR identification from lavender EST libraries
- Expressed Sequence Tag (EST) databases containing approximately 23,000 sequences are developed for L. angustifolia and L. x intermedia plants (Lane et al., 2010; Demissie et al., 2012). The main steps of EST library development include total RNA isolation, mRNA purification and cDNA library construction, and partial sequencing of random cDNA clones at the 5' ends by Sanger sequencing.
- EST libraries are sorted manually based on the sources of two species, L. angustifolia and L. x intermedia, and are used for simple sequence repeats (SSRs) identification using web-based SSR analysis tools (Jung et al., 2008; You et al., 2008) (Figure 1).
- Sorted ESTs in FASTA format are further broken down manually into multiple files with maximum input sequences of 500 ESTs in BatchPrimer3 software (You et al., 2008). For each of input files, SSR screening parameters are set to pick up primer pairs and SSR motifs with minimum lengths of 12 bp for di-, tri- and tetra-nucleotides, 15 bp for penta-nucleotides as well as 18 bp for hexa-nucleotides. Then, additional parameter settings (minimum, optimum and maximum) are carried out for primer length (18, 21 and 25 bp), product size (120, 200 and 300 bp), GC content (40, 50 and 60%), Tm (50, 55 and 60 °C). Finally, screening of SSRs along with primer sets from input sequences are run, and the outputs are displayed as HTML format and saved as tab-delimited text or excel files for further analysis of SSR repeats.
- Sorted EST files are directly imported into the Genome Database for Rosaceae (GDR) SSR mining server (Jung et al., 2008), and the minimum SSR motifs screening parameters are set to 15 bp for mono- and penta-nucleotides, 12 bp for di-, tri- and tetra-nucleotides, as well as 18 bp for hexa-nucleotides. Other primer generating parameters are kept as default parameters of the software. The outputs are then generated as tab-delimited text file format, which are easily converted into excel files for further SSR repeats analysis. [Optional: automated SSR identification and potential primer designing can be made using Perl script MIcroSAtelitte (MISA) identification tool.]
- SSRs with potential primer flanking regions are further filtered out based on primers designed during SSR motif identification using the aforementioned tools.
- All identified SSRs with primer flanked regions are again characterized, targeting the length of repeats as longer category with SSR length of ≥ 18 bp for tri-nucleotides and ≥ 20 bp for di-, tetra-, penta and hexa-nucleotides, or with lower than those demarcations as shorter category.
- To predict the putative functional roles of the repeats under longer category, ESTs/unigenes containing SSRs with primer flanking regions are annotated against public databases using Blast2GO online platform (Conesa et al., 2005). In brief, ESTs containing repeats ≥ 18 bp are filtered and uploaded into Blast2GO, followed by blasting against the public database. Then, mapping of ESTs with Blast hit are continued prior to the final putative functional annotation. Lastly, based on the detected annotations, characterization of gene ontology (GO) annotation and putative functions of SSR containing ESTs are made.
- Depending on the annotated functions or random representation of identified repeats, unigenes/ESTs are selected and used to refine manually the designed primer pairs following accordingly the minimum, optimum and maximum parameters: primer length (18, 21 and 25 bp), product size (120, 200 and 300 bp), GC content (40, 50 and 60%), Tm (50, 55 and 60 °C) and others using OligoAnalyzer 3.1 online software. (Optional: primer designing can be done by any primer designing tools.)
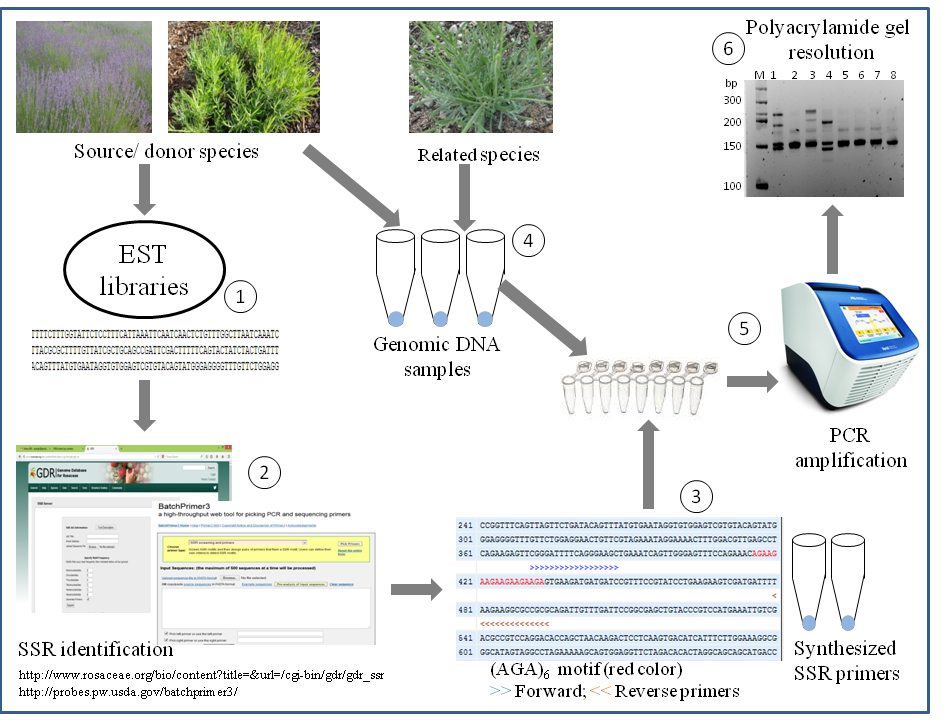
Figure 1. Overview of SSRs analysis from EST libraries of lavender species and validation of identified SSR on source as well as related Lavandula species. Major steps of SSR analysis from ESTs and subsequent validation include: (1) EST library development; (2) SSR motif identification using web-based mining tools; (3) SSR characterization, primer design and custom-synthesis of primers; (4) genomic DNA isolation; (5) PCR amplification of expected SSR motif-containing fragments; and (6) resolution of DNA fragments on polyacrylamide gel for scoring of amplicons as presence (1) or absence (0).
- Expressed Sequence Tag (EST) databases containing approximately 23,000 sequences are developed for L. angustifolia and L. x intermedia plants (Lane et al., 2010; Demissie et al., 2012). The main steps of EST library development include total RNA isolation, mRNA purification and cDNA library construction, and partial sequencing of random cDNA clones at the 5' ends by Sanger sequencing.
- Validating and assessing cross-species transferability of SSRs
- Selected primer sets from aforementioned procedures are custom-synthesized and resuspended in 10 mM Tris (pH 8.0) buffer and stored at -20 °C until use.
- Fresh young leaves (~100 mg) per sample of both source and related species (Figures 1 and 2) are ground with liquid N2 to a fine powder with mortar and pestle, and transferred into 1.5 ml centrifuge tube containing 450 µl cell lysis buffer (GP1 or GPX1) and RNase A (5 µl) from the plant genomic DNA extraction kit. Note that GPX1 is used for samples of mature leaf tissues (Figure 2B), mostly rich in phenolic compounds. The samples are then incubated at 60 °C in water bath for 10-15 min, mixing by inverting every 5 min. The remaining procedures are done according to the kit's instructional manual. Finally, each of the samples is eluted with a total of 40 µl elution buffer with two round elution (25 µl and 15 µl, respectively) after 3-5 min incubations at room temperature. Eluted genomic DNA (gDNA) is quantified using NanoDrop 2000 spectrophotometer (1 µl elution buffer for blank and 1 µl gDNA for each sample). To check the gDNA integrity, 10 µl of the gDNA (100-300 ng) mixed with 1 µl of 10x loading dye and autoclaved water is loaded into each well of an agarose gel (1%) and run on horizontal gel electrophoresis with 1x TAE buffer at 80 V for 30-45 min (Figure 3). DNA image is then visualized and taken by the Gel Imager (Figure 3). Finally, a working solution is made for each sample into 50-70 ng/µl and stored at -20 °C until use.
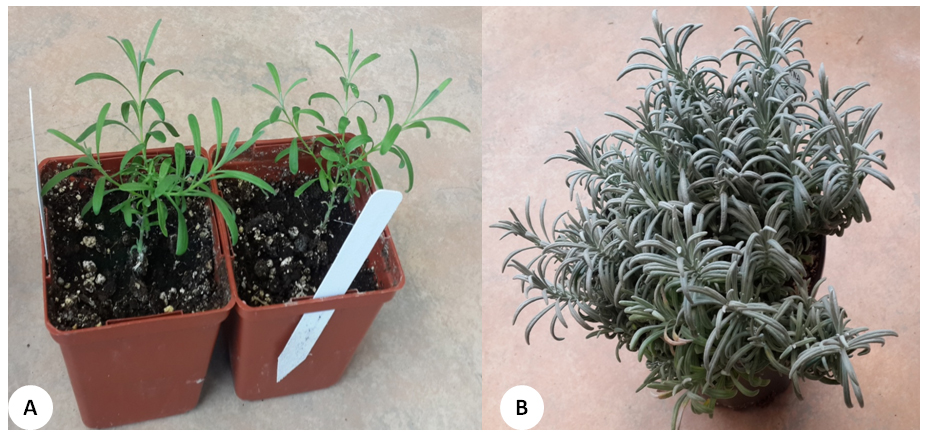

Figure 2. L. x intermedia cv Grosso plants for genomic DNA extraction. A. Young plants. B. Mature plant.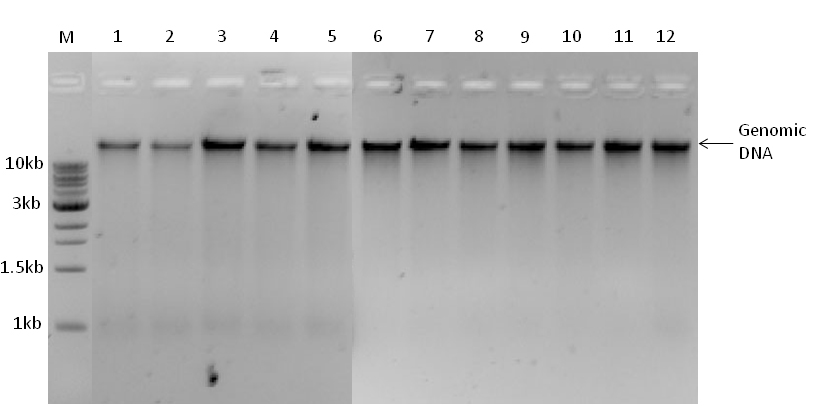
Figure 3. Agarose gel (1%) resolution of genomic DNA samples (200 ng/well) isolated from young leaf tissues of L. x intermedia (lanes 1-5) and L. angustifolia (lanes 6-12) plants. M, 1kb DNA ladder (300 ng). - PCR amplification for each primer set is performed by touch-down PCR using a two-stage amplification program (Figure 1). For the first stage, the Thermocycler is set to 15 min denaturation at 95 °C followed by 11 cycles of 30 sec denaturation at 95 °C, 30 sec annealing at 64-54 or 62-52 °C (depending on the primer type) with dropping of 1 °C in every cycle for annealing and 2 min at 72 °C for extension. For the second stage, the Thermocycler is programmed to perform 24 cycles at 95 °C for 30 sec, 54 °C or 52 °C for 30 sec and 72 °C for 2 min, and completed by a final extension at 72 °C for 10 min.
- The PCR amplified products are separated on a 6% polyacrylamide gel (Figure 4). Samples that are well mixed with loading dye (9 µl PCR products and 1 µl 10x loading dye) are loaded into gel wells using standard 1-200 µl volume pipette tips. One well is left empty to load a DNA ladder mix containing 0.35 µl of 50 bp DNA ladder (1 µg/ µl), 1 µl of 10x loading dye and 8.65 µl of autoclaved water. Once all samples and DNA ladder are carefully loaded, the gel is run in vertical gel electrophoresis using 0.5x TBE buffer at 160 V for approximately 45 min.
- The gel is then carefully transferred into a plastic container with 50 ml 0.5x TBE buffer supplemented with 1 µl SYBR safe and rocked for 30 min. After a quick rinse of the stained gel with fresh 0.5x TBE buffer, a gel image is taken using the Gel Imager (Figure 4).
- SSR primers from either L. angustifolia or L. x intermedia ESTs are validated for their polymorphism within the donor species (same as source species) as well as cross-species transferability to other related species. (Optional: A single amplicon of every targeted repeat can further be gel purified and sequenced to verify the authenticity of the designed primer sets that amplify the desired fragments.)
- Because of the complex polyploidy nature of Lavandula genome, we assess the polymorphism and cross-species amplification based on presence or absence of amplified SSR fragments. If the target fragments with expected sizes are detected, the data is recorded as “presence” (1). However, if the target fragments do not meet the expected size or there is no amplification, it is recorded as “absence” (0) for that particular fragment. As a demonstration, strong LAF15 amplicons across the given samples are assessed (Figure 4). Five fragments (~145, 155, 185, 200, 250 bp) from the two species are detected and scored. The scoring is made for each fragment across 15 samples as "1" for detected fragments and "0" for no or faint amplifications. For "~145 bp" and "~155 bp", the patterns of the scoring from sample-1 to sample-15 are "100100001110100" and "111111111111111" respectively. The remaining three fragments (~185, 200, 250 bp) are scored as "1000000000100100", "000100001100000" and "101000000000000", respectively. These scored data can be managed in excel sheet in line with the formats of the subsequent analysis software. A similar scoring pattern is also used for samples of related species. (Optional: Fluorescent dye-labeled primers can be custom-synthesized and used for automated capillary electrophoresis detection with the help of fluorescent dye tagged to the fragments during amplification with fluorescent dye-labeled primers. This option is also useful to detect the exact numbers and sizes of amplicons per a primer set.)
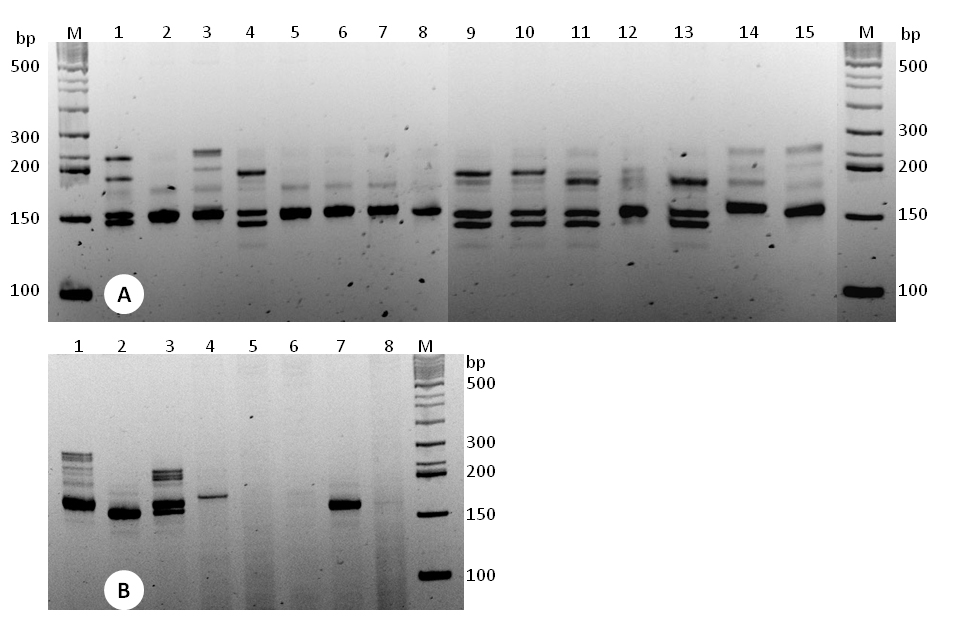
Figure 4. Polyacrylamide gel (6%) resolution of EST-SSR (LAF15) amplicons from (A) samples of L. angustifolia (lanes 1-9) and L. x intermedia (lanes 10-15) plants, and from (B) samples of related species, including L. angustifolia (lane 1) and L. x intermedia (lane 3). M, 50 bp DNA ladder; lane 2, L. latifolia; lane 4, L. buchii; lane 5. L. dentata; lane 6. L. lusitanica; lane 7, L. x ginginsii; lane 8, L. stoechas.
- Selected primer sets from aforementioned procedures are custom-synthesized and resuspended in 10 mM Tris (pH 8.0) buffer and stored at -20 °C until use.
Notes
Mature leaf tissue may contain large amounts of phenolic compounds that can co-purify with genomic DNA, and interfere with the subsequent PCR amplification reactions. Thus, extra caution must be taken on selection of tissue for DNA extraction. Young and fresh leaf tissues (Figure 2A) often give better DNA than mature and dry tissues.
Recipes
- PCR reaction mix, 25 µl
0.3 µM forward primer
0.3 µM reverse primer
70 ng genomic DNA template
2.5 µl 10x PCR buffer
1.5 mM MgCl2
0.5 µg BSA
250 µM dNTPs
1.25 U Taq polymerase
Nuclease-free water - 0.5 M EDTA (pH 8.0)
Add 93.05 g of Na2EDTA in 400 ml of RO water
Stir thoroughly and adjust pH 8.0 with pellet NaOH (the solution becomes transparent when the pH reaches to 8.0)
Bring volume to 0.5 L with RO water
Autoclave and store at RT - 50x TAE buffer, 0.5 L
121 g Tris base (MW = 121.1 g/mol)
28.5 ml glacial acetic acid
50 ml 0.5 M EDTA (pH 8.0)- Dissolve 121 g Tris in 300 ml sterile reverse osmosis (RO) water with gently stirring
- Add EDTA and glacial acetic acid
- Bring the final volume to 0.5 L with sterile RO water
- Labeled properly and autoclave at 121 °C for 30 min
- Store at RT
- Dissolve 121 g Tris in 300 ml sterile reverse osmosis (RO) water with gently stirring
- 1x TAE buffer, 4 L
- Measure 80 ml 50x TAE
- Bring the final volume to 4 L with RO water
- Store at RT
- Measure 80 ml 50x TAE
- 5x TBE buffer, 1 L, pH 8.3
54 g Tris base (MW = 121.1 g/mol)
27.5 g boric acid (MW = 61.83 g/mol)
20 ml of 0.5 M EDTA (pH 8.0)- Dissolve Tris base in 600 ml RO water with gentle stirring
- Add boric acid
- Add EDTA solution
- Adjust pH to 8.3 and bring up the final volume to 1 L
- Autoclave at 121 °C for 30 min and store at RT
- Dissolve Tris base in 600 ml RO water with gentle stirring
- 0.5x TBE working solution, 2 L
- Measure 200 ml 5x TBE
- Bring the final volume to 2 L with RO water
- Store at RT
- Measure 200 ml 5x TBE
- 10% Ammonium persulfate (APS) (v/w)
Dissolve 1 g APS (MW = 228.2 g/mol) in 10 ml sterile RO water
Store in 4 °C (prepared fresh after 30 days) - 1 M Tris (pH 8.0), 0.5 L
- Dissolve 60.55 g Tris base in 400 ml
- Add 20 ml concentrated HCl
- Bring the final volume to 0.5 L with RO water
- Dissolve 60.55 g Tris base in 400 ml
- 10 mM Tris (pH 8.0) buffer, 100 ml
Take 2 ml 1 M Tris (pH 8.0)
Bring the final volume to 100 ml with sterilized RO water
Store at 4 °C - 1% agarose gel
Add 0.5 g of Agarose into 250 ml Erlenmeyer flask containing 50 ml 1x TAE buffer
Boil in microwave with frequent monitoring until it gets clear
Cool down to ~50 °C
Add 1 µl SYBR safe and mix gently
Pour into the gel cast to solidify - 6% polyacrylamide gel, 1.5 mm gel (10 ml)
Add sequentially:- 5.73 RO water
- 1.67 ml Acrylamide 40% solution [Acrylamide and Bis-Acrylamide (19:1)]
- 2.4 ml 5x TBE buffer
- 200 µl 10% APS
- 10 µl TEMED
Wait till completely polymerized - 5.73 RO water
Acknowledgments
This protocol was adapted from the previously published study, Adal et al. (2015). This work was supported through grants and/or in-kind contributions to SSM by UBC, Genome British Columbia, Natural Sciences and Engineering Research Council of Canada, Agriculture and Agri-Food Canada and the BC Ministry of Agriculture (through programs delivered by the Investment Agriculture Foundation of BC).
References
- Adal, A. M., Demissie, Z. A. and Mahmoud, S. S. (2015). Identification, validation and cross-species transferability of novel Lavandula EST-SSRs. Planta 241(4): 987-1004.
- Conesa, A., Gotz, S., Garcia-Gomez, J. M., Terol, J., Talon, M. and Robles, M. (2005). Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18): 3674-3676.
- Demissie, Z. A., Cella, M. A., Sarker, L. S., Thompson, T. J., Rheault, M. R. and Mahmoud, S. S. (2012). Cloning, functional characterization and genomic organization of 1,8-cineole synthases from Lavandula. Plant Mol Biol 79(4-5): 393-411.
- Jung, S., Staton, M., Lee, T., Blenda, A., Svancara, R., Abbott, A. and Main, D. (2008). GDR (Genome Database for Rosaceae): integrated web-database for Rosaceae genomics and genetics data. Nucleic Acids Res 36(Database issue): D1034-1040.
- Lane, A., Boecklemann, A., Woronuk, G. N., Sarker, L. and Mahmoud, S. S. (2010). A genomics resource for investigating regulation of essential oil production in Lavandula angustifolia. Planta 231(4): 835-845.
- You, F. M., Huo, N., Gu, Y. Q., Luo, M. C., Ma, Y., Hane, D., Lazo, G. R., Dvorak, J. and Anderson, O. D. (2008). BatchPrimer3: a high throughput web application for PCR and sequencing primer design. BMC Bioinformatics 9: 253.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Adal, A. M., Demissie, Z. A. and Mahmoud, S. S. (2016). EST-SSR Analysis and Cross-species Transferability Study in Lavandula. Bio-protocol 6(15): e1891. DOI: 10.21769/BioProtoc.1891.
Category
Plant Science > Plant molecular biology > DNA > Genotyping
Molecular Biology > DNA > Genotyping
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










