- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An Improved and Simplified Radial Gel Diffusion Assay for the Detection of Xylanase Activity
Published: Vol 6, Iss 13, Jul 5, 2016 DOI: 10.21769/BioProtoc.1863 Views: 9128
Reviewed by: Zhaohui LiuIgor Cesarino Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Silencing Arbuscular Mycorrhizal Fungal Gene Using Chitosan Nanoparticle-Mediated dsRNA Delivery System
Chumei Yan [...] Xianan Xie
Jun 5, 2025 2638 Views
In Vitro Screening of Microbial Extracts Against the Oomycetes Phytophthora capsici and Pythium ultimum
Mónica Trigal Martínez [...] María Ángeles Vinuesa Navarro
Sep 20, 2025 1263 Views

A Reliable In Planta Inoculation and Antifungal Screening Protocol for Rhizoctonia solani-Induced Sheath Blight in Rice
Alinaj Yasin [...] Palash Deb Nath
Nov 5, 2025 1572 Views
Abstract
Xylanase (E.C. 3.2.1.8) degrades β-1, 4 xylan by cleaving β-1, 4 glycosidic linkages randomly, resulting in the generation of xylose and xylo-oligosaccharides. Xylanases are produced by organisms including fungi, bacteria, yeast, marine algae, protozoans, snails, crustaceans and insects. Xylanases present considerable industrial interest for their use in paper manufacturing, improvement of animal feed digestibility, and clarification of fruit juices. In addition, this enzyme is the component of cell wall-degrading enzymes (CWDEs) during plant–pathogen interaction. Thus, considering their various applications in plant defence and also in industry, the characterization of xylanase activity becomes an important aspect. Conventionally, xylanase activity is determined by radial gel diffusion assay using Congo red staining (Emami and Hack, 2001) and by DNSA assay which is a colorimetric method for xylanase activity (McLauchlan et al., 1999; Kutasi et al., 2001). Comparatively, radial gel diffusion assay using Congo red staining is a qualitative assay whereas DNSA method is a quantitative assay. Moreover, Congo red is a chemical considered as hazardous category 1B (Carcinogenicity) and category 12 (Reproductive toxicity) by the 2012 OSHA Hazard Communication Standard (29 CFR 1910.1200). In the present study, the proposed method enables qualitative detection of xylanase activity using ethanol precipitation in the radial gel diffusion assay which is safer and simpler. The ethanol precipitation in agar plate has been adapted from the method for detecting xylanase activity in polyacrylamide gels (Royer and Nakas, 1990).
Keywords: XylanaseMaterials and Reagents
- 90 x 15 mm Petri plates (SARSTEDT AG, catalog number: 82.1473001 )
- 0.5-cm-diameter drinking straw
- 150 ml Erlenmeyer flask
- Aspergillus niger xylanase (Xylanase M4) (Megazyme) or Trichoderma longibrachiatum xylanase (Xylanase M3) (Megazyme)
- Disodium hydrogen phosphate (Sigma-Aldrich, catalog number: S5136 )
- Citric acid (Sigma-Aldrich, catalog number: C1909 )
- Birch wood xylan (Sigma-Aldrich, catalog number: X0502 )
- Agarose (AppliChem GmbH, catalog number: A8963.0500 )
- Absolute ethanol (VWR International, catalog number: 20821.321 )
- McIlvaine’s buffer (pH 5.0) (see Recipes)
- 95% (v/v) ethanol (see Recipes)
Equipment
- Microwave
- Hallow flat end with 0.5-cm-diameter drinking straw
- 30 °C incubator (Mini BATT 805) (230 V, 50 Hz, 270 W) (Asal Srl, model: 805 )
- Scanner camera (Epson, model: perfection V30 )
Procedure
- Mix 1.0% (w/v) birch wood xylan with 1.0% (w/v) agarose in McIlvaine’s buffer (15 ml/Petri dish) and boil in 150 ml Erlenmeyer flask (approximately 10 min) until the birch wood xylan is dissolved completely; mix thoroughly and then dispense it in a Petri plate (44 mm base diameter x 12 mm depth).
- Allow the birch wood xylan–agarose solution to solidify in the Petri dish and then prepare a 0.5-cm-diameter well on it. The hole in agarose is poked using a hallow flat end of a 0.5-cm-diameter drinking straw.
- Place 30 µl of A. niger or T. longibrachiatum xylanase solution (total volume to be adjusted with McIlvaine’s buffer) in 0.5-cm-diameter wells of birch wood xylan-agarose Petri plate.
- Incubate the Petri plate at 30 °C for 16 h.
- Overlay the plate with 95% ethanol and keep the plate at room temperature for 30 min to reveal the halo representing the degradative activity of xylanase.
- Measure the diameter of the halo using a scale.
- Take the image using a digital camera or scan the Petri plate using a scanner to keep the record of the results (Figure 1).
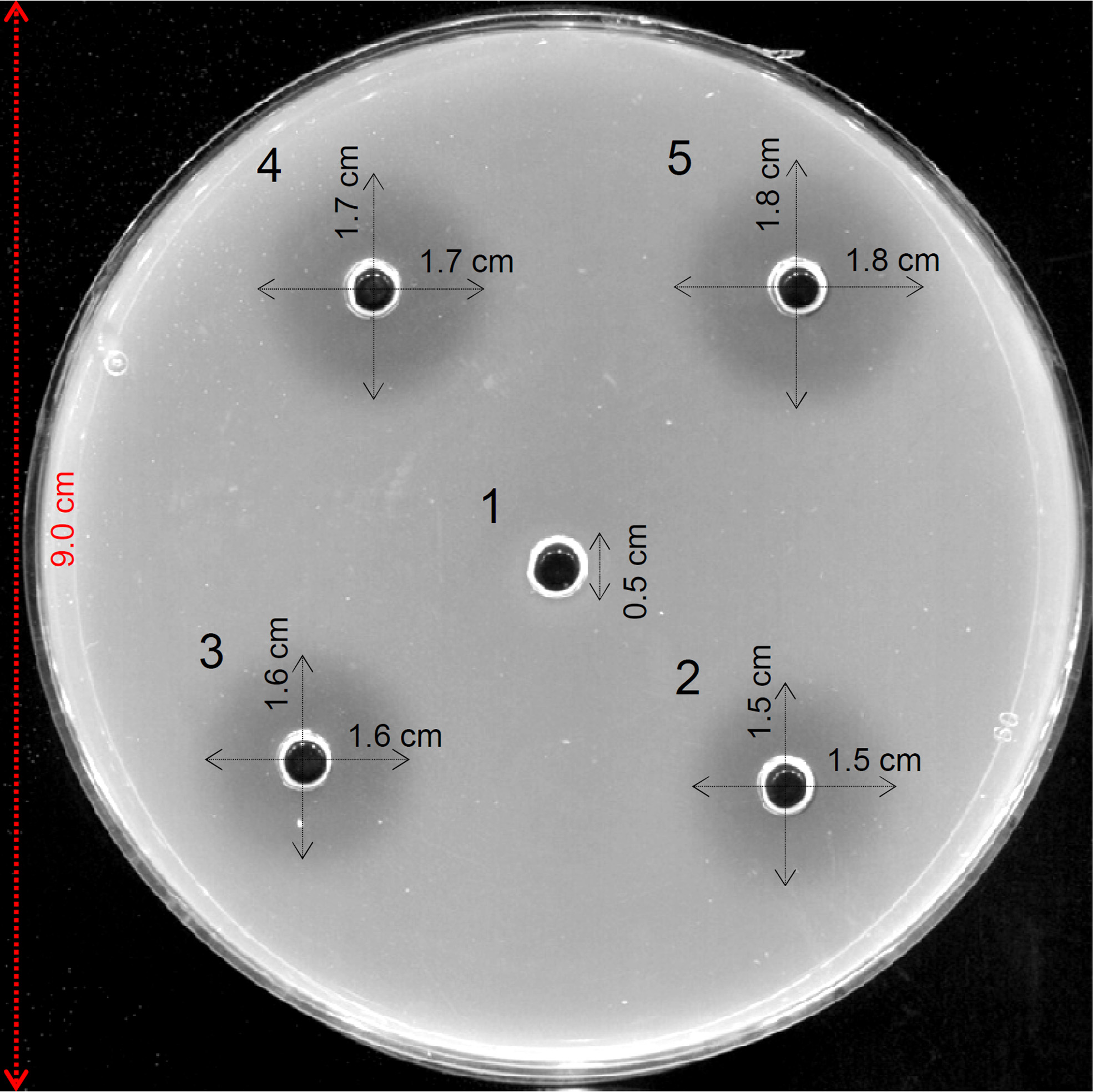

Figure 1. Example of agarose diffusion assays for detection of xylanase activity. Haloes represent xylanase activity. 1: Buffer (negative control); 2: Aspergillus niger xylanase M4 (0.006U); 3: A. niger xylanase M4 (0.012U); 4: A. niger xylanase M4 (0.018U); 5: A. niger xylanase M4 (0.024U).
Recipes
- McIlvaine’s buffer (pH 5.0)
0.2 M disodium hydrogen phosphate
0.1 M citric acid - 95% (v/v) absolute ethanol
Mix 95 ml absolute ethanol with 5% (v/v) distilled water
Acknowledgments
This protocol has been adapted from the previously published by Kalunke et al. (2013). This protocol was designed to determine xylanase inhibition for wheat transgenic plants overexpressing the xylanase inhibitor TAXI‐III.
Research was supported by the Italian Ministry of University and Research (PRIN 2010-2011) to Renato D’Ovidio.
References
- Emami, K., Hack, E. (2001). Characterisation of a xylanase gene from Cochliobolussativus and its expression. Mycol. Res. 105 (3): 352-359.
- Kalunke, R. M., Janni, M., Benedettelli, S. and D’Ovidio, R. (2013). Using biolistics and hybridization to combine multiple glycosidase inhibitor transgenes in wheat. Euphytica 194(3): 443-457.
- Kutasi, J., Bata, A., Brydl, E., Rafai, P. and Jurkovich, V. (2001). Characterisation and effects of a xylanase enzyme preparation extracted from Thermomyces lanuginosus cultures. Acta Vet Hung 49(2): 175-184.
- McLauchlan, W. R., Garcia-Conesa, M. T., Williamson, G., Roza, M., Ravestein, P. and Maat, J. (1999). A novel class of protein from wheat which inhibits xylanases. Biochem J 338 (Pt 2): 441-446.
- Royer, J. C. and Nakas, J. P. (1990). Simple, sensitive zymogram technique for detection of xylanase activity in polyacrylamide gels. Appl Environ Microbiol 56(6): 1516-1517.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kalunke, R. M., Moscetti, I., Tundo, S. and D’Ovidio, R. (2016). An Improved and Simplified Radial Gel Diffusion Assay for the Detection of Xylanase Activity. Bio-protocol 6(13): e1863. DOI: 10.21769/BioProtoc.1863.
Category
Biochemistry > Protein > Activity
Microbiology > Microbial biochemistry > Protein
Plant Science > Plant immunity > Host-microbe interactions
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












