- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
HPLC Analysis of Secreted Organic Acids
Published: Vol 6, Iss 8, Apr 20, 2016 DOI: 10.21769/BioProtoc.1786 Views: 12054
Reviewed by: Maria SinetovaElizabeth LibbyRoman A. Sidorov

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

β-lactamase (Bla) Reporter-based System to Study Flagellar Type 3 Secretion in Salmonella
Fabienne F. V. Chevance and Kelly T. Hughes
Jun 20, 2023 1807 Views
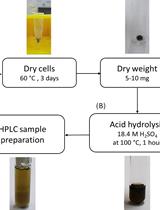
Determination of Poly(3-hydroxybutyrate) Content in Cyanobacterium Synechocystis sp. PCC 6803 Using Acid Hydrolysis Followed by High-performance Liquid Chromatography
Janine Kaewbai-ngam [...] Tanakarn Monshupanee
Aug 20, 2023 1872 Views
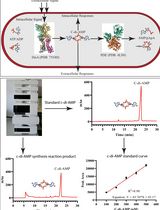
An HPLC-based Assay to Study the Activity of Cyclic Diadenosine Monophosphate (C-di-AMP) Synthase DisA from Mycobacterium smegmatis
Avisek Mahapa [...] Dipankar Chatterji
Dec 20, 2024 1819 Views
Abstract
Under certain growth conditions some microorganisms secrete organic acids into the extracellular medium to relieve the accumulation of excess energy carriers, and/or to reduce toxic concentrations of organic acids. For example, a glycogen-deficient ∆glgC mutant of the cyanobacterium Synechococcus sp. PCC 7002 secretes pyruvate, acetate, α-ketoglutarate, α-ketoisocaproate and succinate (Davies et al., 2014; Jackson et al., 2015). Secretion of these organic acids functions as a putative energy-spilling mechanism in the absence of glycogen, the major carbon and reductant sink in this organism. Identification of secreted organic acids can facilitate the design of metabolic engineering strategies that funnel over-accumulating organic acids towards metabolic pathways that make a product of interest (such as a biofuel). Here, we describe a method for analyzing secreted organic acids in the extracellular media using high-performance liquid chromatography (HPLC). This method was developed for analysis of organic acids secreted by photosynthetic microbes (cyanobacteria and algae) into media, but could be used to analyze organic acids secreted by any microorganism cultivated in liquid medium.
Keywords: Organic acidsMaterials and Reagents
- 2 ml Eppendorf tubes (VWR international, catalog number: 20170-170 )
- PIPETMAN Classic P1000 pipette (Gilson Scientific Ltd., catalog number: F123602 )
- P1000 pipette tips (VWR international, catalog number: 83007-376 )
- 1 ml disposable syringe (BD, catalog number: 309659 )
- Syringe filter, 0.45 µm PTFE membrane (Pall Corporation, catalog number: PN4543 )
- Disposable filter unit (0.45 µm) (Thermo Fisher Scientific, NalgeneTM, catalog number: 166-0045 )
- Screw-thread chromatography vials (VWR International, catalog number: 66009-858 )
- Cyanobacterium Synechococcus sp. PCC 7002
- Liquid culture (cyanobacteria or other microorganism)
- Sulfuric acid (H2SO4) (Merck Millipore Corporation, catalog number: SX1244 )
- Succinate (Sigma-Aldrich, catalog number: S3674 )
- α-ketoglutarate acid (Sigma-Aldrich, catalog number: 75890 )
- Acetic acid (Sigma-Aldrich, catalog number: 338826 )
- Pyruvate (Sigma-Aldrich, catalog number: 107360 )
- α-ketoisocaproate (Sigma-Aldrich, catalog number: 68255 )
Note: It is also named “4-Methyl-2-oxovaleric acid” on Sigma-Aldrich website. - 8 mM H2SO4 (see Recipes)
- 50 mM succinate stock solution (see Recipes)
- 50 mM α-ketoglutarate stock solution (see Recipes)
- 50 mM acetic acid stock solution (see Recipes)
- 50 mM pyruvate stock solution (see Recipes)
- 50 mM α-ketoisocaproate stock solution (see Recipes)
- 10 mM organic acids standard mixture (see Recipes)
Equipment
- Microcentrifuge (Beckman Coulter, catalog number: B30147 )
- Surveyor Plus HPLC (Thermo Fisher Scientific) composed of
- Surveyor LC pump
- Surveyor Autosampler (AS)
- Surveyor Photo Diode Array (PDA) Plus detector
- Surveyor Refractive Index (RI) Plus detector
- Aminex fermentation monitoring column (150 mm by 7.8 mm), stationary phase Polystyrene-divinylbenzene sulfonic acid resin, 9 μM particle size, 8% cross linkage (Bio-Rad Laboratories, catalog number: 1250115 )
- Micro-Guard Cation H Cartridge guard column, 30 x 4.6 mm, hydrogen form, pH range 1-3, for Aminex® hydrogen-form columns (Bio-Rad Laboratories, catalog number: 1250129 )
- Surveyor LC pump
- ChromQuest™ Software Platform (Thermo Fisher Scientific, catalog number: INQSOF012 )
Software
- Chromquest software
Procedure
- Sample preparation
- Using a pipette, remove a 2 ml aliquot from the liquid culture, place in a 2 ml Eppendorf tube, and centrifuge at 13,000 x g for 10 min to pellet the cells.
- Remove and keep the supernatant; this is the extracellular medium that contains secreted organic acids. At this point the supernatant may be frozen at -20 °C for later HPLC analysis or analyzed immediately as described in the following steps.
- Filter 500-700 µl of the sample supernatant into chromatography vials using a 1 ml disposable syringe and a 0.45 µm-pore-size filter to remove particles that may interfere with the organic acid detection and potentially clog values and lines. Repeat individually for each organic acid standard sample that will be used to construct the standard curve. Cap the vials. These are now ready for HPLC analysis.
- Using a pipette, remove a 2 ml aliquot from the liquid culture, place in a 2 ml Eppendorf tube, and centrifuge at 13,000 x g for 10 min to pellet the cells.
- HPLC procedure
- Use 8 mM H2SO4 as the mobile phase for isocratic elution.
- Sequentially increase the column temperature to 35 °C and then 45 °C using the Chromquest software (use a mobile phase flow rate of 0.1 ml min-1). Set the Refractive Index Detector to 50 °C.
- Once the column has reached 45 °C, download a method for measuring samples with the following settings. These settings are a standard protocol for the fermentation monitoring column; the only optimization is the increased run time to ensure the sample has completely eluted from the column.
- Surveyor LC pump: total flow, 0.5 ml/min; pressure limits, 0-1,100 psi; run time, 40 min.
- Surveyor PDA Plus: run time, 40 min; scans, 200-360 nm, scan rate, 1.0 Hz; bandwidth 1 nm.
- Surveyor AS: injection volume, 25 µl; needle height from bottom, 2.0 mm; syringe speed, 8 µl sec-1; flush speed, 70 µl sec-1; flush volume, 500 µl; wash volume, 300 µl; flush/wash source, 8 mM H2SO4 mobile phase; set tray temperature to 10 °C; enable column oven control temperature to 45 °C.
- Surveyor RI Plus: run time, 40 min; data rate, 10 Hz; temperature control set to 50 °C.
- Surveyor LC pump: total flow, 0.5 ml/min; pressure limits, 0-1,100 psi; run time, 40 min.
- Wait 10 min for baseline to stabilize, place vials in the autosampler tray, create a new sequence file for your run, autozero the PDA Plus and RI detectors, and start the run.
- Use 8 mM H2SO4 as the mobile phase for isocratic elution.
- Data analysis
- Typical chromatograms generated by the PDA Plus detector are shown in Figure 1. Similar chromatograms are generated by the RI detector. The organic acids secreted by different strains (Figure 1A) can be identified by comparing retention times to those of known standards (Figure 1B).
- Use the Chromquest software to integrate and quantify the area under each organic acid peak.
- Create a standard curve for each organic acid by plotting the peak area vs the known concentration. Example calibration curves are shown in Figure 2.
- Use the standard curves to calculate the concentrations of each organic acid in the supernatant samples. This will give the concentration of organic acid present in the extracellular media. Data can be normalized to cell count to determine the amount of organic acids secreted per cell.

Figure 1. HPLC chromatograms generated by the PDA detector. A. Extracellular media from wild-type and ∆glgC cells. B. 4 mM mixture of organic acids standards. The retention times for each organic acid using our experimental procedure are as follows: α-ketoglutarate, 5.74 min; pyruvate, 6.52 min; succinate, 7.36 min; acetate, 9.51 min; α-ketoisocaproate, 10.45 min.
Figure 2. Example calibration curves constructed by measuring the peak area of varying concentrations of organic acid standards. A. Calibration curve obtained using PDA detector. B. Calibration curve obtained using RI detector. - Typical chromatograms generated by the PDA Plus detector are shown in Figure 1. Similar chromatograms are generated by the RI detector. The organic acids secreted by different strains (Figure 1A) can be identified by comparing retention times to those of known standards (Figure 1B).
Notes
- Liquid cultures must be of sufficient cell density to produce detectable concentrations of organic acids in the extracellular media. For example, cyanobacterial cultures were inoculated at a dry weight of 0.3 g L-1, and sampled anywhere between a dry weight of 0.3 g L-1 to 2.0 g L-1 for our experiments.
- Both the PDA and RI detectors will detect organic acids and so chromatograms from either detector may be used for organic acid analysis; however, each detector has specific advantages and disadvantages. RI detectors are often preferred because they have universal detection capabilities and respond to the bulk property of the analyte, i.e. its refractive index. However, one limitation of an RI detector is a lack of sensitivity. UV-Vis detectors (such as PDA) are popular because they are a simple, reliable, and sensitive detector. However, PDA detectors are unable to detect most alcohols often a co-secreted metabolite.
- Standards may be run individually (in addition to the standard-curve mixtures) to identify the specific retention time for each organic acid.
Recipes
- 8 mM H2SO4 (pH 2)
0.898 ml H2SO4 (98%)
Add dH2O to 1,000 ml
Filter with 0.45 µm filter unit - 50 mM succinate (stock solution, store frozen)
14.76 mg succinate
Add dH2O to 25 ml - 50 mM α-ketoglutarate (stock solution, store frozen)
18.26 mg α-ketoglutarate
Add dH2O to 25 ml - 50 mM acetic acid (stock solution, store frozen)
71.4 µl acetic acid
Add dH2O to 25 ml - 50 mM pyruvate (stock solution, store frozen)
88.65 µl pyruvate (98%)
Add dH2O to 25 ml - 50 mM α-ketoisocaproate (stock solution, store frozen)
154.19 µl α-ketoisocaproate (>98%)
Add dH2O to 25 ml - 10 mM organic acids standard mixture (store frozen)
Mix 2 ml of each of the five 50 mM organic acid standards above to give a 10 mM solution of mixed organic acids (final volume 10 ml).
Perform a serial dilution of this mixed organic acid solution to generate a range of concentrations appropriate for constructing a standard curve; for example, 5 mM, 2 mM, 1 mM, 0.5 mM, 0.25 mM, 0.1 mM. The range of concentrations of the standard curve should span those concentrations measured in the extracellular samples.
Acknowledgments
This protocol was adapted from the previously published study, D’Adamo et al. (2014) and it was performed by Davies et al. (2014) and Jackson et al. (2015). This work was supported by the US Air Force Office of Scientific Research (grant AFOSR 9550-14-1-0147).
References
- Davies, F. K., Work, V. H., Beliaev, A. S. and Posewitz, M. C. (2014). Engineering limonene and bisabolene production in wild type and a glycogen-deficient mutant of Synechococcus sp. PCC 7002. Front Bioeng Biotechnol 2: 21.
- D'Adamo, S., Jinkerson, R. E., Boyd, E. S., Brown, S. L., Baxter, B. K., Peters, J. W. and Posewitz, M. C. (2014). Evolutionary and biotechnological implications of robust hydrogenase activity in halophilic strains of Tetraselmis. PLoS One 9(1): e85812.
- Jackson, S. A., Eaton-Rye, J. J., Bryant, D. A., Posewitz, M. C. and Davies, F. K. (2015). Dynamics of photosynthesis in a glycogen-deficient glgC mutant of Synechococcus sp. Strain PCC 7002. Appl Environ Microbiol 81(18): 6210-6222.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Davies, F. K., D’Adamo, S. and Posewitz, M. C. (2016). HPLC Analysis of Secreted Organic Acids. Bio-protocol 6(8): e1786. DOI: 10.21769/BioProtoc.1786.
Category
Biochemistry > Other compound > Acid
Microbiology > Microbial biochemistry > Other compound
Microbiology > Microbial metabolism > Other compound
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











