- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Calculation of Microorganism Lag Times as a Measure of Adaptative Capability between Different Growth Conditions
Published: Vol 6, Iss 3, Feb 5, 2016 DOI: 10.21769/BioProtoc.1727 Views: 6971
Reviewed by: Valentine V TrotterDisha SrivastavaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Glucose Starvation, Magnesium Ion Starvation, and Bile Stress Assays
Aryashree Arunima and Mrutyunjay Suar
Sep 20, 2021 2979 Views
Analysis of Heterocyst and Akinete Specific Glycolipids in Cyanobacteria Using Thin-layer Chromatography
Ritu Garg [...] Iris Maldener
Mar 20, 2022 2474 Views
Detecting Photoactivatable Cre-mediated Gene Deletion Efficiency in Escherichia coli
Yuta Koganezawa [...] Miki Umetani
Jun 5, 2023 1957 Views
Abstract
This protocol has been designed as a simple and efficient way to investigate microorganism adaptive capabilities (Enjalbert et al., 2015). It is performed using switch experiments in which cells are initially grown in the first condition (primary cultures), then rapidly switched to the second condition (secondary culture) without centrifugation or quenching. The measurement is based on the capacity of the secondary culture cells to resume growth. This protocol can be utilized for assessing metabolic or stress adaptation of microorganisms.
Keywords: LagMaterials and Reagents
- Sterile Erlen-Meyer flask per switch
- 0.45 µm filter per switch (Minisart 0.45 μm filter) (Sartorius AG)
- Plastic adapter per switch (Tube versilic 4 x 7 mm) (Saint-Gobain or equivalent)
- Sterile syringe 5 ml (Terumo Medical Corporation)
- Microorganism culture in condition 1
- Medium for growth condition 2
Equipment
- Shaker incubator
- Spectrophotometer and cuvettes
Procedure
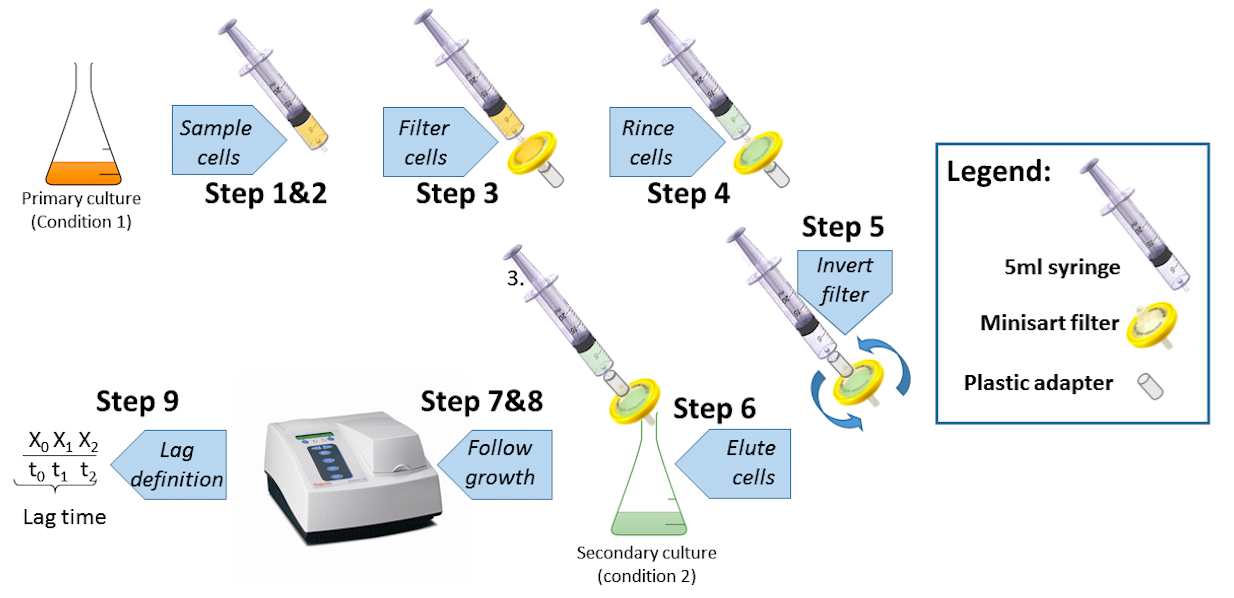
Figure 1. Switch experiments procedure
- In the incubator, pre-warm one empty 5 ml syringe, two 5 ml syringes filled with 3 ml condition 2 medium, the flask filled with 30 ml of condition 2 medium, and the filter with a plastic adapter on the nozzle. Sterility of the media has to be maintained at this step.
- With the empty syringe (needle inner diameter: 0.8 mm), sample cells from the primary culture in condition 1.
Notes:- The volume depends on two factors. (i) It has to be maximized so that the absorbance when eluted in the condition 2 flask can be reproducibly measured at step 6 (for example, 2 ml of E. coli culture at OD600 nm = 3 in the primary culture provide a measurable OD600 nm = 0.2 in the secondary culture). (ii) It has to be minimized to avoid clogging the filter at step 3.
- From step 2 onward, the manipulations have to be performed as rapidly as possible to minimize the culture perturbation (i.e., in less than 2 min). As a control, we suggest performing the same experiment by replacing condition 2 medium by condition 1 and ensuring that the initial growth rate in the secondary culture is equal to the growth rate in the primary culture.
- The volume depends on two factors. (i) It has to be maximized so that the absorbance when eluted in the condition 2 flask can be reproducibly measured at step 6 (for example, 2 ml of E. coli culture at OD600 nm = 3 in the primary culture provide a measurable OD600 nm = 0.2 in the secondary culture). (ii) It has to be minimized to avoid clogging the filter at step 3.
- Attach the filter to syringe filled with 3 ml of pre-warmed condition 2 medium and rinse the cells on filter (the flow-through is discarded).
- Replace the syringe by one of the two syringes filled with 3 ml of pre-warmed condition 2 medium and rinse the cells on the filter with medium 2.
- Attach the second 5 ml syringe filled with 3 ml of pre-warmed condition 2 medium on the other end of the filter using the plastic adapter.
- Elute the cells over the flask filled with 30 ml of condition 2 medium.
- Place the flask in the shaker incubator for 1 min to ensure homogeneity of the solution and measure the initial OD. Immediately place the flask back in the incubator.
- Monitor again the growth of the secondary culture when the cells reach their maximal growth rate on condition 2 (standardized by trial). For example, switching E. coli from a glucose to an acetate based mineral medium required measuring the OD at times 60 and 90 minutes after inoculation by spectrophotometry at OD600 nm (Enjalbert et al., 2015). The measure at 60 min allows to calculate the lag (see step 8), and the calculation of the growth rate between 60 and 90 min ensures that the cells reach their maximal growth rate (this maximum growth rate in condition 2 has to be previously determined).
- Use the following equation to calculate the lag time before maximal growth (see Figure 2 for justification):
(t1-tm) = (t1-t0)-ln(X1/X0)/μmax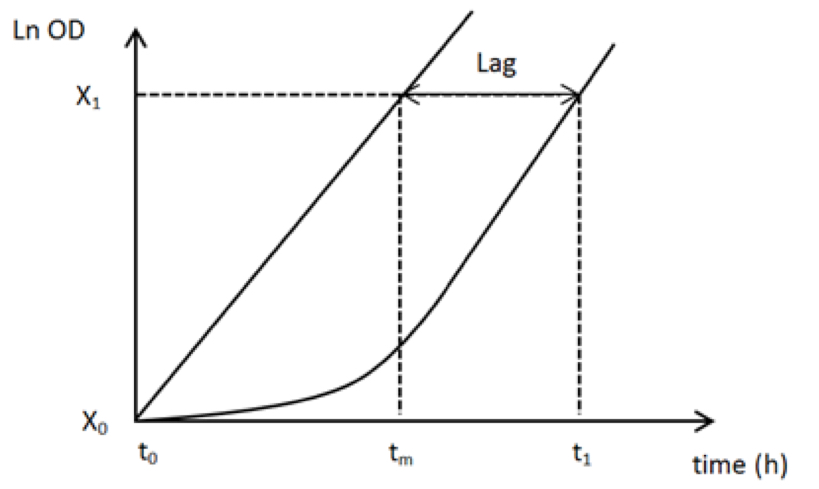
Figure 2. Theoretical growth profiles for the calculation of the lag in the switch experiments. tm is the theoretical time needed to increase the biomass from X0 to X1 if the growth rate is maximal (µmax) from t0. If there is a delay, X1 will be obtained at t1 (i.e., later than tm). From these elements, the lag can be determined as (t1-tm).
By definition,
Eq1: μmax = ln(X1/X0)/(tm-t0)
From which
Eq2: (tm-t0)= ln(X1/X0)/μmax
Since
Eq3: (t1-tm)= (t1-t0)-(tm-t0)
From Eq3 and Eq2,
Eq41: (t1-tm) = (t1-t0)-ln(X1/X0)/μmax
Acknowledgments
B. E. chair was supported by the INRA (Institut National de la Recherche Agronomique) and the INSA (Institut National des Sciences Appliquées) (Program <Chaire d’excellence>
References
- Enjalbert, B., Cocaign-Bousquet, M., Portais, J. C. and Letisse, F. (2015). Acetate exposure determines the diauxic behavior of Escherichia coli during the glucose-acetate transition. J Bacteriol 197(19): 3173-3181.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Enjalbert, B. (2016). Calculation of Microorganism Lag Times as a Measure of Adaptative Capability between Different Growth Conditions . Bio-protocol 6(3): e1727. DOI: 10.21769/BioProtoc.1727.
Category
Microbiology > Microbial physiology > Adaptation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link