- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantification of Low Molecular Weight Thiols in Arabidopsis
Published: Vol 6, Iss 1, Jan 5, 2016 DOI: 10.21769/BioProtoc.1704 Views: 7729
Reviewed by: Samik BhattacharyaMoritz BomerAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
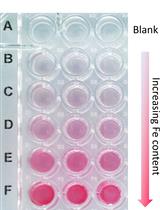
A Quick Method to Quantify Iron in Arabidopsis Seedlings
Chandan Kumar Gautam [...] Wolfgang Schmidt
Mar 5, 2022 3960 Views
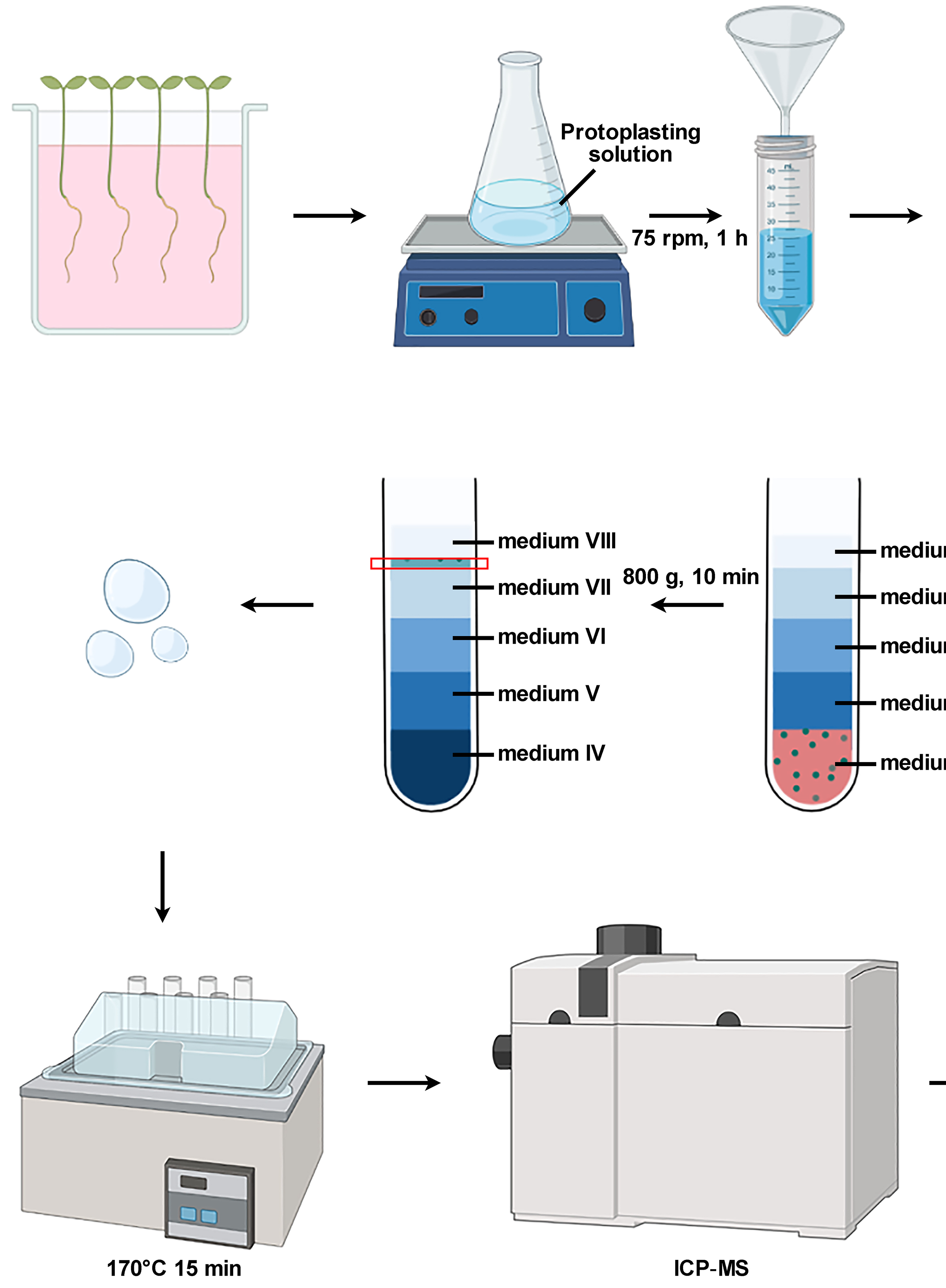
Isolation of Intact Vacuoles from Arabidopsis Root Protoplasts and Elemental Analysis
Chuanfeng Ju [...] Zhenqian Zhang
Mar 5, 2023 2102 Views
High-Performance Liquid Chromatography Quantification of Glyphosate, Aminomethylphosphonic Acid, and Ascorbate in Culture Medium and Microalgal Cells
Juan Manuel Ostera [...] Gabriela Malanga
Apr 5, 2025 1222 Views
Abstract
Low-molecular-weight (LMW) thiols are a class of highly reactive compounds due to their thiol moiety. They play important roles in the maintenance of cellular redox homeostasis, detoxification, and development. Monobromobimane (mBBr) is weakly fluorescent but selectively reacts with thiols to yield highly fluorescent thioethers (mBSR) products, which is especially useful for the quantification of LMW thiols. The stable mBSR products can be separated by high-performance liquid chromatography (HPLC) equipped with a fluorescent detector. The main cellular LMW thiols are L-cysteine, gamma-glutamylcysteine, and glutathione (GSH). The following protocol describes the extraction and quantification of L-cysteine, gamma-glutamylcysteine, and glutathione from Arabidopsis tissues as reported (Xiang and Oliver, 1998; Zhao et al., 2014; Wang et al., 2015) with minor revision. Modifications may be required if the HPLC system or the C18 column is different.
Keywords: CysteineMaterials and Reagents
- 1.5 ml microcentrifuge tube
- 0.2 μm nylon filter (Sigma-Aldrich, catalog number: Z259969 )
- 100 ml glass syringe
- Pipette tip
- Fresh Arabidopsis thaliana tissues (any plant part you want to test, 50-100 mg is sufficient)
- L-cysteine (Sigma-Aldrich, catalog number: 778672 )
- Gamma-glutamylcysteine (γ-Glu-Cys) (Sigma-Aldrich, catalog number: G0903 )
- L-Glutathione reduced (Sigma-Aldrich, catalog number: G4251 )
- Double distilled water (supplied by School of Life Sciences, University of Science and Technology of China)
- Double distilled water
- Hydrochloric Acid (HCl) (Sigma-Aldrich, catalog number: 435570 ) (see Recipes)
- 2-(N-Morpholino)ethanesulfonic acid hydrate (MES) (Sigma-Aldrich catalog number: M8250 ) (see Recipes)
- Ethylenediaminetetraacetic acid disodium salt dihydrate (EDTA.2Na.2H2O) (Sangon Biotech, catalog number: A100105 ) (see Recipes)
- Monobromobimane (mBBr) (Sigma-Aldrich, catalog number: 69898 ) (see Recipes)
- Trifluoroacetic acid (TFA) (Sigma-Aldrich, catalog number: 302031 ) (see Recipes)
- Acetonitrile (OCEANPAK, catalog number: Ac00030281 ) (see Recipes)
Equipment
- Analytic balance (Mettler-Toledo International Inc., model: ML104 )
- Mortar and Pestle
- Refrigerated microcentrifuge (Thermo Fisher Scientific, Eppendorf, model: 5424R )
- 37 °C degree incubator (Shanghai Jinghong Laboratory Instrument, model: GNP-9080 )
- Reverse-phase C18 column (5 µm, 110A, 150 x 4.6 mm) (Phenomenex, model: Gemini C18 column ) or equivalent, guard column (Phenomenex, SecurityGuard Standard, model: AJ0-7597 )
- HPLC equipment (Agilent Technologies, model: 1200 series )
- 200 μl pipette
- pH meter (Thermo Fisher Scientific, Mettler Toledo, model: FE20–FiveEasy PlusTM )
- Ultrasonic cleaner (Shanghai Sonxi Ultrasonic Instrument, model: DS-2510DT )
Software
- Suitable data collection and processing software (Agilent Technologies, model: 1200 ChemStation)
- Standard curve plotting (Microsoft Excel)
Procedure
- Preparation of samples and standards
- Fresh tissues are sampled into a 1.5 ml microcentrifuge tube and weighed with an analytical balance. 50-100 mg is sufficient for each biological sample.
- Samples are ground in the microcentrifuge tube with a mortar and a pestle with 2 volumes of 0.15 M HCl added by 200 μl pipette (for example, 100 mg tissue needs 200 μl).
- The homogenate is centrifuged at 12,000 x g, 4 °C, for 15 min and the supernatant is transferred into a new 1.5 ml microcentrifuge tube. This step can be repeated if necessary to remove insoluble substances.
- Prepare 10 mM stock solutions of L-cysteine, gamma-glutamylcysteine, and glutathione, and then prepare the following standards: 0, 50, 100, 200, 500, 1,000 μM of L-cysteine, gamma-glutamylcysteine, and glutathione by diluting the stocks with 0.15 M HCl.
- 100 μl of the supernatant from step A3 (or standards) is transferred to 1.5 ml microcentrifuge tubes containing 2 μl of 0.5 M EDTA, 2.6 μl of 300 mM mBBr in acetonitrile, and 100 μl of 1.75 M MES (pH 7.4). The mixture was incubated in the dark at 37 °C for 1 h to allow the derivatization reaction to completed.
- The mixture is centrifuged as in step A3 for 5 min before quantification.
- Fresh tissues are sampled into a 1.5 ml microcentrifuge tube and weighed with an analytical balance. 50-100 mg is sufficient for each biological sample.
- Quantification by HPLC (refer to Agilent 1200 HPLC ChemStation Operation)
Select “Edit entire method” from “Method” menu and set parameters according to the following text. For quantification, 50 μl sample from step A6 is injected into the sample chamber of the HPLC system and separated using a reverse-phase C18 column and a flow rate of 0.8 ml/min on an Agilent 1200 HPLC system. Solvents A and B are used to elute the fluorescent derivatives with the gradient shown in the table below. The fluorescent derivatives (mBSR) are detected using fluorescence detector with excitation wavelength at 260 nm and emission at wavelength 474 nm.
Table 1. HPLC elution programTime (min) A (%) B (%) Rate of flow (ml/min) 0.0 90 10 0.8 0.3 85 15 0.8 14.0 80 20 0.8 15.0 0 100 0.8 19.0 0 100 0.8 20.0 90 10 0.8 24.0 90 10 0.8 - Analyze standards and samples using the HPLC program above with at least 3 replicates.
- Peak areas are integrated using the ChemStation software.
Select “Data analysis” from the “View” menu to enter the picture data analysis.
Select “Load signal” from the “File” menu to select your data file.
Select “Integrate” from the “Integration” menu, and then the data is integrated.
- Analyze standards and samples using the HPLC program above with at least 3 replicates.
- Calculations
- Prepare standard curves by plotting the concentration (μM) of the standards (Y-axis) vs the peak areas (X-axis) and add a trendline.
- Determine the peak area for each sample and determine concentration using the trendline. Concentration (μM) = a * peak area + b (a and b are already calculated in the trendline).
- Prepare standard curves by plotting the concentration (μM) of the standards (Y-axis) vs the peak areas (X-axis) and add a trendline.
Recipes
- 0.15 M HCl (stored at room temperature)
1.27 ml HCl (37%)
100 ml double distilled water
Filtered through 0.2 μm nylon filter using 100 ml syringe - 1.75 M MES (pH 7.4) (stored at 4 °C)
3.41 g MES
10 ml double distilled water
Adjust pH to 7.4 using NaOH and filter through 0.2 μm nylon filter using 100 ml syringe - 0.5 M EDTA.2Na.2H2O (stored at room temperature)
18.6 g EDTA.2Na.2H2O
Dissolved in double distilled water, pH adjusted to 8.0 with NaOH, and volume brought to100 ml
Filtered through 0.2 μm nylon filter using 100 ml syringe - 300 mM monobromobimane (stored at -20 °C)
0.025 g monobromobimane
307.3 μl acetonitrile
12,000 x g, 4 °C, 30min to remove insoluble substances - Solvent A: 0.1% (v/v) trifluoroacetic acid (HPLC grade) (freshly prepared)
1 ml trifluoroacetic acid
999 ml double distilled water
Filtered through 0.2 μm nylon filter using 100 ml syringe - Solvent B: 90% (v/v) acetonitrile (HPLC grade) (freshly prepared)
900 ml 100% acetonitrile
100 ml 0.1% (v/v) trifluoroacetic acid
Filtered through 0.2 μm nylon filter using 100 ml syringe
Solvent A and Solvent B need degas in the ultrasonic cleaner for 30 min with loose lid - 10 mM L-cysteine (stored at -20 °C)
0.0606 g L-cysteine
50 ml 0.15 M HCl - 10 mM gamma-glutamylcysteine (stored at -20 °C)
0.1251 g gamma-glutamylcysteine
50 ml 0.15 M HCl - 10 mM glutathione (stored at -20 °C)
0.1537 g glutathione
50 ml 0.15 M HCl - Standards (stored at -20 °C)
0 μM 50 μM 100 μM 200 μM 500 μM 1,000 μM 10 mM 0 μl 5 μl 10 μl 20 μl 50 μl 100 μl 0.15 M HCl 1,000 μl 995 μl 990 μl 980 μl 950 μl 900 μl
Acknowledgments
This protocol was modified from previous work described by Fahey and Newton (1987).
References
- Fahey, R. C. and Newton, G. L. (1987). Determination of low-molecular-weight thiols using monobromobimane fluorescent labeling and high-performance liquid chromatography. Methods Enzymol 143: 85-96.
- Wang, Z., Mao, J. L., Zhao, Y. J., Li, C. Y. and Xiang, C. B. (2015). L-Cysteine inhibits root elongation through auxin/PLETHORA and SCR/SHR pathway in Arabidopsis thaliana. J Integr Plant Biol 57(2): 186-197.
- Xiang, C. and Oliver, D. J. (1998). Glutathione metabolic genes coordinately respond to heavy metals and jasmonic acid in Arabidopsis. Plant Cell 10(9): 1539-1550.
- Zhao, Q., Wu, Y., Gao, L., Ma, J., Li, C. Y. and Xiang, C. B. (2014). Sulfur nutrient availability regulates root elongation by affecting root indole-3-acetic acid levels and the stem cell niche. J Integr Plant Biol 56(12): 1151-1163.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Miao, Z., Wang, Z. and Xiang, C. B. (2016). Quantification of Low Molecular Weight Thiols in Arabidopsis. Bio-protocol 6(1): e1704. DOI: 10.21769/BioProtoc.1704.
Category
Plant Science > Plant biochemistry > Other compound
Biochemistry > Other compound > Thiol
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









