- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparation of Recombinant Galectin-3 for Cancer Studies
Published: Vol 6, Iss 1, Jan 5, 2016 DOI: 10.21769/BioProtoc.1696 Views: 9369
Reviewed by: HongLok LungClara Lubeseder-MartellatoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
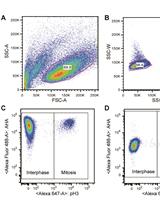
Measuring Protein Synthesis during Cell Cycle by Azidohomoalanine (AHA) Labeling and Flow Cytometric Analysis
Koshi Imami and Tomoharu Yasuda
Apr 20, 2019 9781 Views
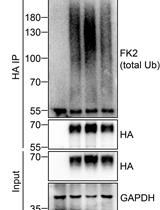
Analysis of the Ubiquitination and Phosphorylation of Vangl Proteins
Di Feng [...] Bo Gao
Oct 20, 2022 3463 Views
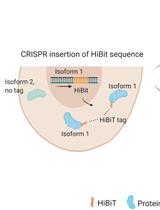
Isoform-specific, Semi-quantitative Determination of Highly Homologous Protein Levels via CRISPR-Cas9-mediated HiBiT Tagging
Kristina Seiler [...] Mario P. Tschan
Jul 20, 2023 2549 Views
Abstract
Galectin-3 is a member of a class of proteins termed Galectins, characterized by their ability to bind glycans containing β-galactose (Cummings and Liu, 2009). Galectin-3 binds preferentially to proteoglycans terminating with N-acetyllactosamine (LacNAc) chains (i.e., tandem repeats of galactose) (Newlaczyl and Yu, 2011). Galectin-3 is unique among the galectins in its chimeric structure. It shares a conserved carbohydrate recognition domain (CRD) with the other galectins, but has a long amino-terminal tail that is thought to be involved in protein aggregation. It can also form homodimers through its CRD (Cummings and Liu, 2009). Galectin-3 has been found to have diverse functions in tumorigenesis including: signaling, apoptosis inhibition, immune suppression, cell growth, and metastasis among others. Galectin-3 is frequently upregulated in cancers (Nangia-Makker et al., 2008). Its function largely depends on its expression and localization properties (Newlaczyl and Yu, 2011). Because of its many roles in cancer-associated processes, establishing a method for Galectin-3 production is valuable for further study of its functions in cancer. Here, we describe how Galectin-3 purification was achieved by cloning of the human Galectin-3 gene into pGEX-2T vector containing the gene for glutathione-S-transferase (GST) upstream of its cloning site. The Galectin-3 gene was cloned into this vector via restriction digests of both the plasmid and the Galectin-3 gene by restriction enzymes BamHI and EcoRI, followed by ligation of the two fragments. The resulting plasmid was then used to transform BL21, an Escherichia coli (E. coli) strain specialized for protein expression. Finally, we discuss how the GST fusion protein was isolated and the recombinant Galectin-3 protein was further purified from the GST.
Keywords: CancerPart I. Protocol for GST-Gal3 purification (Harper and Speicher, 2011)
Materials and Reagents
- Microcentrifuge tubes (Denville Posi-Click Tubes, catalog number: C-2170 )
- 15 ml conical tubes (Thermo Fisher Scientific, Corning CentriStar, catalog number: 05-538-59A )
- 10 ml serological pipettes (Thermo Fisher Scientific, Corning Costar, catalog number: 4488 )
- One Shot® BL21 StarTM (DE3) Chemically Competent E. coli (Thermo Fisher Scientific, catalog number: C6010-03 )
- pGEX 2T-Gal3 vector (kindly provided by Dr. W. Stallcup, UCSD, CA)
- L-broth (Thermo Fisher Scientific, catalog number: BP1426-2 )
- Glutathione-Agarose Beads (Thermo Fisher Scientific, PierceTM, catalog number: 16100 )
- Protease Inhibitor Tablets, EDTA-free (Thermo Fisher Scientific, PierceTM, catalog number: 88665 )
- Ice
- Purified Water
- 10x PBS, diluted to 1x (Thermo Fisher Scientific, Hyclone, catalog number: BP3994 )
- Isopropyl-beta-D-thiogalactopyranoside (IPTG) (Gold Biotechnology, catalog number: 12481C25 )
- Tris-Cl (pH 7.5)
- 150 mM NaCl
- 0.75% CHAPS powder (AG. Scientific, catalog number: C-1019 )
- 0.75% CHAPS buffer (see Recipes)
Equipment
- Microfuge (Beckman Coulter, model: Microfuge 16 )
- 37 °C shaker (FormaOrbital Shaker)
- Shaker (GMI, Lab Companion, model: SK-300 Shaker )
- Rotating Platform (24 rpm) (Benchmark model: MiniMixer )
- Vortex (Thermo Fisher Scientific, catalog number: 12-812 )
Note: Currently, it is “LabX, catalog number: 12-812”. - Centrifuge (Beckman Coulter, model: Avanti J-20 XP )
- Centrifuge rotor (Beckman Coulter, model: JLA-16.250 )
- Centrifuge (Beckman Coulter, model: Allegra 25R )
- Centrifuge rotor (Beckman Coulter, model: TS-5.1-500 )
- 250 ml plastic centrifuge bottles (Sigma-Aldrich, Nalgene®, model: B1033 )
- 2 L flask (Corning, Pyrex®, model: No. 4980 )
- 10 ml serological pipettes (Corning, Costar®, catalog number: 4488 )
Procedure
- Transform BL21 bacteria with pGEX-2T-Gal3 expression vector, pick colonies and verify plasmid DNA with appropriate restriction digests (BamHI and EcoRI). The size of the plasmid is 5,697 bp; the size of the Galectin-3 insert is 753 bp (see map, Supplemental Figure 1).
- Use one colony to inoculate 3 ml clean, autoclaved Luria-broth and grow overnight at 37 °C at 250 rpm until OD600=1.0.
- In a 2 L flask, add 1,000 ml water. Measure 25 g Luria Broth and add to water in the flask. Autoclave before use (see step 2).
- Add 0.3 ml of the transformed bacteria to Luria Broth.
- Grow at 37 °C at 250 rpm for 4-5 h until OD600=0.6.
- Next, induce bacteria with 0.5 mM IPTG (119.2 mg) and allow growth on a shaker at room temperature at 200 rpm overnight for 16-18 h.
- Spin down bacteria into four, 250 ml plastic centrifuge bottles at 5,000 rpm for 15 min at 4 °C until pelleted. Discard supernatant. Place pellets on ice. (If you wish to stop here, the pellets may be stored at -80 °C until lysis. Prior to lysis, warm frozen pellets in 37 °C water bath for 10 min.)
- On ice, prepare CHAPS lysis buffer solution.
- Add two protease inhibitor tablets to 50 ml CHAPS lysis buffer solution and dissolve the tablets into the buffer by vortexing.
- On ice, add 12 ml CHAPS lysis buffer solution to each pellet.
- Break up the pellet by pipetting and vortexing until mixture is smooth. Vortex first to detach pellet from the wall of the centrifuge bottle, then use a 10 ml serological pipette to further disrupt the pellet by pipetting up and down to form a homogenous solution.
- Allow lysis to continue on ice for 30 min. Vortex every five minutes for the duration of the lysis (six times total).
- While bacterial lysis is taking place, add 2 ml glutathione-agarose bead slurry to 13 ml of purified water in 15 ml conical tube. Mix by inverting tube 2 to 3 times. Allow beads to swell for thirty minutes on ice.
- Distribute the bacterial lysate to microfuge tubes (about 1.5 ml lysate per tube, approximately 30 tubes). Alternatively, the lysate can be placed into a 50 ml centrifuge tube.
- Spin microfuge tubes at 14,000 rpm at 4 °C for 15 min. Supernatant should be clear. If using 50 ml centrifuge tube, spin at 3,500 rpm for 15 min at 4 °C. It is recommended, then, that the resulting supernatant be transferred to microfuge tubes and spun at 14,000 rpm for 10 additional minutes.
- Using a micropipette, separate supernatant from the pellets. Discard pellets.
- Pool supernatant from all microfuge tubes into two15 ml conical tubes.
- Gently aspirate excess water from swelled agarose-beads (beads should be settled at bottom of vial; if not centrifuge briefly for 10-20 sec without exceeding 1,000 rpm as high speeds can damage the beads).
- Add 10 ml CHAPS lysis buffer to beads to equilibrate. Mix by inverting the tube several times. Centrifuge briefly without exceeding 1,000 rpm.
- Gently aspirate excess buffer from the agarose beads.
- Add ~1 ml CHAPS lysis buffer to the agarose beads so that bead volume and buffer volume are equal (50% v/v). Mix beads and buffer by inverting the tube several times.
- Add 1 ml of the bead/buffer solution to each 15 ml conical tube containing bacterial lysis supernatant. Tip of pipette should be cut for beads to fit through.
- Mix beads and lysis supernatant by inverting the tube several times.
- Incubate overnight at 4 °C on slowly rotating platform (24 rpm).
- The next day, spin down supernatant and beads briefly without exceeding 1,000 rpm for ~10 sec at 4 °C to settle beads. Remove supernatant and save as it may still contain some of your protein and may be useful if further analysis is necessary.
- Wash GST-protein-bound beads by adding 14 ml 1x PBS to each tube. Mix by inverting. If beads are stuck at the bottom of the tube, gently flick the tube with your finger to mobilize the beads.
- Centrifuge briefly without exceeding 1,000 rpm to allow beads to settle. Remove 1x PBS, and wash again with another 14 ml 1x PBS.
- Remove 1x PBS. Beads should contain your GST-fusion protein.
- Assess the integrity and yield of your rGST-Gal3 protein with a Coomassie stained 12% polyacrylamide SDS-PAGE gel and/or Western blot (see Figure 2).
- Store beads in 50% v/v CHAPS buffer solution (~1 ml beads, ~1 ml CHAPS buffer) at 4 °C until further use or proceed with thrombin cleavage to separate Galectin-3.
Recipes
- 0.75% CHAPS buffer
1.2 g (30 mM) Tris-Cl (pH 7.5)
2.2 g (150 mM) NaCl
1.9 g (0.75%) CHAPS powder
Fill to 250 ml with water
Part II. Protocol for thrombin-mediated cleavage of GST from GST-Gal3
Materials and Reagents
- Microfuge tubes (DENVILLE SCIENTIFIC, catalog number: C-2170 )
- 375 U Thrombin (Merck Millipore Corporation, Calbiochem®, catalog number: 605195 )
- Glutathione-agarose beads bound with rGST-Gal3
- p-aminobenzamidine agarose beads [50% (V/V)] (Sigma-Aldrich, catalog number: A7155 )
- Tween-20 (Thermo Fisher Scientific, Fisher Bioreagent, catalog number: BP337-500 )
- Ice
- 10x PBS, diluted to 1x (Thermo Fisher Scientific, Hyclone, catalog number: BP3994 )
- Commercial Galectin-3 (Peprotech, catalog number: 450-38 , 50 ng)
- Anti-galectin-3 primary antibody (1:800 in PBST with 2.5% BSA, Santa Cruz, 20157)
- Chicken anti-rabbit secondary antibody (1:10,000 in PBST with 2.5% BSA, Santa Cruz, sc-2955)
- PBST (0.1% Tween 20) (see Recipes)
Equipment
- Rotating Platform (24 rpm) (Benchmark Scientific, model: MiniMixer )
- Microfuge (Beckman Coulter, model: Microfuge 16 )
Procedure
- On ice, add 500 μl PBST to 100 μl of the 50% v/v rGST-Gal3 bound agarose beads in CHAPS buffer in microfuge tubes (600 µl total volume/tube).
- On ice, prepare a thrombin solution by diluting 375 U of thrombin into 800 µl PBST.
- Add 400 µl of thrombin solution to each microfuge tube for a final volume of 1 ml per microfuge tube.
- Place tubes on rotating platform (24 rpm) and allow incubation overnight at room temperature.
- The next day, wash p-aminobenzamidine beads in 1x PBS by adding necessary amount of p-aminobenzamidine slurry (250 µl bead slurry per 100 µl rGST-Gal3 bound agarose beads) into a 15 ml centrifuge tube then filling the tube to 15 ml with 1x PBS. Mix by inverting then centrifuge briefly without exceeding 1,000 rpm.
- Remove 1x PBS then repeat wash with fresh 1x PBS.
- Remove PBS, then add enough fresh 1x PBS so that beads and 1x PBS form a 50% v/v solution.
- Centrifuge the bead/thrombin solution at 3,500 rpm for 1 min in tabletop microfuge. Remove supernatant and place into fresh 1.5 ml microfuge tubes.
- Add 250 µl p-aminobenzamidine agarose bead slurry to each supernatant-containing microfuge tube in order to remove thrombin from the solution.
- Incubate the p-aminobenzamidine beads with supernatant for 1.5 h at room temperature on rotating platform (24 rpm).
- Microfuge tubes at 3,500 rpm for ~30 sec.
- Collect supernatant. Proteins will be in supernatant while thrombin, attached to the beads, will be eliminated with the pellet.
- Assess supernatant protein composition with a Coomassie-stained SDS-PAGE and Western blot (Figures 1-2).

Figure 1. Galectin-3 is successfully cleaved from GST through incubation with thrombin protease. Lanes 1-3 show increasing volumes of Galectin-3 (1 µl, 2 µl, 4 µl; approximately 150, 300, 600 ng) cleaved from GST through thrombin treatment. Lane 5 shows commercial Galectin-3. Samples were analyzed by SDS-PAGE on 12% polyacrylamide gel, followed by Western blot. Immunoblot was performed with Rabbit, anti-galectin-3 primary antibody (1:800 in PBST with 2.5% BSA), and chicken anti-rabbit secondary antibody (1:10,000 in PBST with 2.5% BSA).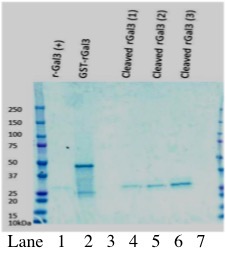
Figure 2. Analysis of thrombin cleavage of rGST-Gal3 by Coomassie-stained gel electrophoresis. Lane 1 shows positive control human Galectin-3 at ~25 kDa. Lane 2 shows GST-Gal3 without cleavage by thrombin. The size of the GST-Galectin3 fusion protein is approximately 50 kDa. Lanes 4-6 show increasing amounts of Galectin-3 cleaved from GST (0.94 µg, 1.88 µg, 3.75 µg as determined by Bradford Assay).
Recipes
- PBST (0.1% Tween 20)
Add 100 µl of Tween-20 per 100 ml of 1x PBS
Acknowledgments
We thank Dr. William Stallcup (Sanford-Burnham Medical Research Institute; Cancer Center; La Jolla, CA USA) for providing the GST-Gal3 expression plasmids. This work was supported by R01 grant CA163722 from the NIH (to EGVM).
References
- Cummings, R. D., and Liu, F. T. (2009). Galectins. In: Varki, A., Cummings, R. D., Esko, J. D. , Freeze, H. H., Stanley, P., Bertozzi, C. R., Hart, G. W. and Etzler, M. E. (eds). Essentials of Glycobiology (2nd ed.). Cold Spring Harbor.
- Harper, S. and Speicher, D. W. (2011). Purification of proteins fused to glutathione S-transferase. Methods Mol Biol 681: 259-280.
- Nangia-Makker, P., Balan, V. and Raz, A. (2008). Regulation of tumor progression by extracellular galectin-3. Cancer Microenviron 1(1): 43-51.
- Newlaczyl, A. U. and Yu, L. G. (2011). Galectin-3--a jack-of-all-trades in cancer. Cancer Lett 313(2): 123-128.
- Wen, Y., Makagiansar, I. T., Fukushi, J., Liu, F. T., Fukuda, M. N. and Stallcup, W. B. (2006). Molecular basis of interaction between NG2 proteoglycan and galectin-3. J Cell Biochem 98(1): 115-127.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tyler, K., Lee, S. and Van Meir, E. G. (2016). Preparation of Recombinant Galectin-3 for Cancer Studies. Bio-protocol 6(1): e1696. DOI: 10.21769/BioProtoc.1696.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Cancer Biology > General technique > Tumor microenvironment > Extracellular matrix
Biochemistry > Protein > Expression
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










