- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation and Flow-cytometric Analysis of Mouse Intestinal Crypt Cells
Published: Vol 5, Iss 21, Nov 5, 2015 DOI: 10.21769/BioProtoc.1635 Views: 17023
Reviewed by: HongLok LungKristina Y. AguileraAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Isolation of Stem Cells, Endothelial Cells and Pericytes from Human Infantile Hemangioma
Lan Huang and Joyce Bischoff
Jan 20, 2020 4966 Views
Low-viscosity Matrix Suspension Culture for Human Colorectal Epithelial Organoids and Tumoroids
Tao Tan [...] Oliver M. Sieber
Apr 20, 2022 4086 Views
Thrombopoietin-independent Megakaryocyte Differentiation of Hematopoietic Progenitor Cells from Patients with Myeloproliferative Neoplasms
Chloe A. L. Thompson-Peach [...] Daniel Thomas
Jan 20, 2023 2379 Views
Abstract
The intestinal epithelial layer forms tubular invaginations into the underlying connective tissue of the lamina propria. These structures, termed crypts, are the basic functional unit of the intestine. Colon crypts and the surrounding lamina propria house different cell types, including epithelial cells, stem cells, enterocytes, goblet cells, as well as cells of the innate and adaptive immune systems (Clevers, 2013; Mowat and Agace, 2014). Here we describe a technique for the isolation of mouse intestinal crypt cells as well as their characterization by flow cytometry analysis (FACS) (Del Reino et al., 2012).
Keywords: Flow cytometryMaterials and Reagents
- Falcon 50 ml conical tubes (VWR International, catalog number: 21008-240 )
- Falcon 15 ml conical tubes (VWR International, catalog number: 21008-216 )
- 10 cm Petri dishes (Corning, Falcon®, catalog number: 353046 )
- Gauze, mesh size approximately 100 μm (Tegosa, catalog number: 10018 )
- Cell strainers (70 μm) (Corning, Falcon®, catalog number: 352350 )
- Cell strainers (40 μm) (Corning, Falcon®, catalog number: 352340 )
- 25G needle (Premier Healthcare & Hygiene, BD Microlance, catalog number: 300600 )
- 5 ml syringe (BD, catalog number: 309649)
- 5 ml round-bottom flow cytometry tube (BD Biosciences, catalog number: 352063 )
- 8- to 10-week old mice (Mus musculus) (male or female) (we used the C57BL/6 strain)
- Hank’s balanced salt solution medium (HBSS) (Life Technologies, Gibco®, catalog number: 2420-091 )
- Penicillin-Streptomycin Solution 100x (Biowest, catalog number: L0022 )
- Ethylenediaminetetraacetic acid (EDTA) (AppliChem GmbH, Panreac, catalog number: 13102/ 131026 )
- Fetal bovine serum (FBS) (Life Technologies, catalog number: GCS0103-500 )
- APC-conjugated anti-CD45 antibody (Beckman Coulter, catalog number: 732158 )
- FITC-conjugated anti-CD4 antibody (Beckman Coulter, catalog number: 731999 )
- PE-conjugated anti-Ly6G antibody (Beckman Coulter, catalog number: 732487 )
- Biotinylated anti-F480 antibody (eBioscience, catalog number: 13-4801 )
- PeyC7-conjugated anti-CD8 antibody (Biolegend, catalog number: 100721 )
- ECD-conjugated streptavidin (Beckman Coulter, catalog number: IM3326 )
- Dispase II (Life Technologies, catalog number: 17105-041 )
Note: Currently, it is “Thermo Fisher Scientific, GibcoTM, catalog number: 17105-041”. - Phosphate-buffered saline (PBS) (see Recipes)
- Disaggregation medium (see Recipes)
Equipment
- Dissection equipment (forceps & scissors)
- Scalpel or razor blade
- 37 °C chamber (Thermo Fisher Scientific, HeraeusTM)
- Refrigerated centrifuge (Thermo Fisher Scientific, HeraeusTM, model: MegafugeTM 2.012 )
- Neubauer chamber cell counter
- Orbital shaking platform (Kühner AG, Lab-Shaker)
- CO2 chamber
- Cytomics fc500 flow cytometer (Beckman Coulter)
- Microscope (ZEISS, model: Axiovert 40CFL )
Procedure
- Isolation of intestinal crypt cells
- Sacrifice mice in a CO2 chamber by trained personnel. Perform the inhalation of CO2 from a pressurized tank in a chamber constructed of clear material (e.g., Plexiglas) (a standard size mouse cage may contain no more than 5 mice) followed by cervical dislocation. According to the Institutional Animal Care and Use Committee, CO2 must be delivered in a controlled manner, at a low-flow rate of 10-30% volume displacement per minute.
- After swabbing the abdomen with alcohol, make a ventral incision. Use forceps and scissors to carefully remove the entire colon (Figure 1). Cut from the cecum to the anus (between 8-10 cm depending on the mouse size), and place it in a 10 cm Petri dish with 15 ml cold PBS to cover the colon.
- Hold the distal part of the colon with the forceps and flush to remove feces with 5-10 ml cold sterile PBS using a 25 G needle attached to a 5 ml syringe. After flushing out feces, place the cleaned colon in a new Petri dish with 10 ml cold PBS.
- Make a longitudinal incision in the colon and rinse again with cold PBS to remove remaining feces.
- Place the open colon in cold HBSS medium on ice until all colons for the experiment have been extracted as in steps A1-4. For good cell yield, we use three mice per test condition.
- Incubate all colons in 5 ml HBSS containing 100 U/ml penicillin and 0.1 mg/ml streptomycin (15 min, room temperature).
- Mince the colon tissue into small pieces of approx. 2 mm x 2 mm on a Petri dish, with the help of a scalpel. Place the small pieces in a 50 ml Falcon tube and incubate in 5 ml of 8 mM EDTA in HBSS (160 μl 0.5 M EDTA in 10 ml HBSS; 15 min, 37 °C).
- Remove the medium and add 5 ml fresh cold HBSS to the colon pieces in the bottom of the falcon tube.
- Dissociate crypt epithelial cells by vigorous hand shaking to obtain a supernatant enriched in crypts. Decant supernatant from tissue debris into a new Falcon 50 ml tube. Add fresh cold HBSS to the sample and repeat the shaking step. Repeat this procedure 3-5 times to increase crypt enrichment.
- To confirm crypt separation, visualize a 20 μl supernatant sample under the microscope (Figure 2).
- Centrifuge supernatants (200 x g, 15 min, 4 °C).
- Remove supernatant. To the pellets, add 5 ml of disaggregation medium, transfer them to a 15 ml Falcon tube and incubate (30 min, 37 °C at 2,500 rpm with orbital shaking) to obtain single-cell suspensions.
- Add 250 μl FBS to the medium (5% final concentration) after disaggregation to terminate the dispase reaction.
- Filter cells sequentially through 100 μm gauze, then 70 and 40 μm cell strainers.
- Collect the cells by centrifugation (150 x g, 10 min, 4 °C), resuspend in 1.5 ml staining buffer (2% FBS in PBS) and count live cells in a cell counter. The expected cell yield from three mice is approx. 2 x 107 cells.
- Stain 1 x 106 intestinal crypt cells with fluorescence-labeled antibodies to the cell markers of interest using standard FACS analysis procedures, and analyze on a flow cytometer.
- Sacrifice mice in a CO2 chamber by trained personnel. Perform the inhalation of CO2 from a pressurized tank in a chamber constructed of clear material (e.g., Plexiglas) (a standard size mouse cage may contain no more than 5 mice) followed by cervical dislocation. According to the Institutional Animal Care and Use Committee, CO2 must be delivered in a controlled manner, at a low-flow rate of 10-30% volume displacement per minute.
- Example of flow cytometry analysis
- Incubate 1 x 106 intestinal crypt cells in 200 μl staining buffer (2% FBS in PBS) with fluorescently labeled anti-CD45, -CD4, -F4/80, -CD8 and -Ly6G antibodies (20 min, 4 °C, protected from light). Dilute the antibodies in PBS. 4-CD45-APC diluted 1/100; CD4-FITC diluted 1/100; F4/80-Biotina diluted 1/200; CD8-PeyC7 diluted 1/200 and Ly6G-PE diluted 1/100.
- Wash twice with 150 μl staining buffer, centrifuge (150 x g, 5 min, 4 °C) and discard supernatant.
- Incubate cells with streptavidin diluted 1:50 (20 min, 4 °C, protected from light).
- Repeat washing step B2.
- Resuspend cells in 200 μl staining buffer and transfer into round-bottom flow cytometry tubes with 220 μl staining buffer.
- Analyze staining with the Kaluza program, gating all positive cells for each antibody in CD45+ cells (Figure 3).
- Incubate 1 x 106 intestinal crypt cells in 200 μl staining buffer (2% FBS in PBS) with fluorescently labeled anti-CD45, -CD4, -F4/80, -CD8 and -Ly6G antibodies (20 min, 4 °C, protected from light). Dilute the antibodies in PBS. 4-CD45-APC diluted 1/100; CD4-FITC diluted 1/100; F4/80-Biotina diluted 1/200; CD8-PeyC7 diluted 1/200 and Ly6G-PE diluted 1/100.
Representative data
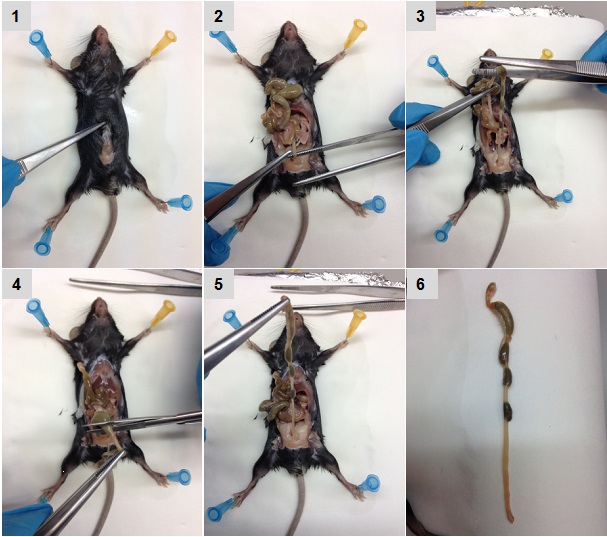
Figure 1. Sequential images of colon isolation. Pictures were acquired sequentially as described in the text.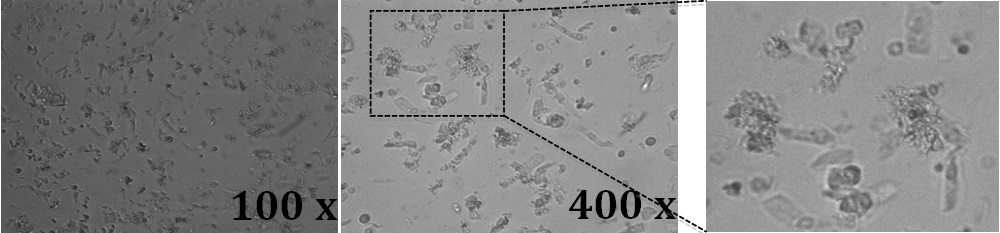
Figure 2. Light microscope images of mouse colon crypts. 20 μl of a suspension containing the crypts were placed directly on glass objective slides together with a glass coverslip. Images were acquired
with a 10x and a 40x objective and a digital camera attached to the microscope. A higher magnification of the field in the dotted squared is shown in the right panel. Crypts are in single or cluster elongated columnar and spherical cells.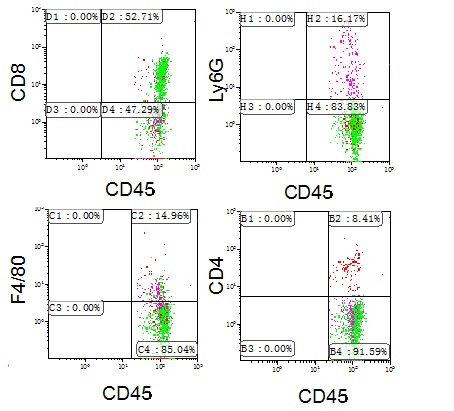
Figure 3. Characterisation of CD45+ cells in mouse colon. Colon crypt cells from mouse were stained with anti-CD45, -CD4, CD8, -Ly6G and -F4/80 antibodies and the percentage of positive cells analysed by flow cytometry. Representative profiles are shown. Other FACS analysis and quantification image examples are published in Cancer Research (Del Reino et al., 2014). For details, refer to Figure 5B.
Recipes
- Phosphate-buffered saline (PBS)
137 mM NaCl
2.7 mM KCI
4.3 mM Na3PO4
1.4 mM K2HPO4 - Disaggregation medium
HBSS
0.4 mg/ml dispase
Penicillin/streptomycin: 100 U/ml penicillin and 0.1 mg/ml streptomycin
Acknowledgments
This work was supported by The Spanish Ministry of Economy and Competiveness (MINECO) (SAF2013-45331-R) and La Marató TV3 Foundation (82031).
References
- Clevers, H. (2013). The intestinal crypt, a prototype stem cell compartment. Cell 154(2): 274-284.
- Del Reino, P., Alsina-Beauchamp, D., Escos, A., Cerezo-Guisado, M. I., Risco, A., Aparicio, N., Zur, R., Fernandez-Estevez, M., Collantes, E., Montans, J. and Cuenda, A. (2014). Pro-oncogenic role of alternative p38 mitogen-activated protein kinases p38gamma and p38delta, linking inflammation and cancer in colitis-associated colon cancer. Cancer Res 74(21): 6150-6160.
- Mowat, A. M. and Agace, W. W. (2014). Regional specialization within the intestinal immune system. Nat Rev Immunol 14(10): 667-685.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Alsina-Beauchamp, D., Reino, P. D. and Cuenda, A. (2015). Isolation and Flow-cytometric Analysis of Mouse Intestinal Crypt Cells. Bio-protocol 5(21): e1635. DOI: 10.21769/BioProtoc.1635.
Category
Cancer Biology > Cancer stem cell > Cell biology assays > Cell isolation and culture
Stem Cell > Adult stem cell > Intestinal stem cell
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link