- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Bioimaging Pipeline to Show Membrane Trafficking Regulators Localized to the Golgi Apparatus and Other Organelles in Plant Cells
Published: Vol 5, Iss 17, Sep 5, 2015 DOI: 10.21769/BioProtoc.1583 Views: 10510
Reviewed by: Arsalan DaudiRenate WeizbauerAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Live Leaf-Section Imaging for Visualizing Intracellular Chloroplast Movement and Analyzing Cell–Cell Interactions
Yuta Kato [...] Mitsutaka Taniguchi
Aug 5, 2025 2376 Views

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1723 Views
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 791 Views
Abstract
The plant Golgi apparatus is composed of numerous stacks of cisterna, designated as cis, medial, and trans Golgi cisternae; these stacks move within the cytoplasm along the actin cytoskeleton. Cis cisternae receive secretory products from endoplasmic reticulum (ER) and they subsequently progress through the stack to the trans cisternae, where they are sorted to other destinations, including cell wall, plasma membrane (PM), vacuoles, and chloroplasts. In addition, the plant Golgi apparatus plays a role of glycosylating proteins as well as synthesizing cell wall polysaccharides, such as hemicelluloses and pectins. This protocol describes procedures for imaging fluorescently-tagged proteins localized to the plant Golgi apparatus of Arabidopsis seedlings using confocal laser microscopy (CLSM), total internal reflection fluorescence microscope (TIRF), and immunogold labeling of high-pressure frozen/freeze substituted samples by transmission electron microscopy (TEM). We particularly focus on long-term time lapse imaging and protein localization in subdomains within the Golgi. This protocol can be also used for other organelles, tissues, and plant species.
Materials and Reagents
- ½X Murashige and Skoog (MS) medium (Sigma-Aldrich, catalog number: M5519 )
- Bacto Agar [Becton, Dickinson and Company (BD), catalog number: 2140101 ]
- Arabidopsis thaliana seedlings, expressing Golgi-localized proteins fused to a fluorescent protein (e.g. 35S::ST-mRFP in Col, 35S::ERD2-GFP in Col, pGNL1:: GNL1-YFP in gnl1, pGNOM::GNOM-GFP in gnom)
- Inhibitors [such as BrefeldinA (BFA) (Sigma-Aldrich, catalog number: B7651 ), monensin (Sigma-Aldrich, catalog number: M5273 ), salinomycin (Sigma-Aldrich, catalog number: S4526 )]
- Dimethyl sulfoxide (DMSO) (Wako Chemicals USA, catalog number: 046-2198 )
- Freezing planchets type B (Ted Pella, catalog number: 39201 )
- Sucrose (Wako Chemicals USA, catalog number: 196-00015 )
- 1.8 ml cryovials [for example: Nunc® cryotubes (Sigma-Aldrich, catalog number: V7884-450EA )]
- Methacrylate-based resin (Lowicryl HM20 resin kit) (Electron Microscopy Sciences, catalog number: 14340 )
- Sodium phosphate monobasic (NaH2PO4) (Sigma-Aldrich, catalog number: S8282-500G )
- Sodium phosphate dibasic (Na2HPO4) (Sigma-Aldrich, catalog number: S7907-100G )
- Liquid nitrogen
- 0.5% Formvar solution (Electron Microscopy Sciences, catalog number: 15820 )
- Tween-20 (Sigma-Aldrich, catalog number: P1379-25ML )
- Primary antibodies against fluorescent tag [e.g. green fluorescent proteins(GFP)]
- Secondary antibodies conjugated to gold nanoparticles (5, 10, or 15 nm in diameter)
- Cryo-substitution solution (see Recipes)
- Lead citrate (see Recipes)
- Uranyl actetate solution (see Recipes)
- Phosphate-buffer saline (PBS) stock solution (10x) (see Recipes)
- 0.1% Tween-20 in PBS (PBS-T-0.1%) (see Recipes)
- 0.5% Tween-20 in PBS (PBS-T-0.5%) (see Recipes)
- Blocking buffer (see Recipes)
- HM20 resin solutions (see Recipes)
Equipment
- Coverwell® silicone imaging chambers (2.5 mm deep, 20 mm diameter) (Electron Microscopy Sciences, catalog number: 70326-16 )
- Nickel slot grids (Electron Microscopy Sciences, catalog number: 2015-Ni )
- 0.12-0.17 mm thick coverslips (Matsunami Glass, catalog number: C022221 )
- Light Forceps (Hammacher, catalog number: HWC 118-10 )
- Lab-Tek Chambered Coverglass (Thermo Fisher Scientific, catalog number: 155361 )
- 0.9-1.2 mm thick Glass slides (Matsunami Glass, catalog number: S24410 )
- Parafilm
- 22 °C plant growing chamber
- Inverted confocal Microscope (Olympus, model: FV1200 )
Note: equipped with 473 nm or 559 nm diode laser and with water immersion 63x / 1.20 numerical aperture objectives or oil immersion 100x / 1.40 numerical aperture objectives - TIRF Microscope equipped with oil immersion CFI Apo TIRF 100x H / 1.49 numerical aperture objectives (Nikon, model: Eclipse TE2000-E with TIRF2 system)
- High-pressure freezer (Leica, model: EM HPM100 or Bal-tec/RMC/ABRA Fluid AG, model: HPM 010 )
- Automated freeze-substitution and low-temperature resin embedding/polymerization system (for example, Leica, model: AFS2 )
- Ultramicrotome (for example, Leica, model: EM UC7 ultramicrotome )
- Glass knife maker (for example, Leica, model: EM KMR3 )
- Ultra 45° diamond knife or other types of wet diamond knives (Diatome AG)
- Transmission electron microscope (for example, FEI, model: Tecnai 12 )
Procedure
- Live imaging of Golgi apparatus by using Confocal microscope
This protocol is suitable for performing the long-term live imaging of plant cells.- Put seeds on ½X MS media solidified with 0.8% Bacto agar and grow them vertically at 22 °C for 4-5 days under continuous light.
- Collect 4-5 days old Arabidopsis seedlings and transfer them onto ½X MS media solidified with 0.8% Bacto agar, including inhibitor or control DMSO solvent.
Note: To prevent damaging root cells of Arabidopsis seedlings, transferring Arabidopsis seedlings by holding cotyledons with light forceps are recommended. Inhibitor stocks are dissolved in DMSO and thus corresponding amount of DMSO should be contained in control medium. Working concentration of inhibitors can vary depending on chemicals and cell types analyzed but it is normally 10-100 µM.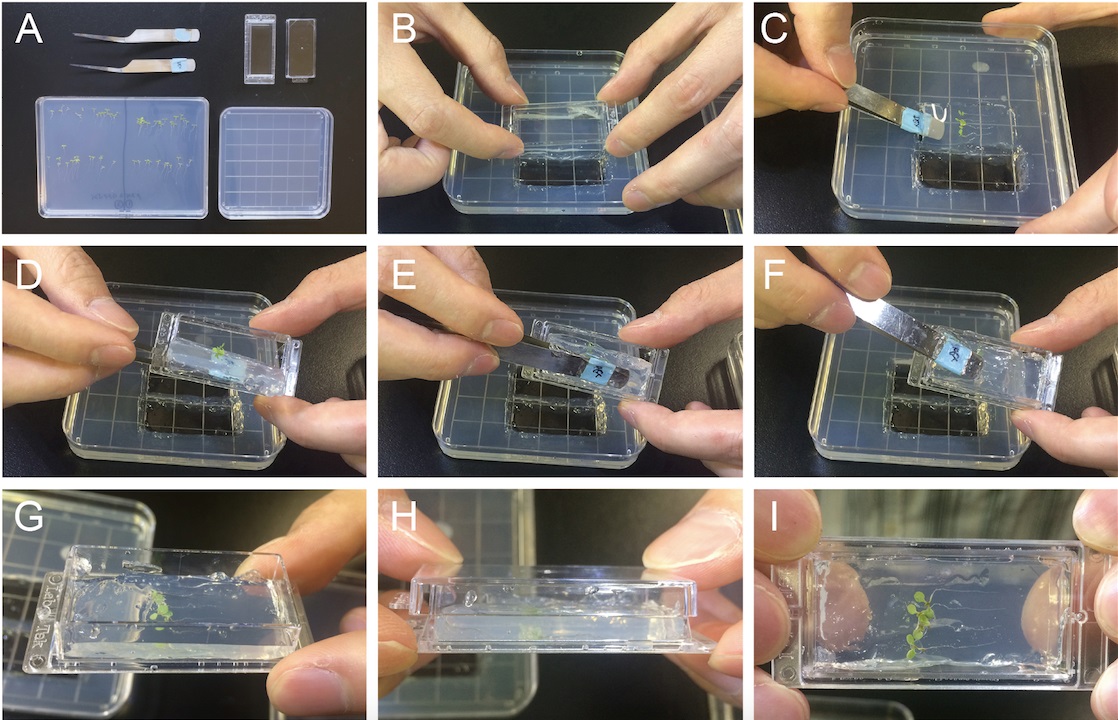
Figure 1. Preparing specimen for long-term live cell imaging using Arabidopsis seedlings. A. Required materials. B-I. Sample preparation for imaging Arabidopsis cells. - Transfer Arabidopsis seedlings and solid media onto the Lab-Tek Chambered Coverglass. Seedlings should be sandwiched between coverslips and solid media (see Video 1 and Figure 1).
Video 1. Specimen preparation for long-term live cell imaging using Arabidopsis seedlings - Place prepared Lab-Tek Chambered Coverglass on the inverted confocal laser microscope equipped with 63x or 100x objectives.
- Focus specimen and observe cells.
Note: Observing cells located close to the coverslip, such as epidermal cells, can provide better image. For performing long-term time lapse imaging, be sure that the objective does not dry out. To prevent water immersion objective from drying, place one drop of Zeiss Immersol W directly onto the objectives or coverslips as immersion medium. As a marker of Golgi apparatus, fluorescently labeled ERD2, ST, COPI proteins can be used (Naramoto et al., 2014). Please note that this experimental setup allows for limited gas exchange, which might affect several processes in the plant. Alternatives for long-term imaging include imaging chambers or microfluidic devices for plants (Busch et al., 2012).
- Put seeds on ½X MS media solidified with 0.8% Bacto agar and grow them vertically at 22 °C for 4-5 days under continuous light.
- Detailed live-imaging analysis of Golgi-localized proteins by using TIRF microscopy
This protocol is suitable for visualizing the detailed subcellular localization of proteins that localized around cell surface.- Place seeds on ½X MS solid media (0.8% Bacto agar) and grow them vertically on 22 °C for 7 days under the continuous light.
- Transfer 7-day-old Arabidopsis seedlings into ½X MS liquid medium and incubate them for longer than 30 min. If necessary, inhibitor or DMSO control solvent can be added.
Note: Incubation time can vary depending on the experiments but it is normally from 30 min to 2 h. - Excise root or hypocotyl segments from 7-day-old Arabidopsis seedlings by using fine scissors or surgical knives (see Figure 2).
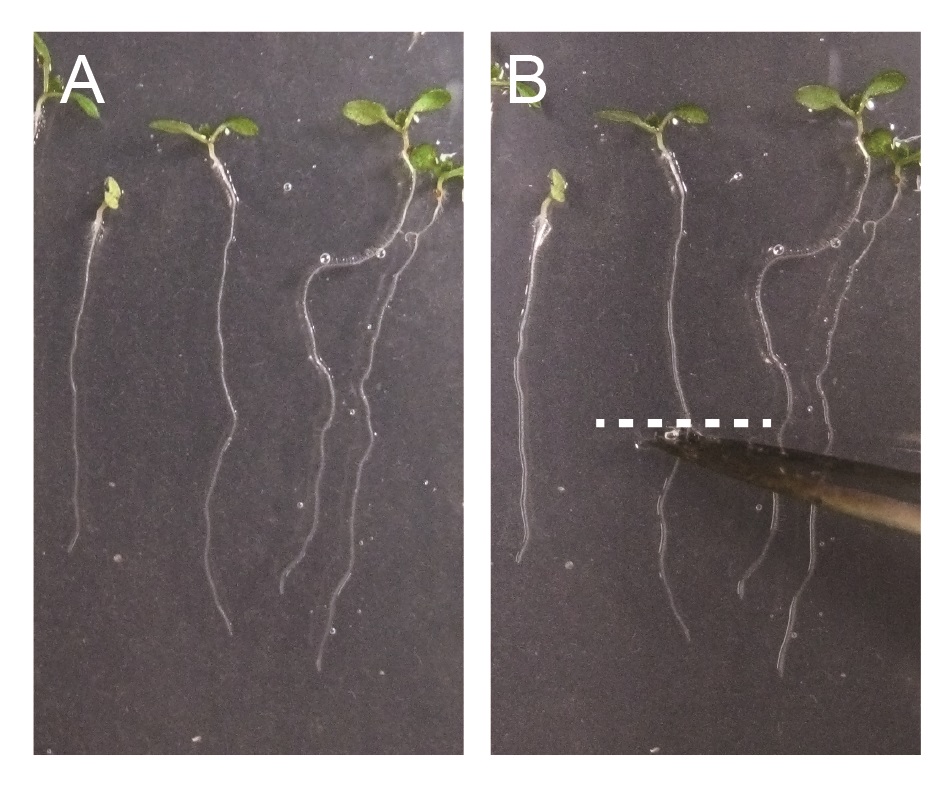
Figure 2. Excision of Arabidopsis roots for TIRF microscopy imaging. A. 7-day-old Arabidopsis seedlings. B. 7-day-old Arabidopsis seedlings, excised by scissor. Excision position at roots or hypocotyls are indicated by dashed white line. - Mount root or hypocotyl segments with liquid media using glass slides and cover slips.
- Place prepared specimen on the TIRF microscope equipped with an oil immersion CFI Apo TIRF 100x H/1.49 numerical aperture objective.
- Focus the specimen and observe cells.
Note: Only cells in contact with the surface of coverslip can be imaged. For the roots, cells in elongation zone, especially those close to the differentiation zone, are suitable for observation. Please see example TIRF image of pGNOM::GNOM-GFP that localize at Golgi apparatus (see Video 2).
Video 2. Live imaging of GNOM-GFP localized at Golgi apparatus by TIRF microscopy as described in this protocol
- Place seeds on ½X MS solid media (0.8% Bacto agar) and grow them vertically on 22 °C for 7 days under the continuous light.
- Immunogold labeling of Golgi localized proteins by using TEM
- Submerge seedlings grown on ½ MS plates and expressing a fluorescently-tagged Golgi-localized protein in a drop of 0.1 M sucrose. Cut 1 mm-long root segments and place them inside freezing planchet (Figure 3A) filled with 0.1 M sucrose. Besides transgenic roots expressing fluorescently-tagged proteins, process also wild-type samples to use as negative controls during immunolabeling.
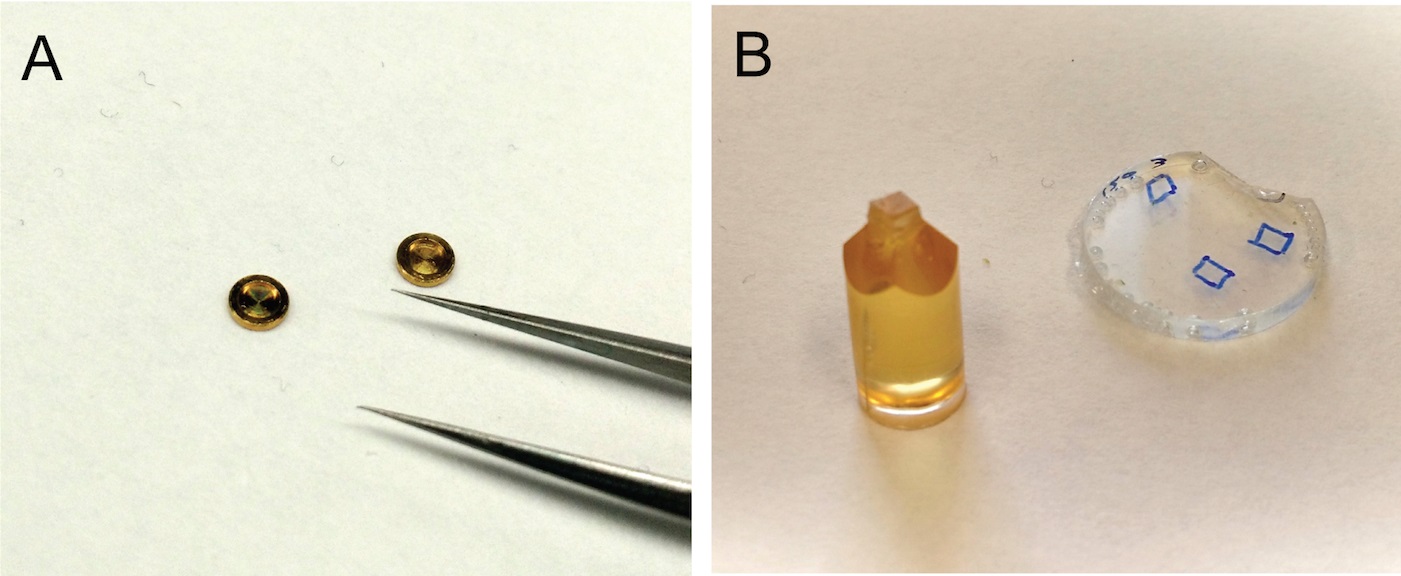

Figure 3. Preparation of root samples for immunogold labeling. A. Freezing planchets for high-pressure freezing. B. Mounting of HM20 resin-embedded roots on stubs for sectioning. - Place another freezer planchet on top, flat side down, to close the chamber.
- Freeze the two planchets containing the root segments in a high-pressure freezer.
- Under liquid nitrogen, separate the two planchets and transfer the one containing the frozen roots to a cryovial with cryosubstitution solution.
- Place cryovial with cryosubstitution solution and samples into a cryosubstitution device pre-cooled to -90 °C. Samples should remain at -90 °C for at least 3 days.
- Raise the temperature of the cryosubstitution device to -60 °C. Remove cryosubstitution solution and rinse samples with fresh, pre-cooled (-60 °C) anhydrous acetone 3-4 times. Remove empty freezing planchets using pre-cooled tweezers (all root segments should be detached from the planchets and free inside in the cryovial with acetone).
- Remove acetone and add pre-cooled (-60 °C) 30% HM20 solution for at least 3 h. Repeat this step with pre-cooled (-60 °C) 60% HM20 solution (3 h) and 100% HM20 (3 h). At least 3 more changes with 100% HM20 are recommended. During the entire embedding procedures, samples should remained at -60 °C.
- Assemble embedding chambers by attaching Coverwell® imaging chambers to a glass slide. Fill embedding chambers with fresh 100% HM20 and transfer roots from cryovial to embedding chambers. Cover with a glass coverslip.
- Polymerize HM20 resin at -50 °C with UW light for 48 h.
- Once polymerized, remove resin blocks from embedding chambers. Locate roots within resin block with the help of a dissecting microscope. Cut pieces or resin containing roots and mount them on top of resin stubs using super glue (Figure 3B).
- With a razor blade trim the resin block around the root. Start sectioning with a glass knife in an ultramicrotome until reaching the root. Switch to a diamond knife; collect 70 nm-thick sections on Formvar coated-nickel grids.
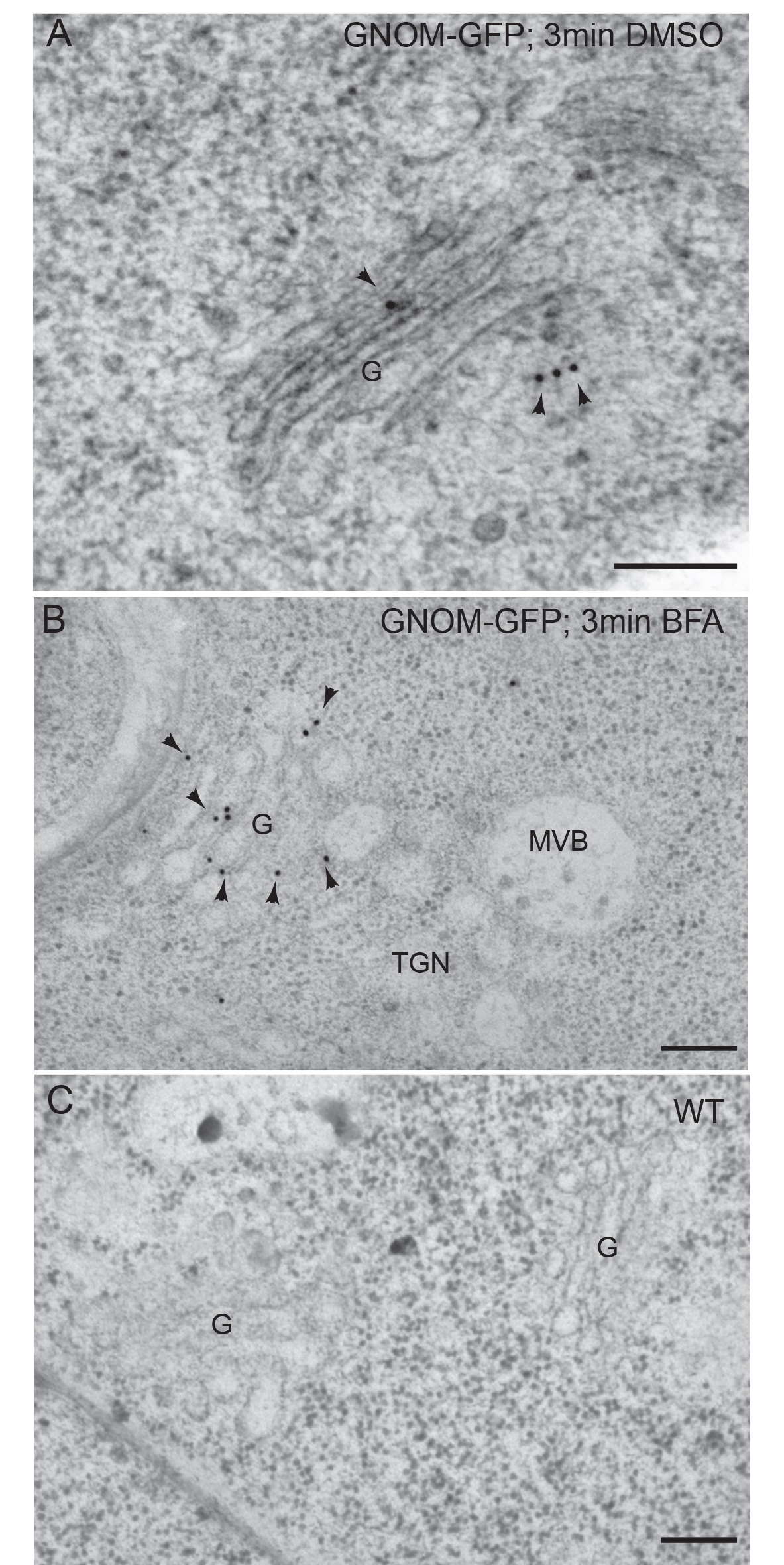
Figure 4. Immunogold labeling of GNOM-GFP with anti-GFP antibodies on high-pressure frozen-freeze substituted Arabidopsis roots. A. Detection of GNOM-GFP in transgenic roots treated for 3 min in DMSO. B. Detection of GNOM-GFP in transgenic roots treated for 3 min in BFA. C. Negative control: wild type roots treated with DMSO for 3 min. Arrowheads indicate gold particles. G, Golgi; MVB, multivesicular body. TGN, Trans Golgi Network. Scale bars = 200 nm. Modified from Naramoto et al. (2014) Copyright American Society of Plant Biologists. - Place 10 μl drops of blocking buffer on a piece of parafilm and float nickel grids on top of drops (the root sections should be in contact with the blocking buffer) for 15-20 min.
- Transfer grids to 10 μl drops of primary antibody diluted in blocking buffer (recommended dilutions: 1:10 to 1:50) for 1 h.
- Rinse grids under a stream of PBS-T-0.5% buffer for 1 min. Blot grids with filter paper and float them on 10 μl drops of secondary antibody solution (recommended dilutions 1:10 to 1:100) for 1 h.
- Rinse grids with PBS-T-0.5% buffer for 1 min followed by a rinse with distilled water.
- Stain root sections with uranyl acetate solution for 10 min followed by lead citrate for 5 min.
- Observed sections in a transmission electron microscope (Figure 4).
- Submerge seedlings grown on ½ MS plates and expressing a fluorescently-tagged Golgi-localized protein in a drop of 0.1 M sucrose. Cut 1 mm-long root segments and place them inside freezing planchet (Figure 3A) filled with 0.1 M sucrose. Besides transgenic roots expressing fluorescently-tagged proteins, process also wild-type samples to use as negative controls during immunolabeling.
Recipes
- Cryo-substitution solution
0.2% glutaraldehyde plus 0.2% uranyl acetate in acetone
Place 1.5 ml in a 2 ml cryovial and store individual aliquots in liquid nitrogen - Lead citrate
- Wear gloves to handle lead nitrate
Boil 50 ml of distilled water to remove CO2 (CO2 dissolved in water can cause this solution to precipitate) and let the water cool down at room temperature - Add 0.33 g of lead nitrate to 10 ml of boiled distilled water and mix gently until the lead nitrate crystals completely dissolve
- Add 0.44 g of sodium citrate and mixed gently; the solution will become milky white
- Add 2 ml of 1 N sodium hydroxide solution freshly prepared with boiled water; the solution will become transparent
- Bring up volume to 12.5 ml with boiled distilled water
- Store solution at either room temperature or 4 °C
- Wear gloves to handle lead nitrate
- Uranyl actetate solution
2% uranyl acetate (w/v) in 30% methanol (v/v) - Phosphate-buffer saline (PBS) stock solution (10x)
1.76 g of NaH2PO4, 11.49 g of Na2HPO4, 85 g sodium chloride in 1 L of distilled water, pH 6.8 (store at room temperature) - 0.1% Tween-20 in PBS (PBS-T-0.1%)
Add 100 μl of Tween-20 to 100 ml of 1x PBS
Wash Tween-20 out of the pipette tip by pipetting up and down several times into the PBS-T-0.1% solution - 0.5% Tween-20 in PBS (PBS-T-0.5%)
Add 5 ml of Tween-20 to 1 L of 1x PBS - Blocking buffer
10% (w/v) nonfat dry milk in PBS-T-0.1%
For a 10 ml volume, add PBS-T-0.1% to 1 g of nonfat dry milk up to 10 ml final volume - HM20 resin solutions
Prepare HM20 resin in the hood according to manufacturer’s instructions
Make 30% and 60% HM20 resin solutions (v/v) by mixing with the appropriate volume of anhydrous acetone
Acknowledgments
This work was supported by Grant for Basic Science Research Projects from The Sumitomo Foundation to SN; the Japanese Society for the Promotion of Science (JSPS; 30612022 to S.N.); the Ministry of Education, Culture, Sports, Science and Technology in Japan [NC-CARP project (to S.N.)] and U. S. National Science Foundation grant MCB1157824 (to MSO).
References
- Busch, W., Moore, B. T., Martsberger, B., Mace, D. L., Twigg, R. W., Jung, J., Pruteanu-Malinici, I., Kennedy, S. J., Fricke, G. K., Clark, R. L., Ohler, U. and Benfey, P. N. (2012). A microfluidic device and computational platform for high-throughput live imaging of gene expression. Nat Methods 9(11): 1101-1106.
- Naramoto, S., Otegui, M. S., Kutsuna, N., de Rycke, R., Dainobu, T., Karampelias, M., Fujimoto, M., Feraru, E., Miki, D., Fukuda, H., Nakano, A. and Friml, J. (2014). Insights into the localization and function of the membrane trafficking regulator GNOM ARF-GEF at the Golgi apparatus in Arabidopsis. Plant Cell 26(7): 3062-3076.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Naramoto, S., Dainobu, T. and Otegui, M. S. (2015). A Bioimaging Pipeline to Show Membrane Trafficking Regulators Localized to the Golgi Apparatus and Other Organelles in Plant Cells. Bio-protocol 5(17): e1583. DOI: 10.21769/BioProtoc.1583.
Category
Plant Science > Plant cell biology > Cell imaging
Cell Biology > Cell imaging > Confocal microscopy
Cell Biology > Cell imaging > Electron microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link