- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
DNA in situ Hybridizations for VEGFA Gene Locus (6p12) in Human Tumor Tissue
Published: Vol 5, Iss 15, Aug 5, 2015 DOI: 10.21769/BioProtoc.1553 Views: 9990
Reviewed by: HongLok LungMartin V KolevOmar Akil

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Enhanced-ice-COLD-PCR for the Sensitive Detection of Rare DNA Methylation Patterns in Liquid Biopsies
Florence Mauger and Jörg Tost
Dec 5, 2019 6441 Views

Single-cell qPCR Assay with Massively Parallel Microfluidic System
Marta Prieto-Vila [...] Yusuke Yamamoto
Mar 20, 2020 5802 Views

In-Cell Western Protocol for Semi-High-Throughput Screening of Single Clones
Arpita S. Pal [...] Andrea L. Kasinski
Aug 20, 2022 3509 Views
Abstract
Over the last decades numerous regulators of angiogenesis have been identified and characterized. Among the others the vascular endothelial growth factor (VEGFA) appears undoubtedly important in several pathophysiological processes. Moreover, VEGFA represents one of the most attractive targets of anticancer therapy, given its major role in the growth and development of different tumor types. Here we describe a method to detect the copy number variation (CNV) status of the VEGFA gene by fluorescence in situ hybridization (FISH). FISH analysis is a reliable method for investigating VEGFA amplification or increased gene copy number and may represent an alternative method to immunohistochemical analysis for investigating the deregulation of VEGFA expression levels.
Keywords: VEGFAMaterials and Reagents
- BAC clone No. RPCIB753M0921Q (GenomeCUBE)
- LB medium (Sigma-Aldrich, catalog number: L3522-250G )
- Large-construct kit (QIAGEN, catalog number: 12462 )
- SP6 (EuroFins MGW Operon LLC A Eurofins Genomics Company, catalog number: SP050-1 ) and T7 (EuroFins MGW Operon LLC A Eurofins Genomics Company, catalog number: SP401 ) primers
- AluI restriction enzyme (Life Technologies, InvitrogenTM, catalog number: 45200-029 )
- Cy3-dUTP (GE Healthcare Dharmacon, catalog number: 45-000-738 )
- BioPrime array CGH kit (module) (Life Technologies, InvitrogenTM, catalog number: 18095-012 ) (it includes 2.5 Random primers solution, 10x dUTP nucleotide mix, Exo-Klenow fragment)
- Amicon Ultra-0.5 centrifugal filter unit with Ultracel-30 membrane (Merck KGaA, catalog number: UFC503096 )
- Human Cot-1 DNA (Life Technologies, InvitrogenTM, catalog number: 15279-011 )
- DAPI solution (1 µg/ml) (Abbott Laboratories, catalog number: 06J49-001 )
- Spectrum Green-labeled Chr6 centromeric probe (CEP6) and hybridization buffer (Abbott Laboratories, catalog number: 06J37-016 and 07J36-001 )
- Micron-centrifugal filters (Merck KGaA, catalog number: MRCF0R030 )
- TE Buffer, 1x Solution 500 ml (USB, catalog number: 75893 )
- 20x SSC (Abbott Laboratories, catalog number: 02J10-032 )
- 0.3 % NP40 (Abbott Laboratories, catalog number: 07J05-001 )
- Pretreatment reagent VP 2000 (Abbott Laboratories, catalog number: 02J06-030 )
- Protease 1 (250 mg) (Abbott Laboratories, catalog number: 02J08-032 )
- Ethanol (Sigma- Aldrich, catalog number: 0 2860 )
- Rubber cement (Carfa AG, catalog number: 00494 )
- Protease 1 (250 mg in 500 ml protease buffer) (Sauter et al., 1995)
- Wash buffer (0.4 SSC + 0.3% NP40-pH 7-7.5) (Sauter et al., 1995)
Equipment
- Incubator for slides (Memmert GmbH + Co.KG, Mode: ll IPP 300 )
- Pipette (Eppendorf)
- Nanodrop spectrophotometer assay (Thermo Fisher Scientific, model: NanoDrop 2000 )
- Slide (Thermo Fisher Scientific, catalog number: No. 10143352 )
- Cover slide (Biosystem 24 x 50 mm, catalog number: 14071135638 and R. Langenbrinck 21 x 26 mm, catalog number: 01-2126 )
- Hotplate for denaturation (Medax GmbH & Co.KG, catalog number: 12895 )
- Zeiss fluorescence microscope using a 63x objective (Carl Zeiss AG, model: Axioskop 40 )
- AxioVision software (Carl Zeiss AG, model: SE64 Rel.4.9 )
Procedure
- How to make the probe
- Inoculate starter culture with the BAC clone (e.g. glycerol stock) in 2-5 ml LB medium, containing the appropriate selective antibiotic (chloramphenicol).
- Incubate over-night 25/30 °C, shaking at 200 rpm.
- Dilute 0.5 ml of the starter culture in 500 ml selective LB medium (1/500 to 1/1,000), containing the appropriate antibiotic (chloramphenicol).
- Incubate over-night with shaking (200 rpm).
- Isolate the BAC-DNA using the Large-Construct Kit according to the manufacturer’s instructions.
- Verify BAC identity by sequencing using 1 μg of isolated DNA and 20 pmol of SP6, T7 primers, respectively.
- Digest ~1 μg BAC-DNA with AluI restriction enzyme according to the manufacturer’s instructions.
- Suspend the digested BAC-DNA in sterile water to a final volume of 21 μl (BioPrime array CGH kit).
- Add 20 μl of 2.5x Random primers solution (BioPrime array CGH kit) to each sample and incubate at 95 °C for 5 min.
- On ice, add 5 μl of 10x dUTP Nucleotide Mix (BioPrime array CGH kit).
- Add 3 μl of Cy3-dUTP.
- Add 1 μl of Exo-Klenow fragment (BioPrime array CGH kit).
- Mix gently and perform a quick spin down (5 sec) and incubate at 37 °C for 2 h.
- Add 5 μl of Stop Buffer to each tube and place on ice (BioPrime array CGH kit).
Labeling mix Reagent Added per Sample Volume Added per Sample 2.5x Random primers solution (BioPrime array CGH kit) and incubate at 95 °C for 5 min 20 µl On ice, 10x dUTP Nucleotide Mix 5 µl Cy3-dUTP 3 µl Exo-Klenow fragment 1 µl Mix gently and perform a quick spin down (5 sec) then incubate at 37 °C for 2 h Stop Buffer to each tube and place on ice (long term storage at -20 °C) 5 µl - Label the Micron tubes on both the lid and the side with sample identifiers, as the lids tend to break. Also label the side of the filter with an identifier.
- Load the Micron filter into the 1.5 ml tubes provided with the kit, so that the filter is sifting on the ridges inside the tube.
- Pipette 450 μl of TE 1x into each filter.
- Load the samples into the filters.
- Centrifuge at 8,000 x g for 10 min at room temperature.
- Check the filter (visual inspection by eyes) to make sure the Cy3-Dye labelled DNA has stuck to it.
- Add 500 μl of TE to the filter and centrifuge at 8,000 x g for 10 min.
- Discard the flow through and place in a new 1.5 ml filter tube upside-down to elute.
- Centrifuge at 8,000 x g for 1 min.
- Check the filter to make sure the entire sample is eluted from the filter (store samples short-term at 4 °C. Store long-term at -20 °C).
- Add 20 μl of Human Cot-1 DNA.
- Add 5 μl of CEP 6.
- Add 35 μl of hybridization buffer (keep in ice or store at -20 °C).
- Inoculate starter culture with the BAC clone (e.g. glycerol stock) in 2-5 ml LB medium, containing the appropriate selective antibiotic (chloramphenicol).
- How to prepare the tissue
- Pretreat FFPE tissue microarrays or FFPE whole tissue sections (7 μm thick) 10 min in the Xylene-1 min in 100% EtOH-1 min in 90% EtOH-1 min in 70% EtOH-1 min in H2O.
- Continue pretreating the tissue sections for 15 min in the pre-warmed paraffin pretreatment reagent VP2000 (80 °C).
- Immerse the tissue sections for 1 min in the H2O.
- Immerse the tissue sections for 2 h into the pre-warmed 37 °C protease buffer (Protease 1 - 250 mg in 500 ml Protease Buffer).
- Immerse the tissue sections for 1 min in the H2O.
- Immerse the tissue sections for 1 min in the 70% EtOH-1 min in the 90% EtoH-1 min in the 100% EtOH-1 min in the H2O.
- Let the tissue section air-dry at room temperature for approximately 10 min.
- Pretreat FFPE tissue microarrays or FFPE whole tissue sections (7 μm thick) 10 min in the Xylene-1 min in 100% EtOH-1 min in 90% EtOH-1 min in 70% EtOH-1 min in H2O.
- Hybridization
- Apply 10-20 μl of the probe on the tissue sections and cover it.
- Apply the rubber cement to prevent the probe from leaking.
- Leave the tissue sections for 20 min at 75 °C.
- Incubate the tissue sections over-night at 37 °C.
- Uncover the tissue sections.
- Wash the tissue sections in the wash buffer at room temperature 5 min.
- Wash the tissue sections in the wash buffer, pre-warmed at 48 °C between 2-5 min (when you have background wash longer and with higher temperature).
- Immerse the tissue sections 1 min in the H2O.
- Let the tissue sections dry protecting it from the light.
- Add 10-20 μl of DAPI solution (1,000 ng/ml) to the tissue sections.
- Cover the tissue sections with cover glass 24 x 50 mm.
- Place the tissue sections 15 min at 4 °C.
- Evaluate the tissue sections using a fluorescence microscope (63x objective).
- Count a minimum of 100 tumor nuclei signals in four separate regions of the tissue section independently.
- Apply 10-20 μl of the probe on the tissue sections and cover it.
Representative data
FISH results (representative pictures are shown in Figure 1) were interpreted according to: (1) absolute Chr6 copy number and (2) the ratio VEGFA gene/Chr6 copy number. We classified as not amplified samples with a VEGFA/Chr6 ratio of <1.8 and an average VEGFA copy number <4.0 signals per cell; equivocal with a VEGFA/Chr6 ratio <2.0 and an average VEGFA copy number ≥ 4.0 and <6.0 signals per cell and amplified with a VEGFA/Chr6 ratio of <2.2 and an average VEGFA copy number of ≥6.0 signals per cell or a VEGFA/Chr6 ratio ≥2.2 with an average VEGFA copy number of ≥4.0 signals per cell, as proposed by the ASCO/CAP guidelines for HER2 amplification in breast cancer (Wolff et al., 2013). Polysomy of Chr6 was defined as an average of the Chr6 copy number. When the average was included between 2.26 and 3.75, the polysomy 6 was defined as low whereas, when the average was >3.75 the polysomy 6 was defined high (Andreozzi et al., 2014; Sauter et al., 1995; Wolff et al., 2007; Salido et al., 2005; Torrisi et al., 2007; Tapia et al., 2007; Ma et al., 2005).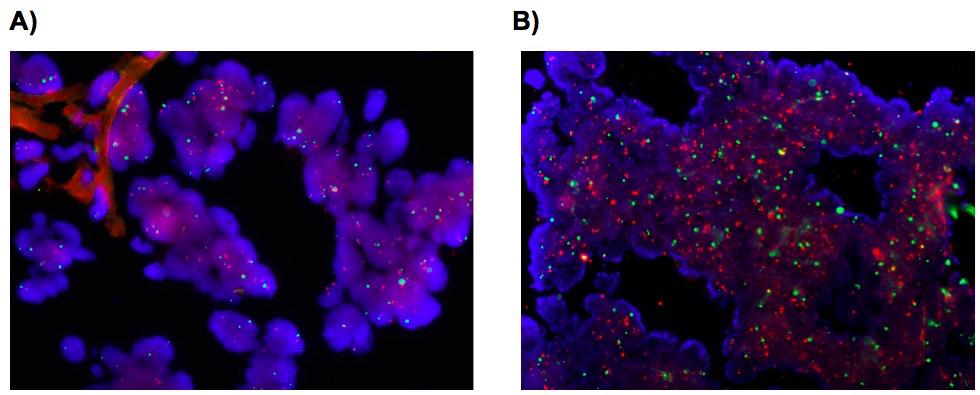
Figure 1. Fluorescence in situ hybridization (FITC+Rhodamine+DAPI) of the VEGFA gene locus 6p12 in colorectal cancer (CRC), (A) non-amplified CRC sample, (B) amplified CRC sample. Red dots highlight VEGFA gene, green dots highlight Chr6 centromere.
Acknowledgments
We would like to acknowledge Matthias Matter, MD, PhD, Cristina Quintavalle, PhD and Christian Ruiz, PhD, for their suggestions and critical comments during the establishment phase of this methodology. This protocol represents a modified enhanced type of the previously used version (Sauter et al., 1995; Vlajnic et al., 2011; Andreozzi et al., 2014).
References
- Andreozzi, M., Quagliata, L., Gsponer, J. R., Ruiz, C., Vuaroqueaux, V., Eppenberger-Castori, S., Tornillo, L. and Terracciano, L. M. (2014). VEGFA gene locus analysis across 80 human tumour types reveals gene amplification in several neoplastic entities. Angiogenesis 17(3): 519-527.
- Ma, Y., Lespagnard, L., Durbecq, V., Paesmans, M., Desmedt, C., Gomez-Galdon, M., Veys, I., Cardoso, F., Sotiriou, C., Di Leo, A., Piccart, M. J. and Larsimont, D. (2005). Polysomy 17 in HER-2/neu status elaboration in breast cancer: effect on daily practice. Clin Cancer Res 11(12): 4393-4399.
- Salido, M., Tusquets, I., Corominas, J. M., Suarez, M., Espinet, B., Corzo, C., Bellet, M., Fabregat, X., Serrano, S. and Sole, F. (2005). Polysomy of chromosome 17 in breast cancer tumors showing an overexpression of ERBB2: a study of 175 cases using fluorescence in situ hybridization and immunohistochemistry. Breast Cancer Res 7(2): R267-273.
- Sauter, G., Moch, H., Carroll, P., Kerschmann, R., Mihatsch, M. J. and Waldman, F. M. (1995). Chromosome-9 loss detected by fluorescence in situ hybridization in bladder cancer. Int J Cancer 64(2): 99-103.
- Tapia, C., Savic, S., Wagner, U., Schonegg, R., Novotny, H., Grilli, B., Herzog, M., Barascud, A. D., Zlobec, I., Cathomas, G., Terracciano, L., Feichter, G. and Bubendorf, L. (2007). HER2 gene status in primary breast cancers and matched distant metastases. Breast Cancer Res 9(3): R31.
- Torrisi, R., Rotmensz, N., Bagnardi, V., Viale, G., Curto, B. D., Dell'orto, P., Veronesi, P., Luini, A., D'Alessandro, C., Cardillo, A., Goldhirsch, A. and Colleoni, M. (2007). HER2 status in early breast cancer: relevance of cell staining patterns, gene amplification and polysomy 17. Eur J Cancer 43(16): 2339-2344.
- Vlajnic, T., Andreozzi, M. C., Schneider, S., Tornillo, L., Karamitopoulou, E., Lugli, A., Ruiz, C., Zlobec, I. and Terracciano, L. (2011). VEGFA gene locus (6p12) amplification identifies a small but highly aggressive subgroup of colorectal cancer [corrected] patients. Mod Pathol 24(10): 1404-1412.
- Wolff, A. C., Hammond, M. E., Hicks, D. G., Dowsett, M., McShane, L. M., Allison, K. H., Allred, D. C., Bartlett, J. M., Bilous, M., Fitzgibbons, P., Hanna, W., Jenkins, R. B., Mangu, P. B., Paik, S., Perez, E. A., Press, M. F., Spears, P. A., Vance, G. H., Viale, G., Hayes, D. F., American Society of Clinical, O. and College of American, P. (2013). Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol 31(31): 3997-4013.
- Wolff, A. C., Hammond, M. E., Schwartz, J. N., Hagerty, K. L., Allred, D. C., Cote, R. J., Dowsett, M., Fitzgibbons, P. L., Hanna, W. M., Langer, A., McShane, L. M., Paik, S., Pegram, M. D., Perez, E. A., Press, M. F., Rhodes, A., Sturgeon, C., Taube, S. E., Tubbs, R., Vance, G. H., van de Vijver, M., Wheeler, T. M., Hayes, D. F., American Society of Clinical, O. and College of American, P. (2007). American society of clinical oncology/college of american pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25(1): 118-145.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Andreozzi, M., Quagliata, L., Burmeister, K., Arabi, L., Schneider, S., Tornillo, L. and Terracciano, L. (2015). DNA in situ Hybridizations for VEGFA Gene Locus (6p12) in Human Tumor Tissue. Bio-protocol 5(15): e1553. DOI: 10.21769/BioProtoc.1553.
Category
Cancer Biology > General technique > Genetics > Gene expression
Cell Biology > Cell staining > Nucleic acid
Molecular Biology > DNA > DNA labeling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










