- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Phosphopeptide Purification Protocol for the Moss Physcomitrella paten
Published: Vol 5, Iss 14, Jul 20, 2015 DOI: 10.21769/BioProtoc.1527 Views: 9567
Reviewed by: Valentine V TrotterArsalan Daudi

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
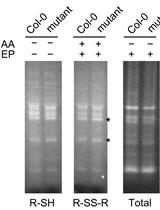
Detection of Disulfides in Protein Extracts of Arabidopsis thaliana Using Monobromobimane (mBB)
Shin‐nosuke Hashida and Maki Kawai-Yamada
Mar 5, 2019 6430 Views
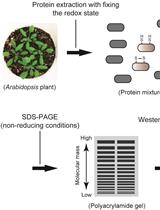
Simple Method to Determine Protein Redox State in Arabidopsis thaliana
Keisuke Yoshida and Toru Hisabori
Jun 5, 2019 7483 Views
In vitro Auto- and Substrate-Ubiquitination Assays
Hye Lin Park [...] Gyeong Mee Yoon
Apr 5, 2022 3557 Views
Abstract
Protein phosphorylation is one of the most common post-translational modifications in eukaryotic cells and plays a critical role in a vast array of cellular processes. Efficient methods of protein extraction and phosphopeptide purification are required to ensure the detection of high quality of proteins. In our hands, phenol extraction of proteins and TiO2 chromatography enrich phosphorylated peptides more efficiently than other methods in the moss Physcomitrlla patens (P. patens).
Materials and Reagents
- Seven-day old protonemata of the moss P. patens
- Liquid nitrogen
- Tris-HCl (pH 7.5)-saturated phenol (AMRESCO, catalog number: K168 )
- Ammonium acetate (Sigma-Aldrich, catalog number: A1542 )
- Methanol (Sigma-Aldrich, catalog number: 34860 )
- DTT (Promega Corporation, catalog number: V3155 )
- Acetone (Sigma-Aldrich, catalog number: 650501 )
- Iodoacetamide (Sigma-Aldrich, catalog number: I6125 )
- Ammonium bicarbonate (pH 8.5) (Sigma-Aldrich, catalog number: 40867 )
- Trypsin (Promega Corporation, catalog number: V5111 )
- Sucrose (Sigma-Aldrich, catalog number: S5390 )
- Tris (Sigma-Aldrich, catalog number: T1378 )
- EDTA (Sigma-Aldrich, catalog number: E6635 )
- 1,4-dithiothreitol (DTT) (Promega Corporation, catalog number: V3155)
- Protease inhibitors cocktail (Sigma-Aldrich, catalog number: P9599 )
- Phosphatase inhibitors cocktail 2 (Sigma-Aldrich, catalog number: P0044 )
- NH4OH (Sigma-Aldrich, catalog number: 320145 )
- Urea (Sigma-Aldrich, catalog number: U6504 )
- CHAPS (Sigma-Aldrich, catalog number: C9426 )
- Acetonitrile (ACN) (Sigma-Aldrich, catalog number: 34851 )
- Trifluoroacetic acid (TFA) (Sigma-Aldrich, catalog number: 302031 )
- Glutamic acid (Sigma-Aldrich, catalog number: G0355000 )
- Protein extraction buffer (see Recipes)
- Protein resuspension buffer (see Recipes)
- Loading buffer (see Recipes)
- Washing buffer I (see Recipes)
- Washing buffer II (see Recipes)
- Elution buffer (see Recipes)
Equipment
- Porcelain Mortar and Pestle (90 mm)
- Tubes and tube holder (2 ml)
- Titanium dioxide micro-columns (320 µm x 50 mm) (Column Technology, Freemont)
- Centrifuge
- Vortex
- Freeze dryer (Christ, model: Alpha 1-4 LSC )
Procedure
This procedure includes proteins extraction, protein digestion and phosphopeptide purification. The following should all be carried out at cold room temperature except for special instructions.
- Add protease inhibitor mix (protease inhibitors and phosphatase inhibitors) to the ice-cold protein extraction buffer according to the manufacturer’s instructions.
- Pour liquid nitrogen into mortar until nearly full in order to pre-cool the mortar and pestle.
- Add the fresh or frozen (-80 °C freezer) tissue (about 1 g) into mortar, grind them to fine powder in liquid nitrogen, and then thaw on ice for 10-15 min.
- Add 2 ml ice-cold protein extraction buffer into mortar, homogenize by grinding on ice for 10 min, and then transfer 0.8 ml to each tube.
- Add an equal volume of ice-cold Tris-HCl (pH 7.5)-saturated phenol to each tube, vortex tubes for 5-10 min.
- Centrifuge samples at 13,000 rpm for 15 min at 4 °C.
- Collect phenol phase and mixed with three volumes of 100 mM ammonium acetate in methanol.
- Precipitate proteins from mixture of phenol phase and 100 mM ammonium acetate overnight at -20 °C.
- Centrifuge proteins at 13,000 rpm for 15 min at 4 °C.
- Rinse proteins 3 times with ice-cold acetone containing 13 mM DTT and then lyophilize by freeze dryer for 5-10 min.
- The protein concentration was determined according to Peterson (1977) using BSA as a standard.
- Dissolve 500 µg protein power in 100 µl protein resuspension buffer at room temperature.
- Reduce protein (500 µg protein in 100 µl resuspension buffer) with 20 mM DTT at 37 °C for 2.5 h and alkylate protein with 40 mM iodoacetamide for 40 min at room temperature in the dark.
- Precipitate the protein mixture with 1 ml 100% acetone overnight at -20 °C.
- Collect protein by centrifugation, dissolve protein in 100 mM ammonium bicarbonate (pH 8.5), and digest protein with trypsin (50:1 w/w, protein to trypsin ratio) at 37 °C for 16 h.
- Lyophilize the digested peptide mixture, and then dilute in a 200 µl loading buffer.
- Rinse the titanium dioxide micro-columns (TiO2 columns, 320 µm x 50 mm) with 100 µl loading buffer.
- Load the 200 µl samples onto TiO2 columns by applying air pressure created with a plastic syringe.
- Wash the TiO2 columns with 100 µl loading buffer, 100 µl washing buffer I, and finally with 100 µl washing buffer II.
- Elute the columns which bound phosphopeptides with 100 µl of elution buffer.
- Pool and lyophilize the eluates which is purification of phosphopeptides [Representative data is shown in Wang et al. (2014a) and Wang et al. (2014b)].
Recipes
- Protein extraction buffer
250 mM sucrose
20 mM Tris-HCl (pH 7.5)
10 mM EDTA 1 mM 1,4-dithiothreitol
Protease inhibitors cocktail
Phosphatase inhibitors cocktail 2 - Protein resuspension buffer
8 M urea
4% CHAPS
65 mM DTT
40 mM Tris-HCl (pH 7.5) - Loading buffer
65% ACN and 2% TFA solution saturated with glutamic acid
- Washing buffer I
65% ACN
0.5% TFA
- Washing buffer II
65% ACN
0.1% TFA
- Elution buffer
300 mM NH4OH
50% ACN
Acknowledgments
This work was supported by grants from Project of Ministry Science & Technology of China (2013CB967300, 2007CB948201) to Prof. He, grant from National Natural Science Foundation of China (No. 31371243, 30970250), China Postdoctoral Science Foundation and State Education Ministry Scientific Research Foundation for the Returned Overseas Chinese Scholars to Dr. Wang.
References
- Peterson, G. L. (1977). A simplification of the protein assay method of Lowry et al. which is more generally applicable. Anal Biochem 83(2): 346-356.
- Wang, X., Qi, M., Li, J., Ji, Z., Hu, Y., Bao, F., Mahalingam, R. and He, Y. (2014a). The phosphoproteome in regenerating protoplasts from Physcomitrella patens protonemata shows changes paralleling postembryonic development in higher plants. J Exp Bot 65(8): 2093-2106.
- Wang, X. Q., Yang, P. F., Liu, Z., Liu, W. Z., Hu, Y., Chen, H., Kuang, T. Y., Pei, Z. M., Shen, S. H. and He, Y. K. (2009). Exploring the mechanism of Physcomitrella patens desiccation tolerance through a proteomic strategy. Plant Physiol 149(4): 1739-1750.
- Wang, X., Zhou, S., Chen, L., Quatrano, R. S. and He, Y. (2014b). Phospho-proteomic analysis of developmental reprogramming in the moss Physcomitrella patens. J Proteomics 108: 284-294.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wang, X. and He, Y. (2015). A Phosphopeptide Purification Protocol for the Moss Physcomitrella paten. Bio-protocol 5(14): e1527. DOI: 10.21769/BioProtoc.1527.
Category
Plant Science > Plant biochemistry > Protein > Modification
Plant Science > Plant biochemistry > Protein > Labeling
Systems Biology > Proteomics > Phosphoproteomics
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










