- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Virus-induced Gene Silencing (VIGS) in Barley Seedling Leaves
Published: Vol 5, Iss 12, Jun 20, 2015 DOI: 10.21769/BioProtoc.1506 Views: 14464
Reviewed by: Samik BhattacharyaPablo Bolanos-VillegasSollapura J. Vishwanath

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Real-time PCR Analysis of PAMP-induced Marker Gene Expression in Nicotiana benthamiana
Fan Liu [...] Yuanchao Wang
Oct 5, 2018 9326 Views
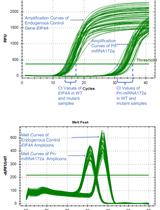

RNA Stability Measurements Using RT-qPCR in Arabidopsis Seedlings
Tianran Jia and Brandon H. Le
Jul 20, 2020 7036 Views
Laser-Assisted Microdissection and High-Throughput RNA Sequencing of the Arabidopsis Gynoecium Medial and Lateral Domains
Valentín Luna-García and Stefan de Folter
Sep 5, 2024 2112 Views
Abstract
Virus induced gene silencing (VIGS) is one of the most potent reverse genetics technologies for gene functional characterisation. This method exploits a dsRNA-mediated antiviral defence mechanism in plants. Using this method allows researchers to generate rapid phenotypic data in a relatively rapid time frame as compared to the generation of stable transformants. Here we describe a simple method for silencing a target gene in barley seedling leaves using vectors based on the Barley Stripe Mosaic Virus (BSMV).
Materials and Reagents
- Bacterial strain: Escherichia coli strain DH5 α
- Taq DNA polymerase (Life Technologies, InvitrogenTM, catalog number: 11304-011 )
- pGEM-T easy cloning kit (Promega Corporation, catalog number: A1380 )
- QIAGEN QIAprep Centrifuge Miniprep Kit (QIAGEN, catalog number: 27106 )
- Recombinant Taq DNA polymerase (Life Technologies, InvitrogenTM, catalog number: 10342-020 )
- Plasmid vectors: pα42 (pα), pβ42.sp1 (pβ), pSL038-1 (pγ), pSL038-PDS (pγ-PDS).
Note: See Scofield and Brandt (2012) for the plasmid maps (provided by Prof. Steven R. Scofield, https://link.springer.com/protocol/10.1007/978-1-61779-882-5_7/figures/1). - Luria-Bertani (LB) liquid broth (Sigma-Aldrich L3152) and solid media containing 1.2% (w/v) agar (OXOID, catalog number: LP0013 )
- Ampicillin (100 mg/l working concentration) (Sigma-Aldrich, catalog number: A0166-5G )
- Restriction enzymes:
PacI (New England Biolabs, catalog number: R0547S )
MluI (New England Biolabs, catalog number: R0198S )
SpeI (New England Biolabs, catalog number: R0133S ) - Ethanol (Sigma-Aldrich, catalog number: 459844 )
- sodium acetate (Sigma-Aldrich, catalog number: S2889 )
- mMessage mMachine T7 in vitro transcription kit (Ambion Sigma-Aldrich, catalog number: AM1344 )
- John Innes compost No 2 (Westland Horticulture)
- Glycine (Sigma-Aldrich, catalog number: 410225 )
- K2HPO4 dibasic (Sigma-Aldrich, catalog number: P3786 )
- Sodium pyrophosphate decahydrate (Sigma-Aldrich, catalog number: 221368 )
- Bentonite (Aldrich, catalog number: 28,523-4 )
- Celite (Fluka, catalog number: 22141 )
- 5x GP buffer (see Recipes)
- FES buffer (see Recipes)
Equipment
- Plant growth chambers (24 °C, 16/8 photoperiod and 55% humidity) (CambridgeHOK containment glasshouse)
- Sterile culture tubes (Falcon, catalog number: 352057 )
- Centrifuge tubes (SARSTEDT AG, catalog number: 72.695.500 )
- Filter paper (Whatman, catalog number: 1001-090 )
- Cooled centrifuge (Eppendorf)
- Shaking incubators for cultures (New Brunswick Scientific)
- Gel apparatus (Helixx Mupid-exU) and image system (Fusion Fx vilber lourmat)
- Thermal cycler (MJ Research PTC200)
- Gel electrophoresis chamber (Helixx Mupid-exU)
- Autoclave (Priorclave)
Software
- Primer3 software (version 0.4.0; http://frodo.wi.mit.edu/primer3/)
Procedure
- Seed germination and plant growth
- Place barley seeds in a 9-cm diameter petri plate containing 2 pieces of filter paper (90 mm diameter) and 6ml of sterile water.
- Cover the petri plates with aluminum foil and stratify in the dark for 2 days at 4 °C.
- Transfer plates to 21 °C in dark for 2 days to germinate seeds.
- Transplant etiolated seedlings to 3-inch pots containing John Innes compost No 2 at a density of 2 seedlings/pot.
- Grow the plants at 22 °C under long day conditions (16 h/8 h), bottom watering every second day. Relative humidity was maintained at 70%.
- Place barley seeds in a 9-cm diameter petri plate containing 2 pieces of filter paper (90 mm diameter) and 6ml of sterile water.
- Virus-induced gene silencing
Note: BSMV is a single-stranded RNA virus with three genome components termed α, β, and γ RNA. A transcribed sequence representing a fragment of the gene to be silenced is inserted immediately downstream of the termination codon of the γ open reading frame that is encoded within a DNA plasmid. RNA is synthesized from α, β, and γ encoded within DNA plasmids. Barley seedlings are infected with all three RNA fragments leads to synthesis of viral dsRNA, which activates the anti-viral RNA silencing pathway, resulting in silencing of the target barley gene. Here we showed VIGS of barley Brassinosteroid-Insensitive-1 (BRI1) gene using two constructs targeting non-overlapping gene fragments. A BSMV γ RNA construct containing a 185 bp-fragment of the barley phytoene desaturase (PDS) gene that protects chlorophyll from photo-bleaching was used as a positive control for the VIGS experiment.- Two independent fragments of BRI1 gene were amplified from genomic DNA of barley cv. Akashinriki using the fragment-specific primers HvBri1A-F/R or HvBri1B-F/R (Table 1). PCR was undertaken in 10 µl volume consisting 1 µl 10X high fidelity PCR buffer, 0.4 µl 50 mM MgSO4, 0.2 µl 10 mM dNTP Mix, 0.2 µl each of 5 μM forward and reverse primer, 0.05 µl 5 U/μl platinum Taq DNA polymerase, 30 ng genomic DNA and sterile Milli-Q H2O to 10 µl. PCR reactions were conducted in a Peltier thermal cycler DNA engine and the PCR program consisted of an initial denaturation step at 94 °C for 2 min, 30 cycles of denaturation 94 °C for 30 sec, annealing at 60 °C for 30 sec, extension at 68 °C for 45 sec and a final extension step at 72 °C for 5 min
Table 1. Primers used for VIGS of BRI1 geneVIGS primer name Forward primer (5΄- 3΄) Reverse primer (5΄- 3΄) HvBRI1:A CGATTAATTAAGCGGAGGCAGAAGAATGA CGACCCGGGGTCACCCTGGCCACTCAC HvBRI1:B CGATTAATTAAGTGAGTGGCCAGGGTGAC CGACCCGGGTGGATGATGTGCGGAATG HvBRi1:RT CAACGATGCTCAAGGTGATG CCGGTGGTCATCTTCCTAAT HvRNAH GCACAGGGAATCGTCAAAGT TCAAAACAACACAACATCGAAGT pGamma TGATGATTCTTCTTCCGTTGC TGGTTTCCAATTCAGGCATCG
Primers were design using the Primer3 software (version 0.4.0; http://frodo.wi.mit.edu/primer3/).
*Note: When designing the primers for gene silencing makes sure that the PCR products will not contain restriction sites for the enzymes used to linearize the plasmids at a later stage. - The amplified gene fragments were cloned into the pGEM-T vector using the pGEM-T Easy cloning kit and were transformed in to E. Coli DH5α via electroporation.
- The recombinant pGEM-T vectors carrying the silencing fragments were then digested with PacI and SmaI. The inserts were purified by gel extraction and then cloned into PacI and SmaI -digested γ RNA vector pSL038-1.
- The resulting recombinant pSL038-1-BRI1A (pγ-BRI1A) and pSL038-1-BRI1B (pγ-BRI1B) plasmids harbouring the BRI1 gene fragments were transformed in to E. coli DH5α via electroporation and cloned products were subsequently sequenced by Macrogen Inc. (Korea) using the vector-specific primers pGamma-F/R (Table 1) to check the orientation of silencing fragment. Positive clones with correct orientations were stored at -80 °C as glycerol stocks.
- E. coli glycerol stocks carrying plasmid pα, pβ, pγ, pγ-PDS, pγ-BRI1A and pγ-BRI1B were streaked on LB-agar plates supplemented with 100 mg/L ampicillin and cultures grown overnight at 37 °C.
- A single colony from each plate was inoculated into 10 ml LB broth supplemented with 100 mg/L ampicillin and cultures were grown overnight at 37 °C.
- Plasmid extraction was carried out using the QIAGEN QIAprep Centrifuge Miniprep Kit as per the product protocol, but excluding RNaseA from buffer P1 (any residual RNaseA carried over in the plasmid DNA preparation will interfere with in vitro transcription at later stages).
- The plasmids pα, pγ, pγ-PDS, pγ-BRI1A and pγ-BRI1B were linearized with MluI and the plasmid pβ was linearized with SpeI. The linearized plasmids were precipitated overnight using ½ volume 5 M ammonium acetate and 2 volume absolute ethanol followed by centrifuging at 18,000 x g for 15 min. DNA pellet was washed with 70% ethanol, air-dried and resuspended in 50 µl RNase-free water. (For 50 seedlings, 1 µg linearized plasmids of each).
- Capped in vitro transcripts were prepared from the linearized plasmids pα, pβ, pγ, γ-PDS, pγ-BRI1A and pγ-BRI1B using the mMessage mMachine T7 in vitro transcription kit following the manufacturer’s protocol. The prepared capped transcripts were checked on a 1% agarose gel. Any smearing of the smaller band indicates degradation of the RNA transcript. There should be a faint band at approximately 10,000 bp and a bright band at approximately 3,000 bp.
- For each plant, VIGS inoculum was prepared by mixing 9 µl of FES Buffer with 0.35 µl each of the α, β and the relevant γ-based transcript [γ, γ-PDS (positive control), γ-BRI1A or γ-BRI1B]. FES buffer served as a mock treatment.
- The first leaf of 10-day-old seedlings was inoculated by rubbing with 10 µl of transcript mixtures or FES (mock treatment) in between thumb and index finger. Care was taken not to damage the leaf.
- After 14 days plants were assessed for visual symptoms of gene silencing. In the case of PDS-silenced plants, the leaves appeared photo bleached (Figure 1). The BRI1-silenced plants showed dwarfing symptoms, as expected (Figure 2).
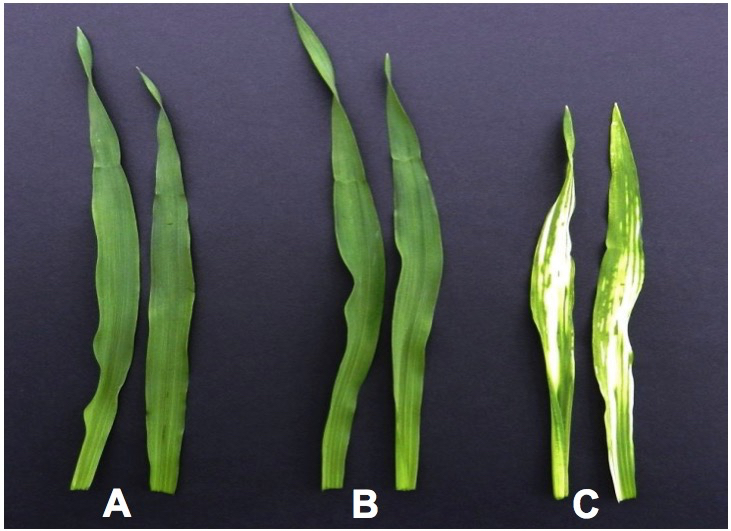

Figure 1. Silencing of PDS gene in barley results in photobleaching and causes the leaves to turn white. A. FES treatment (negative control) B. BSMV:00 (mock treatment) C. BSMV:PDS (positive control).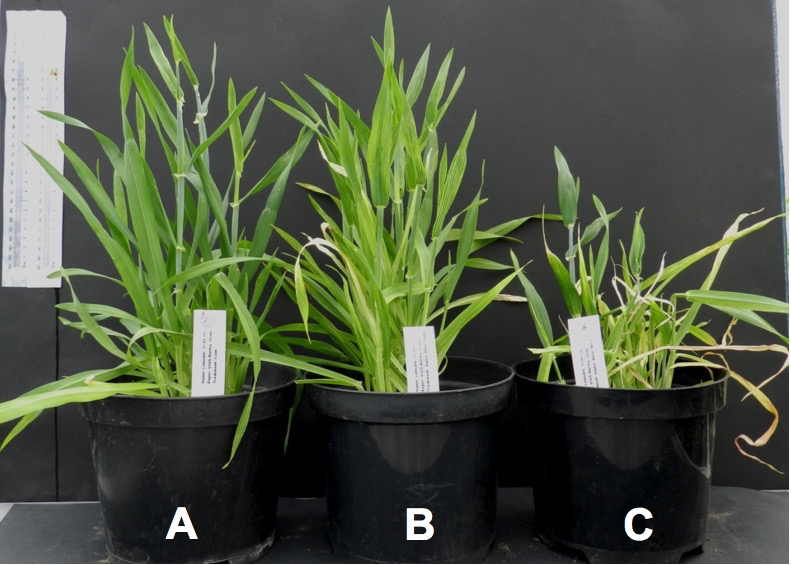
Figure 2. Silencing of HvBRI1 resulted in stunting of plant growth in barley plants. A. FES treatment (negative control) B. BSMV:00 (mock treatment) C. BSMV:HvBRI1. - To confirm the silencing, the 3rd leaf was harvested and flash-frozen in liquid N2 and stored at -70 °C prior to RNA extraction. Total RNA was extracted from plants using the TriZolTM protocol as given by the manufacturer.
- Gene silencing was quantified by real-time RT-PCR using primers specific to BRI1 (HvBRi1:RT, Table 1) and relative to that of the RNA helicase housekeeping gene (HvRNAH, Table 1). Note: It is important to design the real-time primers outside the gene fragment that was cloned into the pγ plasmid.
- Two independent fragments of BRI1 gene were amplified from genomic DNA of barley cv. Akashinriki using the fragment-specific primers HvBri1A-F/R or HvBri1B-F/R (Table 1). PCR was undertaken in 10 µl volume consisting 1 µl 10X high fidelity PCR buffer, 0.4 µl 50 mM MgSO4, 0.2 µl 10 mM dNTP Mix, 0.2 µl each of 5 μM forward and reverse primer, 0.05 µl 5 U/μl platinum Taq DNA polymerase, 30 ng genomic DNA and sterile Milli-Q H2O to 10 µl. PCR reactions were conducted in a Peltier thermal cycler DNA engine and the PCR program consisted of an initial denaturation step at 94 °C for 2 min, 30 cycles of denaturation 94 °C for 30 sec, annealing at 60 °C for 30 sec, extension at 68 °C for 45 sec and a final extension step at 72 °C for 5 min
Notes
- Precaution should be taken not to mix the γ–PDS with other γ samples.
- Gloves must be changed for every treatment
- When applying the transcripts to the leaf, be gentle by not damaging the leaf too much.
Recipes
- 5x GP buffer (500 ml)
18.77 g glycine
26.13 g K2HPO4 dibasic
Bring to 500 ml with ddH2O and use immediately - FES buffer (500 ml)
100 ml GP buffer
5 g sodium pyrophosphate decahydrate
5 g bentonite
5 g celite
Bring to 500 ml with ddH2O
Aliquot into 50 ml volumes and autoclave (121 °C for 20 min)
Aliquots can be stored at room temperature under sterile condition
Acknowledgments
This work was supported by the Science Foundation Ireland research fund (IN10/IN.1/B3028) and Department of Agriculture Research Stimulus Grant RSF 07 513. Part of the procedures were adapted from a previously described methods by Ali et al. (2014), Holzberg et al. (2002) and Scofield et al. (2005).
References
- Ali, S. S., Gunupuru, L. R., Kumar, G. B., Khan, M., Scofield, S., Nicholson, P. and Doohan, F. M. (2014). Plant disease resistance is augmented in uzu barley lines modified in the brassinosteroid receptor BRI1. BMC Plant Biol 14: 227.
- Holzberg, S., Brosio, P., Gross, C. and Pogue, G. P. (2002). Barley stripe mosaic virus-induced gene silencing in a monocot plant. Plant J 30(3): 315-327.
- Scofield, S. R. and Brandt, A. S. (2012). Virus-induced gene silencing in hexaploid wheat using Barley stripe mosaic virus vectors. In: Watson, J. M. and Wang, M. B. (eds). Antiviral Resistance in Plants: Methods and protocols. Springer, 93-112.
- Scofield, S. R., Huang, L., Brandt, A. S. and Gill, B. S. (2005). Development of a virus-induced gene-silencing system for hexaploid wheat and its use in functional analysis of the Lr21-mediated leaf rust resistance pathway. Plant Physiol 138(4): 2165-2173.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Gunupuru, L. R., Ali, S. S., Doohan, F. M. and Scofield, S. R. (2015). Virus-induced Gene Silencing (VIGS) in Barley Seedling Leaves. Bio-protocol 5(12): e1506. DOI: 10.21769/BioProtoc.1506.
Category
Plant Science > Plant molecular biology > RNA > Transcription
Plant Science > Plant immunity > Disease bioassay
Molecular Biology > RNA > RNA interference
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










