- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Lectin Binding Analysis of Streptococcus mutans Glycoproteins
Published: Vol 5, Iss 7, Apr 5, 2015 DOI: 10.21769/BioProtoc.1431 Views: 9577
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Characterization of Protein Domain Function via in vitro DNA Shuffling
Kathy Hiu Laam Po [...] Sheng Chen
Jun 5, 2018 6647 Views
In vitro and in vivo Assessment of Protein Acetylation Status in Mycobacteria
Krishna K. Singh [...] Deepak K. Saini
Jul 5, 2019 5683 Views
In vitro Glutamylation Inhibition of Ubiquitin Modification and Phosphoribosyl-Ubiquitin Ligation Mediated by Legionella pneumophila Effectors
Alan G. Sulpizio [...] Yuxin Mao
Nov 5, 2020 3947 Views
Abstract
Bacterial glycoproteins are of increasing interest due to their abundance in nature and importance in health and infectious diseases. However, only a very small fraction of bacterial glycoproteins have been characterized and its post-translational modification machinery identified. While analysis of glycoproteins can be achieved through various techniques, this is often limited by the specific characteristics of individual proteins such as type and level of glycosylation. Lectins are sugar-binding proteins that recognize specific glycoconjugates in a manner similar to antigen-antibody interactions. Here, we describe a simple method for the detection of glycoproteins using lectin-based Western blot analysis, which can be applied to different organisms and coupled with various other strategies for complementary analysis.
Materials and Reagents
- Bacterial whole cell lysates
- Brain heart infusion medium (BHI) (BD Biosciences, catalog number: 237500 )
- Biotinylated lectin [Vector Laboratories, for Wheat Germ Agglutinin (WGA), catalog number: B-1025]
- HRP-conjugated streptavidin (Cell Signaling Technology, catalog number: 3999 )
- Tween 20 (Sigma-Aldrich, catalog number: P2287 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A9418 )
- Enhanced chemiluminiscent detection kit (GE Healthcare, catalog number: RPN2108 )
- Autoradiography films (Carestream Health, catalog number: 864-6770 )
- NaCl (J.T.Baker®, catalog number: 3628-01 )
- KCl (J.T.Baker®, catalog number: 3045-01 )
- Na2HPO4 (J.T.Baker®, catalog number: 3827-01 )
- KH2PO4 (J.T.Baker®, catalog number: 3246-01 )
- Tris base (J.T.Baker®, catalog number: 4109-02 )
- Sodium dodecyl sulfate (SDS) (Sigma-Aldrich, catalog number: L3771 )
- Glycine (Thermo Fisher Scientific, catalog number: BP381-1 )
- 30% Acrylamide/Bis solution (Bio-Rad Laboratories, catalog number: 161-0158 )
- Tetramethylethylenediamine (TEMED) (Life Technologies, catalog number: 15524-010 )
- Ammonium persulfate (Bio-Rad Laboratories, catalog number: BP179-25 )
- Glycerol (Alfa Aesar, catalog number: A16205 )
- Methanol (BDH Chemicals, catalog number: BDH1135-4LP )
- Bromophenol blue (Sigma-Aldrich, catalog number: B5525 )
- 1x phosphate buffer solution (PBS) (see Recipes)
- Resolving gel (see Recipes)
- Stacking gel (see Recipes)
- 2x sample buffer (see Recipes)
- Running buffer (see Recipes)
- Transfer buffer (see Recipes)
- Blocking solution (see Recipes)
- Primary solution (see Recipes)
- Secondary solution (see Recipes)
- Washing solution (see Recipes)
Equipment
- 0.1 mm glass beads (Research Products International)
- Polyvinylidene fluoride (PVDF) membranes (EMD Millipore, catalog number: P2563-10EA )
- 3 mm Whatman paper (GE Healthcare)
- CO2 Incubator
- Sterile culture tubes (15 ml)
- Screw cap tubes (1.7 ml)
- Bead beater (BioSpec Products)
- Protein electrophoresis running apparatus
- Protein transfer apparatus
- Power supply
- Rocker
- Film developer
- pH meter
Procedure
- Preparation of whole cell protein lysates
- Grow bacterial cells [Streptococcus mutans (S. mutans) OMZ175] in 10 ml BHI broth at 37 °C in a humidified 5% CO2 atmosphere.
- After 18 h, stationary phase cultures (OD600~0.9) are washed twice (2x) by centrifuging at 4,000 rpm for 10 min, discarding supernatants and resuspending pelleted cells with 1x PBS in original culture volume. After the second wash, cells are concentrated 5x by adding five times less the original culture volume of 1x PBS. The concentrated suspension is then transferred to a 1.7 ml screw cap tube containing approximately 500 μl of glass beads.
- Whole cell protein lysates are obtained using a bead-beater at maximum setting (3,450 oscillations per min) and the suspensions are homogenized for three cycles of 30 sec with 2 min of incubation on ice between cycles.
- Alternative mechanical methods to obtain whole cell protein lysates such as sonication or French-press, with or without freeze-thawing cycles, are also acceptable.
- Grow bacterial cells [Streptococcus mutans (S. mutans) OMZ175] in 10 ml BHI broth at 37 °C in a humidified 5% CO2 atmosphere.
- Protein lysates are separated on a 10% SDS-PAGE and subsequently transferred to a PVDF membrane.
- For gel preparation, the cassette is assembled with glass and silica plates with 0.75-1.0 mm spacers and placed on a caster.
- Resolving gel is prepared, poured and let to polymerize for approximately 30 min.
- Stacking gel is prepared, poured and a comb with desired number of wells inserted. Allow gel to polymerize (~ 30 min).
- Remove cassette from caster and place on running apparatus.
- Prepare samples by mixing equal volume of sample and 2x sample buffer and boil for 10 min. Incubate on ice for 1 min, quick spin tube to settle any sediment and load sample on gel.
- Run SDS-PAGE at 100 V for 2 h.
- Transfer “sandwich” is assembled (Figure 1) in the presence of transfer buffer. Gel is removed from plates and placed in assembling “sandwich”.
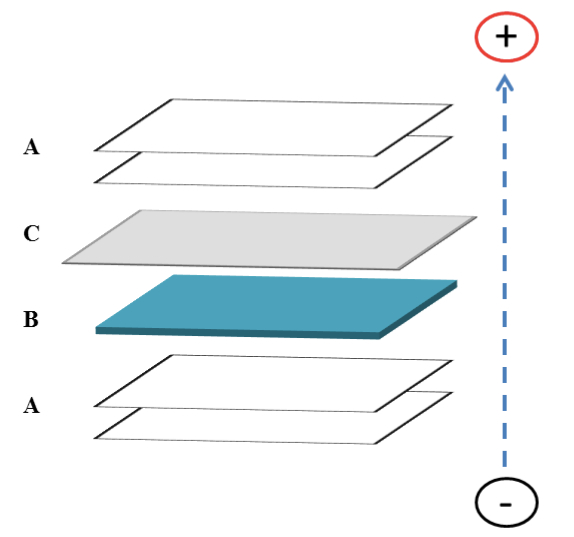
Figure 1. Schematic of transfer “sandwich” assembling for Western blotting. In the presence of transfer buffer, place two (2) pieces of Whatman paper A soaked in transfer buffer followed by the gel with separated proteins B, the PVDF membrane C soaked in methanol then transfer buffer, and two (2) more pieces of soaked Whatman papers. Arrow shows the direction of protein transfer.
- An oversized piece of PVDF membrane, to cover the entire gel, is soaked in methanol first followed by transfer buffer and then placed on top of the gel.
- The rest of the transfer “sandwich” is completed and proteins are transferred to the membrane for 1 h at 100 V in the presence of transfer buffer and an ice unit block to absorb heat produced during transfer.
- Sandwich is disassembled, membrane is removed and any extra piece of membrane that is bigger than the gel is cut out.
- For gel preparation, the cassette is assembled with glass and silica plates with 0.75-1.0 mm spacers and placed on a caster.
- Block membrane with 5% BSA solution for 18-20 h at 4 °C with slow rocking. Wash membrane 3x by adding 20 ml of washing solution and slow rocking for 5 min to remove excess BSA. Then, incubate the membrane with the primary solution (biotinylated lectin in 0.5% BSA) at room temperature for 1 h with slow rocking.
- Wash membrane 3x followed by the addition of the secondary solution (HRP-streptavidin in 0.5% BSA) and incubate at room temperature for 1 h with slow rocking.
- After incubation with secondary solution, wash the membrane 4x to visualize biotinylated lectins bound to glycoproteins using enhanced chemiluminescent detection kit, mix equal volumes of detection solution 1 and detection solution 2 (for a final volume of 0.125 ml per cm2 of membrane) and incubate for 1 min at room temperature.
- Drain excess detection solution and place membrane in an autoradiography cassette for film exposure. Due to the variability of carbohydrate content on glycoproteins, different exposure of the membrane to the film should be performed.
Representative data
A representative image of Cnm analysis for different strains of S. mutans with anti-rCnmA and biotinylated-WGA is shown below.
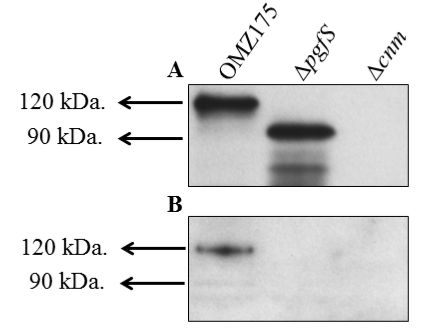
Figure 2. Example of lectin binding analysis of S. mutans. Cnm is a surface protein of invasive S. mutans that undergoes PgfS-dependent glycosylation (Aviles-Reyes et al., 2014). A. Size difference in Cnm due to PgfS modification can be observed using anti-rCnmA. B. The presence of glycoconjugates attached to Cnm in OMZ175 was determined using biotinylated wheat germ agglutinin.
Notes
- Due to the variability of the type and the level of glycosylation for each protein, protocol optimization may be required for successful identification of glycoproteins by lectin analysis.
- Control glycoproteins with known reactivity to specific lectins should be included in each experiment.
Recipes
- 1x PBS
137 mM NaCl
2.7 mM KCl
10 mM Na2HPO4
2 mM KH2PO4
- Resolving gel (10 % Tris glycine SDS-PAGE; 10 ml)
4.0 ml of H2O
3.3 ml of 30% acrylamide
2.5 ml of 1.5 M Tris (pH 8.8)
0.1 ml of 10% SDS
0.1 ml of 10% APS
8 μl of TEMED
- Stacking gel (3 ml)
2.1 ml of H2O
0.5 ml of 30% acrylamide
0.38 ml of 0.5 M Tris (pH 6.8)
0.03 ml of 10% SDS
0.03 ml of 10% APS
6 μl of TEMED
- 2x sample buffer (200 ml)
50 ml of 0.5 M Tris (pH 6.8)
8 g of SDS
40 ml of glycerol
H2O to 200 ml
Add bromophenol blue until dark
- Running buffer
25 mM Tris
192 mM glycine
0.1% SDS
pH 8.3
- Transfer buffer (1 L)
200 ml of methanol
3.0 g of Tris
14.4 g of glycine
H2O to 1 L
- Blocking solution
5% BSA (w/v) in 1x PBS
- Primary solution
0.5% BSA (w/v) in 1x PBS
0.1% Tween 20
20 μg/ml biotinylated lectin
- Secondary solution
0.5% BSA (w/v) in 1x PBS
0.1% Tween 20
- Washing solution
1x PBS + 0.1% Tween 20
Acknowledgments
We would like to thank Fred Hagen for technical support. This work has been supported by grants from the American Heart Association (10GRNT420049) and NIH-NIDCR (R01 DE022559). A.A.-R. was supported by NIH-NHLBI (F31 HL124951).
References
- Avilés-Reyes, A., Miller, J. H., Simpson-Haidaris, P. J., Hagen, F. K., Abranches, J. and Lemos, J. A. (2014). Modification of Streptococcus mutans Cnm by PgfS contributes to adhesion, endothelial cell invasion, and virulence. J Bacteriol 196(15): 2789-2797.
- Nothaft, H. and Szymanski, C. M. (2010). Protein glycosylation in bacteria: sweeter than ever. Nat Rev Microbiol 8(11): 765-778.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Avilés-Reyes, A., Lemos, J. A. and Abranches, J. (2015). Lectin Binding Analysis of Streptococcus mutans Glycoproteins. Bio-protocol 5(7): e1431. DOI: 10.21769/BioProtoc.1431.
Category
Microbiology > Microbial biochemistry > Protein > Modification
Biochemistry > Protein > Immunodetection > Western blot
Biochemistry > Carbohydrate > Glycoprotein
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













