- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
RNA Isolation from Synechocystis
Published: Vol 5, Iss 6, Mar 20, 2015 DOI: 10.21769/BioProtoc.1428 Views: 24178
Reviewed by: Maria SinetovaClaudia CatalanottiAksiniya Asenova

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Purification of Bacterial RNA from Infected Macrophages
Lior Lobel [...] Anat A. Herskovits
Nov 20, 2015 11219 Views
Single Genome Sequencing of Expressed and Proviral HIV-1 Envelope Glycoprotein 120 (gp120) and nef Genes
David J. Nolan [...] Michael S. McGrath
Jun 20, 2017 10053 Views
Abstract
The protocol describes the procedure of total RNA isolation from cells of the cyanobacterium Synechocystis sp. PCC 6803. This protocol is also applicable to Synechococcus elongatus PCC 7942 and PCC 6301, Thermosynechococcus vulcanus, and other unicellular and filamentous species of cyanobacteria that do not have thick polysaccharide-containing outer layers. For the latter, Trizol-containing protocols should be adapted. The yield of RNA depends on optical density of cyanobacterial culture and may reach up to 10-20 µg of total RNA per 1 ml of cell culture. RNA isolated by this method can be used for Northern blot hybridization, RT-qPCR, microarrays and Next Generation Sequencing.
Keywords: SynechocystisMaterials and Reagents
- Cyanobacterial culture (50 ml) (OD750 ~ 2) grown in BG11 medium (Rippka, 1988)
- 95% ethanol for molecular biology (keep at -20 °C)
- 70% ethanol for molecular biology (30 ml of RNAse free water and 70 ml 95% ethanol, keep at -20 °C)
- 70% ethanol for sterilization
- Phenol BioUltra (for molecular biology, TE-saturated, ~73%) (Sigma-Aldrich, catalog number: 77607 ) (keep at 4 °C)
- 1 M Tris-HCl (Life Technologies, catalog number: 15568-025 )
- 0.5 M EDTA-Na2 (pH 8.0) (Life Technologies, catalog number: AM9260G )
- RNase-free autoclaved water
- Chloroform (Sigma-Aldrich, catalog number: C2432 ) (keep at 4 °C)
- 10 M lithium chloride solution (Fluka, catalog number: 83268 ) (chill on ice before use)
- RNAse-free DNAse
- Agarose for molecular biology (Sigma-Aldrich, catalog number: A9539 )
- TAE electrophoresis buffer (Sigma-Aldrich, catalog number: T9650 )
- RNA ladder (Life Technologies, InvitrogenTM, catalog number 15620-016 )
- TE buffer (see Recipes)
- 50/100 TE buffer (see Recipes)
- 10 M LiCl solution in water (see Recipes)
- Cell fix solution (see Recipes)
Equipment
- Saran wrap (any type)
- Autoclave bag (Sigma-Aldrich, catalog number: Z692212 )
- Safe-Lock tubes (2.0 ml) (Eppendorf, catalog number: 0030 120.094 )
- 50 ml plastic conical polypropylene tubes (sterile, nuclease-free, nonpyrogenic, autoclavable) (Life Technologies, catalog number: AM12501 or Corning, Falcon )
- Autoclave
- Refrigerated centrifuge with bucket rotor that fits 50 ml conical tubes (Eppendorf, model: 5804 R and Swing-bucket rotor A-4-44 with 50 ml Falcon adapters)
- Refrigerated microcentrifuge preset to 4 °C (Eppendorf, model: 5415 R )
- Water bath preset to 65 °C
- Subzero refrigerators: -20 °C and -70 °C
- Fume hood
- Ice bath
- Multi tube automated vortex (for example, Micro tube mixer MT-400, Tomy Digital Biology)
- UV-spectrophotometer (preferably, Nanodrop)
- Agarose gel electrophoresis equipment (VWR Mini Gel II Complete Horizontal Electrophoresis System, catalog number: 95043-650)
Note: Workplace, pipettes, gloves etc. should be cleaned with 70% ethanol before RNA isolation procedures. Conical tubes, Eppendorf tubes, pipette tips should be sterilized by autoclaving in autoclave bags before use.
Procedure
- Probes fixation
- Prepare 25 ml portions of cell fix solution in 50 ml plastic conical polypropylene tubes. Keep on ice. Pour 25 ml of cyanobacterial cell culture into 25 ml of ice-cold cell fix solution (Figure 1.1). Stir well and proceed to centrifugation or store samples at -20 °C up to 1 month. For 50-ml of cyanobacterial cultures, two 50 ml tubes should be used. Alternatively, 100-200 ml centrifuge bottles resistant to phenol and ethanol may be used.
- Harvest fixed cells by centrifugation at 3,000 x g for 5-10 min at 4 °C (Figure 1.2).
- Discard cell fix solution leaving 0,5-1 ml of solution to resuspend the pellet. Resuspend the pellet by vortexing or pipetting.
- Transfer a suspension into a sterile 2 ml Eppendorf tube. If 50 ml tubes are used for cell fixation and harvesting, combine two probes from 25 ml tubes in one 2 ml Eppendorf tube.
- Centrifuge at 3,000-5,000 x g, 4 °C for 3-5 min and completely remove supernatant (Figure 1.3). Leave pelleted cells on ice.
Note: Probes fixation is applied to kill cells immediately and to avoid possible degradation of RNA. This is extremely important when temperature-dependent changes in specific mRNAs are under study.
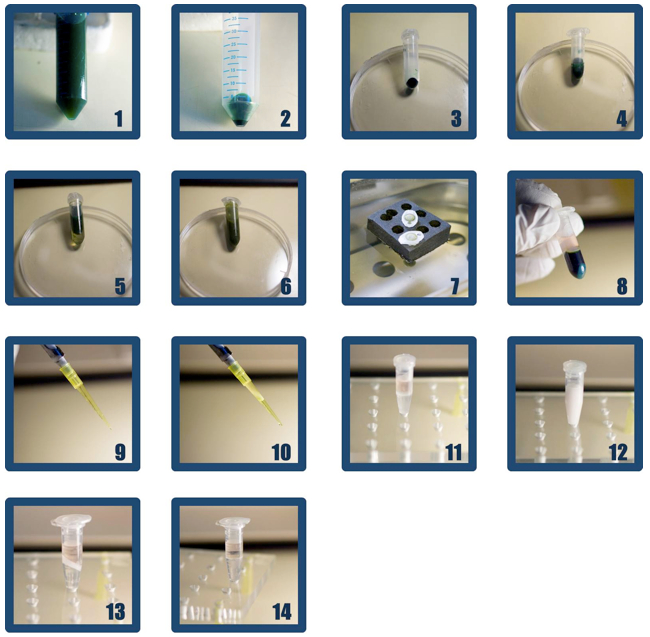

Figure 1. Steps of RNA extraction. 1. Fixation in Cell Fix Solution. 2. Fixed cells after centrifugation. 3. Cell pellet at the bottom of the tube. 4. Cells resuspended in 500 µL of TE-50/100 5. Cell suspension after addition of 1ml of phenol. 6. Cell suspension after vortexing. 7. Incubation in 65°C water bath. 8. Four phases after centrifugation (from bottom to top): blue pellet (if it is green, hot phenol cell lysis was not complete; in this case resuspend again and start from step A4); dark green phenol phase; protein interphase (might be invisible); pink TE-50/100 phase with nucleic acids. 9. Carefully transfer upper TE-50/100 phase into new tube. Use yellow tip. 10. Do not take interphase and phenol phase into the tip. 11. Addition of 300 µl chloroform and 300 µl phenol to TE-50/100-fraction. 12. Homogenous white colored emulsion after vortexing. 13. Three phases after centrifugation (from bottom to top): phenol/chloroform (lower phase); protein interphase; pink TE-50/100 phase (upper phase). This phase should be transferred into a fresh 1.5 ml Eppendorf tube. 14. Repeat steps A9-13 until interphase disappears.
- Prepare 25 ml portions of cell fix solution in 50 ml plastic conical polypropylene tubes. Keep on ice. Pour 25 ml of cyanobacterial cell culture into 25 ml of ice-cold cell fix solution (Figure 1.1). Stir well and proceed to centrifugation or store samples at -20 °C up to 1 month. For 50-ml of cyanobacterial cultures, two 50 ml tubes should be used. Alternatively, 100-200 ml centrifuge bottles resistant to phenol and ethanol may be used.
- Cell lysis by hot phenol
- Resuspend the pellet in 500 µl of sterile ice-cold 50/100 TE-buffer by vortexing or pipetting (Figure 1.4).
- Add 1 ml of phenol, vortex and incubate in a water bath preheated at 65 °C for 10 min (Figure 1.5-1.7). Regularly shake probes (3-4 times during 10 min of incubation) to prevent phenol phase separation.
- Centrifuge samples at top speed (16,000 x g) at 15-20 °C for 10 min. The pellet in the bottom phase should be blue-colored meaning cell lysis is complete (Figure 1.8).
- Transfer upper aqueous phase to a new Eppendorf tube (Figure 1.9-1.10).
- Resuspend the pellet in 500 µl of sterile ice-cold 50/100 TE-buffer by vortexing or pipetting (Figure 1.4).
- Purification of nucleic acids by phenol/chloroform
- Add 300 µl of TE-saturated phenol and 300 µl of chloroform to the probe (collected aqueous phase) (Figure 1.11).
- Vortex for 1-2 min and centrifuge at 16,000 x g, 4 °C for 5 min (Figure 1.12-1.13). For multiple samples apply multi tube automated vortex (for example, Micro tube mixer MT-400).
- Transfer upper aqueous phase to a new tube.
- Repeat this procedure 2-3 times until the interphase (a whitish, denatured protein-containing layer between phenol phase and aqueous phase that contains nucleic acid) disappears completely (Figure 1.14).
- Add 300 µl of TE-saturated phenol and 300 µl of chloroform to the probe (collected aqueous phase) (Figure 1.11).
- Precipitation of nucleic acids
- Add 2-3 volumes of 95% ethanol prechilled at -20-30 °C to a final aqueous phase (~ 1,5 ml of ethanol to ~ 0.5 ml of a probe), stir well and incubate at -20 °C from 2-3 h to overnight.
- At this stage a probe can be stored for several months under ethanol at -20 °C.
- Centrifuge at top speed, 4 °C for 10 min, carefully discard liquid.
- Wash the pellet of nucleic acids with 1 ml of 70% cold ethanol by intensive vortexing.
- Centrifuge at top speed, 4 °C for 5-15 min and completely remove ethanol by aspiration.
- It is important to carry on all operations with RNA on ice.
- Add 2-3 volumes of 95% ethanol prechilled at -20-30 °C to a final aqueous phase (~ 1,5 ml of ethanol to ~ 0.5 ml of a probe), stir well and incubate at -20 °C from 2-3 h to overnight.
- Separation of RNA from DNA by LiCl precipitation
- Add 800 µl of 50/100 TE buffer to the pellet. Incubate for 15 min on ice.
- Vortex probe 3-4 times to dissolve the pellet completely.
- Add 200 µl of 10 M LiCl solution and mix by vortexing. Leave probes on ice for 2 h.
- Centrifuge at top speed, 4 °C, for 15-20 min.
- Carefully remove supernatant by aspiration.
- Wash the RNA preparation by adding 1 ml of ice-cold 70% ethanol and intensive vortexing of the pellet. Centrifuge at top speed, 4 °C, for 15 min.
- Repeat 70% ethanol washing.
- Carefully remove all ethanol and air-dry the probe(s) at room temperature in a tube rack covered by Saran Wrap. If vacuum drier is used, please, take care not to overdry the pellet, because it will become impossible to solubilize it. Dissolve RNA in 50-100 µl of RNAse-free water.
- Add 800 µl of 50/100 TE buffer to the pellet. Incubate for 15 min on ice.
- Evaluation of RNA quantity and quality
- Take 1-2 µl of RNA and check its concentration with Nanodrop spectrophotometer.
- LiCl does not allow to obtain RNA that are completely free from DNA, but the quality of RNA is usually enough for nothern blot analysis. If necessary, purify RNA sample(s) additionally with DNAse according to the manufacturer instructions.
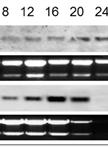
- Check RNA quality by agarose gel electrophoresis. Electrophoretically resolved total cyanobacterial RNA is visible as 3 major bands that correspond to 23S, 16S and 5S RNAs. Smear on lines indicate RNA was degraded during steps of isolation (Figure 2).

Figure 2. Total RNA samples isolated from Synechocystis sp. PCC 6803 at 1 µg per lane were separated by electrophoresis in 1% non-denaturing agarose gel and stained with ethidium bromide. Typically 3 major (or more) bands corresponded to 23S, 16S and 5S RNAs should be seen. Lanes 1-3 represent isolated total RNA of good quality, and lanes 4-6 with smeared bands demonstrate low quality partly degraded RNA. For northern blotting, RNA should be separated in denaturing formaldehyde-containing 1.2% agarose gel, and transferred onto nitrocellulose or nylon membrane (lanes 7, 8). It is recommended to run two extra lanes of RNA sample (lane 7) and RNA ladder (lane 8), which will be cut off and stained with methylene blue to determine the size and quality of RNA after blotting.
- Take 1-2 µl of RNA and check its concentration with Nanodrop spectrophotometer.
Notes
Solution of LiCl contains DNA, and it’s important to remove supernatant completely. Also DNA in LiCl solution can be precipitated by 2 volumes of ethanol or 0.8 volumes of 2-isopropyl alcohol (Lab Scan, C19C11X).
Recipes
- TE buffer
10 mM Tris-HCl (pH 7.5-8.0)
1 mM EDTA-Na2
- 50/100 TE buffer
50 mM Tris-HCl (pH 7.5-8.0)
100 mM EDTA
After preparation, this buffer should be sterilized by autoclaving at 121 °C, 1.5 atm., 15 min
- 10 M LiCl solution in water
Note: It should be autoclaved, aliquoted in sterile conditions, and stored sterile.
- Cell fix solution
0.5% phenol in 95% ethanol
25-ml aliquots in autoclaved 50-ml tubes should be stored at -20 °C
Acknowledgments
This work was supported by a grant from Russian Science Foundation No. 14-24-00020 to D.A.L. and by a grant from Russian Foundation for Basic Research No. 14-94-01446a to K.S.M.
References
- Kiseleva, L. L., Serebriiskaya, T. S., Horvath, I., Vigh, L., Lyukevich, A. A. and Los, D. A. (2000). Expression of the gene for the delta9 acyl-lipid desaturase in the thermophilic cyanobacterium. J Mol Microbiol Biotechnol 2(3): 331-338.
- Mironov, K. S., Sidorov, R. A., Kreslavski, V. D., Bedbenov, V. S., Tsydendambaev, V. D. and Los, D. A. (2014). Cold-induced gene expression and omega (3) fatty acid unsaturation is controlled by red light in Synechocystis. J Photochem Photobiol B 137: 84-88.
- Rippka, R. (1988). Isolation and purification of cyanobacteria. Methods Enzymol 167: 3-27.
- Sambrook, J., Fritsch, E. F. and Maniatis, T. (1989). Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press.
- Sato, N. (1995). A family of cold-regulated RNA-binding protein genes in the cyanobacterium Anabaena variabilis M3. Nucleic Acids Res 23(12): 2161-2167.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mironov, K. S. and Los, D. A. (2015). RNA Isolation from Synechocystis. Bio-protocol 5(6): e1428. DOI: 10.21769/BioProtoc.1428.
Category
Microbiology > Microbial genetics > RNA > RNA extraction
Molecular Biology > RNA > RNA extraction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link