- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Differentiation of Naturally Produced Extracellular Membrane Vesicles from Lipid Aggregation by Glucuronoxylomannan Immunogold Transmission Electron Microscopy in Bacillus subtilis
Published: Vol 5, Iss 5, Mar 5, 2015 DOI: 10.21769/BioProtoc.1408 Views: 11050
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Extraction of Bacterial Membrane Vesicle and Phage Complex by Density Gradient Ultracentrifugation
Shangru Li [...] Tianyuan Jia
Aug 20, 2024 2091 Views
Mycobacterium smegmatis Ribosome Purification, Co-sedimentation, and Subunit Association Assay
Aneek Banerjee [...] Jayati Sengupta
May 20, 2025 1621 Views
Preparation and Negative Staining for Visualization of Cyanoglobule Lipid Droplets Using Transmission Electron Microscopy
Febri A. Susanto [...] Peter K. Lundquist
Dec 5, 2025 1338 Views
Abstract
Recently, membrane vesicle (MV) production was described in Gram-positive bacteria, which harbor a variety of components such as toxins, antibiotic resistance proteins, proteases, DNA, and immune modulators. Free lipids have the ability to form micelles, thus it is important to rule out spontaneous association of lipids into vesicle-like structures and rather, that MVs are produced naturally by a metabolically active cell. Here, we describe a protocol utilizing the polysaccharide, glucuronoxylomannan (GXM) from Cryptococcus neoformans (C. neoformans) as a marker to differentiate naturally produced MVs from vesicles that form spontaneously in the Gram-positive model organism, Bacillus subtilis (B. subtilis). MVs are purified from bacterial cultures grown in the presence of GXM; MVs naturally produced by cells would not contain GXM in the lumen whereas vesicular structures forming in the media could encapsulate GXM and this can be visualized via immunogold transmission electron microscopy.
Materials and Reagents
- Purified glucuronoxylomannan (GXM) (protocol included)
- YPD broth (Difco, catalog number: 242820 )
- C. neoformans H99 fungal strain
- 100 kDa, 50 kDa, 10 kDa, 1 kDa cut-off EMD Millipore Ultrafiltration Membranes (Millipore, catalog number: 14442AM )
- B. subtilis 168 bacterial strain
- BHI broth (Difco, catalog number: 299070 )
- 8% glutaraldehyde (Polysciences, catalog number: 111-30-8 )
- 16% paraformaldehyde (Electron Microscopy Sciences, catalog number: 15700 )
- 0.5 M sodium cacodylate (pH 7.4) (Electron Microscopy Sciences, catalog number: 11650 )
- 100% ethanol
- Lowicryl HM-20 monostep resin (Electron Microscopy Sciences, catalog number: 14340 )
- Gelatin (Thermo Fisher Scientific)
- Aurion Donkey Block (Electron Microscopy Sciences, catalog number: 25599 )
- α-GXM monoclonal antibody 18B7 (mouse monoclonal IgG1) (Casadevall lab-generated)
- BSA-c (Electron Microscopy Sciences, catalog number: 25557 )
- 10 nm conjugated secondary Ab (donkey α-mouse) (Electron Microscopy Sciences, catalog number: 25814 )
- 4% uranyl acetate (aq) (SPI Supplies, catalog number: 615-44-0 )
- 1x phosphate buffered saline (PBS) (see Recipes)
Equipment
- Express PLUS Membrane Filters - Pore (0.22 μm) (Millipore, catalog number: SCGVU01RE )
- Ultracentrifugation tubes (thickwall, polyallomer/pollypropylene, 3.5 ml, 13 x 51 mm) (Beckman Coulter, catalog number: 349623 )
- 200 mesh nickel grids (Polysciences, catalog number: 24916 )
- Centrifuge capable of 15,000 x g
- TLA 100.3 rotor (Beckman Coulter, catalog number: 349490 )
- Amicon ultrafiltration system (Millipore, catalog number: 5124)
- Optima TL ultracentrifuge (Beckman Coulter, discontinued comparable to B11229)
- Sonicator (sonic dismembrator) (Thermo Fisher Scientific, model: 100 cpn-214-161 )
- Freeze substitution system (RMC, model: FS-7500 )
- Reichert Ultracut UCT Ultramicrotome
- JEOL 100CXII or JEOL 1200EX Electron Microscopes
Procedure
- Purification of GXM
- Grow 1 L of YPD broth inoculated with C. neoformans strain H99 overnight at 37 °C with shaking (~18 h 200 rpm).
- Spin out cells by centrifuging 15,000 x g for 20 min at 4 °C.
- Filter cell-free supernatant with 0.22 μm Express PLUS Membrane Filters to remove debris.
- Sequentially concentrate the cell-free supernatant with 100 kDa, 50 kDa, 10 kDa, and 1 kDa cut-off EMD Millipore Ultrafiltration Membranes. The jelly that forms on the filters is GXM (save jelly that is <10 kDa and >1 kDa) (see Figure 1 for concentrator set-up).
- Grow 1 L of YPD broth inoculated with C. neoformans strain H99 overnight at 37 °C with shaking (~18 h 200 rpm).
- Purification of gram-positive extracellular vesicles
- Inoculate 100 ml of BHI broth with fractionated GXM <10 kDa and B. subtilis strain 168.
- Grow overnight at 37 °C with shaking (~18 h 200 rpm).
- Centrifuge culture to remove cells at 15,000 x g for 20 m at 4 °C.
- Filter cell-free supernatant with 0.22 μm Express PLUS Membrane Filters to remove debris.
- Concentrate cell-free supernatant to small volumes (approximately 6 ml) with ultrafiltration system and 100 kDa cut-off EMD Millipore Ultrafiltration Membranes (Figure 1).
- Ultracentrifuge concentrated supernatant at 100,000 x g for 1 h at 4 °C in thickwall, polyallomer, 3.5 ml, 13 x 51 mm tubes to pellet vesicles (Figure 2).
- Wash/resuspend vesicle pellet with 500 μl PBS and repeat spin (repeat wash twice).
- Remove supernatant from sample without disturbing pellet.
- For sonication (production of disrupted/unnatural vesicles-positive control) resuspend pellet in PBS and sonicate (20 sec sonication and 30 sec incubation at 4 °C on ice, repeated 4 times at power 2).
- Ultracentrifuge again to pellet vesicles and carefully remove supernatant.
- For sonication (production of disrupted/unnatural vesicles-positive control) resuspend pellet in PBS and sonicate (20 sec sonication and 30 sec incubation at 4 °C on ice, repeated 4 times at power 2).
- Carefully add 4% paraformaldehyde, 0.1% glutaraldehyde in 0.1 M sodium cacodylate without disturbing pellet.
- Incubate for 1 h at room temperature (RT).
- Inoculate 100 ml of BHI broth with fractionated GXM <10 kDa and B. subtilis strain 168.
- Sample preparation for immunogold labeling
- After fixing for 1 h at RT, wash with 0.1 M sodium cacodylate 3 times.
- Enrobe samples in 5% gelatin.
- Dehydrate through a graded ethanol series in a RMC FS 7500; progressively lowering the temperature 5 °C /h from 4 °C to -50 °C.
- Use Lowicryl HM-20 monostep resin to embed sample.
- Polymerize embedded sample with UV light for 48 h at -50 °C.
- Use Reichert Ultracut UCT to cut 80 nm ultrathin sections onto 200 mesh nickel grids.
- After fixing for 1 h at RT, wash with 0.1 M sodium cacodylate 3 times.
- Immunogold labeling of GXM in vesicle preparations
- Block with Aurion Donkey Block for 1 h.
- Incubate samples with the primary α-GXM mAb antibody, 18B7, in 0.1% BSA-c/PBS incubation buffer for ~18 h at 4 °C.
- Wash samples with 0.1% BSA-c/PBS.
- Incubate samples with the secondary donkey α-mouse IgG 10 nm immunogold-conjugated antibody for 2 h at RT.
- Wash samples with BSA-C.
- Post-fix samples for 5 m at 25 °C with 2% glutaraldehyde/PBS to stabilize the gold particles.
- Counterstain samples for 15 min 25 °C with 4% uranyl acetate (aq).
- Also make control samples with an irrelevant IgG antibodies and without the 1o antibody (18B7).
- Block with Aurion Donkey Block for 1 h.
Representative data
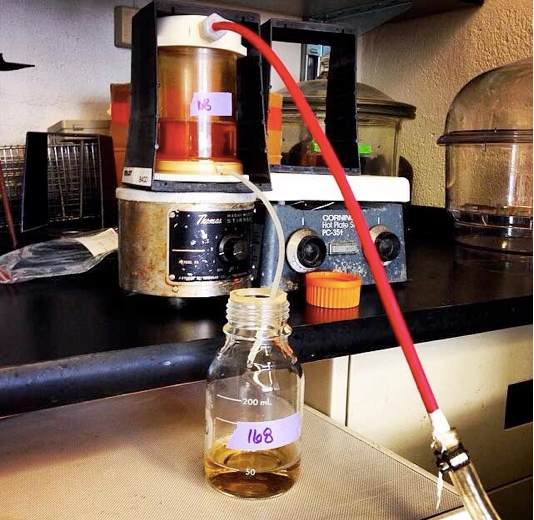

Figure 1. Amicon ultrafiltration setup. The ultrafiltration cells are connected to compressed nitrogen gas. The gas applies pressure, forcing the supernatant through the 100 kDa filter membrane. Vesicles stay in the concentrated supernatant while smaller proteins and liquid supernatant flow into waste container.

Figure 2. Beckman ultracentrifuge tubes. Ultracentrifugation tubes utilized to pellet vesicles.
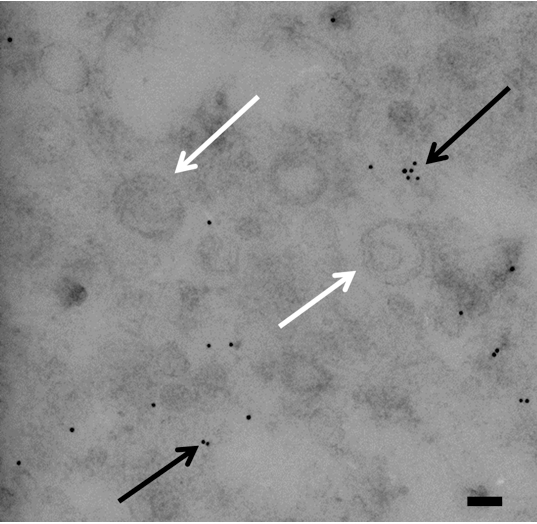
Figure 3. Transmission electron micrograph of immunogold labeling of GXM in vesicles from Bacillus stubtilis. Micrograph shows vesicles and gold particles. Gold particles indicate the presence of GXM. Notice all of the gold is outside of vesicles. White arrows indicate vesicles and black arrows indicate gold particles. Scale bar=100 nm
Vesicles are not incredibly stable so it is suggested to process them immediately after isolation. Only Beckman polyallomer/propylene ultracentrifugation tubes can be used for IEM preparation.Polycarbonate tubes cannot be used for this procedure but can be used when purifying vesicles for other experiments. The sonicated (unnatural) vesicles serve as a positive control. It is expected that vesicles from the sonicated vesicle preparation will contain significantly more gold particles than the naturally produced vesicles. One may also expect that sonicated vesicles have a smaller and less variable diameter than that of natural vesicles.
Notes
B. subtilis strain 168 concentrates very quickly but other strains of bacteria may not. B. subtilis strain 168 produces a large quantity of recoverable vesicles and only requires 100 ml of culture whereas other bacteria may require volumes larger than 1 L. Concentrating to small volumes allows for less ultracentrifugation spins and cuts down on the use of the ultracentrifuge tubes. Ultracentrifuge tubes can be washed and re-used but be aware there may be some contamination.
Recipes
- 1 L 1x phosphate buffered saline (PBS) (pH 7.4)
137 mM NaCl
2.7 mM KCl
10 mM Na2HPO4
1.8 mM KH2PO4
Bring to pH 7.4 with HCl
Dissolve in H2O up to 1 L
Acknowledgments
Funding from NIH Grant Numbers: HL059842, AI033774, AI033142, AI052733 and Center for AIDS Research at Albert Einstein College of Medicine.
Protocol is adapted from Brown et al. (2014) and vesicle purification is adapted from Prados-Rosales et al. (2014).
References
- Brown, L., Kessler, A., Cabezas-Sanchez, P., Luque-Garcia, J. L. and Casadevall, A. (2014). Extracellular vesicles produced by the Gram-positive bacterium Bacillus subtilis are disrupted by the lipopeptide surfactin. Mol Microbiol 93(1): 183-198.
- Prados-Rosales, R., Brown, L., Casadevall, A., Montalvo-Quirós, S. and Luque-Garcia, J. L. (2014). Isolation and identification of membrane vesicle-associated proteins in Gram-positive bacteria and mycobacteria. MethodsX (1): 124-129.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Brown, L., Perumal, G. and Casadevall, A. (2015). Differentiation of Naturally Produced Extracellular Membrane Vesicles from Lipid Aggregation by Glucuronoxylomannan Immunogold Transmission Electron Microscopy in Bacillus subtilis. Bio-protocol 5(5): e1408. DOI: 10.21769/BioProtoc.1408.
Category
Microbiology > Microbial cell biology > Cell imaging
Microbiology > Microbial cell biology > Organelle isolation
Cell Biology > Cell imaging > Electron microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









