- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cyclohexane Diamine Tetraacetic Acid (CDTA) Extraction of Plant Cell Wall Pectin
Published: Vol 4, Iss 24, Dec 20, 2014 DOI: 10.21769/BioProtoc.1357 Views: 12535
Reviewed by: Marisa RosaRenate WeizbauerAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1935 Views
New Approach to Detect and Isolate Rhamnogalacturonan-II in Arabidopsis thaliana Seed Mucilage
Dayan Sanhueza and Susana Saez-Aguayo
Sep 5, 2025 1285 Views
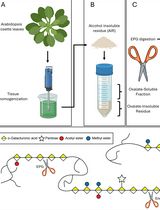
Detailed Method for the Purification of Rhamnogalacturonan-I (RG-I) in Arabidopsis thaliana
Liang Zhang [...] Breeanna R. Urbanowicz
Feb 5, 2026 241 Views
Abstract
The goal of this procedure is to extract pectin from plant cell walls. Pectins are galacturonic acid containing polymeric sugars that are important components of plant cell walls. Various procedures aimed at studying plant cell wall components require the extraction of pectin. Pectin is synthesized in the Golgi apparatus in a highly esterified fashion and is de-esterified in the cell wall (Mohnen, 2008). Pectin is generally water soluble. De-esterified pectin can form so-called “egg-box structures” in the presence of Ca2+ ions (Mohnen, 2008; Harholt et al., 2010). Pectin in these “egg-box structures” is cross-linked and less soluble. Cyclohexane diamine tetraacetic acid (CDTA) chelates Ca2+ ions and hence allows extraction of Ca2+ cross-linked pectin from cell walls.
Materials and Reagents
- Plant material or cell wall preparation of choice (see Note 1)
- Cyclohexane diamine tetraacetic acid (CDTA) (Fluka, catalog number: 34588 )
- Tris-base (Sigma-Aldrich, catalog number: T6066 )
- Sodium hydroxide (Thermo Fisher Scientific, catalog number: S318 )
- Liquid nitrogen
- CDTA extraction buffer (see Recipes)
Equipment
- Liquid nitrogen container
- Paint shaker (e.g. Harbil, model: 5G-HD) and ball bearings (e.g. 3 mm diameter steel beads) (mortar and pestle or bead beater)
- Media bottle
- Single-channel pipettor
- Microfuge tubes appropriate for use at 95 °C and for freezing in liquid N2 (Thermo Fisher Scientific, catalog number: 02-707-355 )
Note: We use 2 ml screw cap tubes.
- Pipette tips
- Water bath or incubator
- Vortex mixer
- Microcentrifuge capable of spinning at 10,000 x g
- Optional: Screw cap tubes or lid locks
Procedure
- Pre-heat water bath or incubator to 95 °C and pre-heat CDTA extraction buffer.
- Collect plant tissue in tubes - we use leaf material from approx. 4 week old Arabidopsis thaliana plants. Three fully expanded Arabidopsis leaves weigh about 250 mg.
- Freeze tubes containing the tissue in liquid nitrogen (tissue can be stored at -80 °C).
- Pulverize frozen tissue using ball bearings and a paint shaker, mortar and pestle or a bead beater. The method used to pulverize tissue is not critical. Keep tissue frozen. If paint shaker is used three cycles of three minutes each are generally sufficient to pulverize tissue.
- Add 1 ml of CDTA extraction buffer to 250 mg of frozen pulverized Arabidopsis tissue.
- Place tubes at 95 °C. Use caution, the liquid will become very hot and lids may open; therefore use screw cap tubes or lid locks to secure lids.
- Incubate for 15 min, vortex every 5 min.
- Centrifuge at 10,000 x g for 10 min at room temperature.
- The supernatant contains the pectin.
- Supernatant can be frozen and freeze dried for downstream applications.
Notes
- We directly use pulverized Arabidopsis leaf material (Bethke et al., 2014). Alternatively crude cell wall preparations like alcohol insoluble residue (Gille et al., 2009) preparations etc. can be used (pre-heat the water bath and extraction buffer and proceed to step 5).
- Different groups have reported different extraction times and temperatures e.g. 15 min at 95 °C (Siedlecka et al., 2008) or 4 h at room temperature (Moller et al., 2008). In our experience about twice the amount of pectin could be extracted when extraction was performed for 15 min at 95 °C as compared to the longer extraction at room temperature.
- Different groups have utilized similar protocols for the extraction of pectin from various monocotyledonous and dicotyledonous plants e.g. cucumber, tomato, celery (Jarvis et al., 1982), poplar (Siedlecka et al., 2008), soybean (Huisman et al., 2001) or wheat (Wiethölter et al., 2003).
- The extraction buffer can be stored at room temperature for several weeks.
- Pectin extracted using this procedure can be used for various downstream application including dot-blot analysis with pectin specific antibodies, sugar analysis, ion-exchange chromatography or determination of degree of esterification of pectin (Bethke et al., 2014; MacDougall et al., 1997).
Recipes
- CDTA extraction buffer
50 mM CDTA
50 mM Tris-base
Adjust pH to 7.2 using sodium hydroxide
Acknowledgments
This work was funded by the Division of Chemical Sciences, Geosciences, and Biosciences, Office of Basic Energy Sciences of the U.S. Department of Energy through Grant DE-FG02-05ER15670 to J.G.
Many labs have used similar protocols in the past. We have adapted this protocol from Siedlecka et al. (2008).
References
- Bethke, G., Grundman, R. E., Sreekanta, S., Truman, W., Katagiri, F. and Glazebrook, J. (2014). Arabidopsis PECTIN METHYLESTERASEs contribute to immunity against Pseudomonas syringae. Plant Physiol 164(2): 1093-1107.
- Gille, S., Hansel, U., Ziemann, M. and Pauly, M. (2009). Identification of plant cell wall mutants by means of a forward chemical genetic approach using hydrolases. Proc Natl Acad Sci U S A 106(34): 14699-14704.
- Harholt, J., Suttangkakul, A. and Vibe Scheller, H. (2010). Biosynthesis of pectin. Plant Physiol 153(2): 384-395.
- Huisman, M. M. H., Fransen, C. T. M., Kamerling, J. P., Vliegenthart, J. F. G., Schols, H. A. and Voragen, A. G. J. (2001). The CDTA-soluble pectic substances from soybean meal are composed of rhamnogalacturonan and xylogalacturonan but not homogalacturonan. Biopolymers 58(3): 279-294.
- Jarvis, M. C. (1982). The proportion of calcium-bound pectin in plant cell walls. Planta 154(4): 344-346.
- MacDougall, A. J., Rigby, N. M., Ring, S. G. (1997). Phase separation of plant cell wall polysaccharides and its implications for cell wall assembly. Plant Physiol 114(1): 353-362.
- Mohnen, D. (2008). Pectin structure and biosynthesis. Curr Opin Plant Biol 11(3): 266-277.
- Moller, I., Marcus, S. E., Haeger, A., Verhertbruggen, Y., Verhoef, R., Schols, H., Ulvskov, P., Mikkelsen, J. D., Knox, J. P. and Willats, W. (2008). High-throughput screening of monoclonal antibodies against plant cell wall glycans by hierarchical clustering of their carbohydrate microarray binding profiles. Glycoconj J 25(1): 37-48.
- Siedlecka, A., Wiklund, S., Peronne, M. A., Micheli, F., Lesniewska, J., Sethson, I., Edlund, U., Richard, L., Sundberg, B. and Mellerowicz, E. J. (2008). Pectin methyl esterase inhibits intrusive and symplastic cell growth in developing wood cells of Populus. Plant Physiol 146(2): 554-565.
- Wiethölter, N., Graeβner, B., Mierau, M., Willats, W. G. T., Knox, J. P., Moerschbacher, B. M. (2003). Isolation and characterisation of the homogalacturonan from type II cell walls of the commelinoid monocot wheat using HF-solvolysis. Carbohydrate Res 338(5):423-431.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Bethke, G. and Glazebrook, J. (2014). Cyclohexane Diamine Tetraacetic Acid (CDTA) Extraction of Plant Cell Wall Pectin. Bio-protocol 4(24): e1357. DOI: 10.21769/BioProtoc.1357.
- Bethke, G., Grundman, R. E., Sreekanta, S., Truman, W., Katagiri, F. and Glazebrook, J. (2014). Arabidopsis PECTIN METHYLESTERASEs contribute to immunity against Pseudomonas syringae. Plant Physiol 164(2): 1093-1107.
Category
Plant Science > Plant biochemistry > Carbohydrate
Biochemistry > Carbohydrate > Polysaccharide
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










