- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of the Electrogenicity of Bile Salt/H+ Antiport in Escherichia coli
Published: Vol 4, Iss 21, Nov 5, 2014 DOI: 10.21769/BioProtoc.1279 Views: 8256
Reviewed by: Aksiniya AsenovaKanika GeraAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

β-lactamase (Bla) Reporter-based System to Study Flagellar Type 3 Secretion in Salmonella
Fabienne F. V. Chevance and Kelly T. Hughes
Jun 20, 2023 1802 Views
Determination of Poly(3-hydroxybutyrate) Content in Cyanobacterium Synechocystis sp. PCC 6803 Using Acid Hydrolysis Followed by High-performance Liquid Chromatography
Janine Kaewbai-ngam [...] Tanakarn Monshupanee
Aug 20, 2023 1866 Views
An HPLC-based Assay to Study the Activity of Cyclic Diadenosine Monophosphate (C-di-AMP) Synthase DisA from Mycobacterium smegmatis
Avisek Mahapa [...] Dipankar Chatterji
Dec 20, 2024 1816 Views
Abstract
The transmembrane proton gradient (ΔpH) is the primary source of energy exploited by secondary active substrate/H+ antiporters to drive the electroneutral transport of substrates across the Escherichia coli (E. coli) inner membrane. Such electroneutral transport results in no net movement of charges across the membrane. The charge on the transported substrate and the stoichiometry of the exchange reaction, however, can result in an electrogenic reaction which is driven by both the ΔpH and the electrical (∆Ψ) components of the proton electrochemical gradient, resulting in a net movement of electrical charges across the membrane. We have shown that the major facilitator superfamily transporter MdtM - a multidrug efflux protein from E. coli that functions physiologically in protection of bacterial cells against bile salts - imparts bile salt resistance to the bacterial cell by coupling the exchange of external protons (H+) to the efflux of bile salts from the cell interior via an electrogenic antiport reaction (Paul et al., 2014). This protocol describes, using fluorometry, how to detect electrogenic antiport activity of MdtM in inverted membrane vesicles of an antiporter-deficient strain of E. coli TO114 cells by measuring transmembrane ∆Ψ. The method exploits changes that occur in the intensity of the fluorescence signal (quenching and dequenching) of the probe Oxonol V in response to changes in membrane potential due to the MdtM-catalysed sodium cholate/H+ exchange reaction. The protocol can be adapted to detect activity of any secondary active antiporter that couples the electrogenic translocation of H+ across a biological membrane to that of its counter-substrate, and may be used to unmask otherwise camouflaged transport activities and physiological roles.
Keywords: Membrane transportMaterials and Reagents
- E. coli TO114 (gift of Prof. Hiroshi Kobayashi, Chiba University, Japan)
- E. coli BW25113 (National BioResource Project, Japan)
- pBAD/Myc-His A expression vector (Life Technologies, catalog number: V440-01 )
- L-(+)-arabinose (Sigma-Aldrich, catalog number: A3256 )
- Carbenicillin (Carbenicillin Direct)
- Agar (Sigma-Aldrich, catalog number: A1296 )
- Tryptone (Fluka, catalog number: T7293 )
- Yeast extract (Fluka, catalog number: 92114)
- Potassium chloride (Thermo Fisher Scientific, catalog number: BP366 )
- Bis-(3-phenyl-5-oxoisoxazol-4-yl)pentamethine oxonol (Oxonol V) (Life Technologies, Molecular Probes®, catalog number: O-266 )
Note: In our original work we used Oxonol V supplied by Cambridge Bioscience Ltd. However, this product no longer appears in their catalogue. - Nigericin (Sigma-Aldrich, catalog number: N7143 )
- Valinomycin (Sigma-Aldrich, catalog number: 94675 )
- BisTris propane (BTP) (Sigma-Aldrich, catalog number: B6755 )
- Phenylmethanesulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: P7626 )
- Deoxyribonuclease I (DNase) from bovine pancreas (Sigma-Aldrich, catalog number: DN25 )
- Carbonyl cyanide 3-chlorophenylhydrazone (CCCP) (Sigma-Aldrich, catalog number: C2759 )
- Sodium DL-lactate solution 50% aqueous (VWR International, catalogue number: 27927.298 )
- Magnesium sulphate heptahydrate (MgSO4.7H2O) (Thermo Fisher Scientific, catalogue number: M/1000/60 )
- Sodium cholate hydrate (Sigma-Aldrich, catalog number: C1254 )
- Sodium D-gluconate (Sigma-Aldrich, catalog number: S2054 )
- Potassium D-gluconate (Sigma-Aldrich, catalog number: G4500 )
- Absolute ethanol (Thermo Fisher Scientific, catalogue number: E/0650DF/17 )
- High purity (18 MΩ) Millipore or AnalR water
- D-sorbitol (Sigma-Aldrich, catalog number: S1876 )
- Sucrose (Sigma-Aldrich, catalog number: 84097 )
- DL-dithiothreitol(Sigma-Aldrich, catalog number: 43815 )
- TRIZMA base (Sigma-Aldrich, catalog number: T1503 )
- Sulphuric acid (Sigma-Aldrich, catalog number: 320501 )
- LBK agar (see Recipes)
- LBK liquid medium (see Recipes)
- Tris/sorbitol/dithiothreitol/sucrose (TSDS) buffer (see Recipes)
- Transport assay buffer (see Recipes)
- 2 M sodium D-gluconate stock solution (see Recipes)
- 2 M potassium D-gluconate stock solution (see Recipes)
Equipment
- Temperature-controlled shaking incubator for bacterial growth
- Large ice bucket
- Petri dishes for bacterial colony growth
- 100 ml conical flasks
- 250 ml conical flasks (x2)
- 5,000 ml conical flask
- Refrigerated, large capacity centrifuge and rotor for harvesting bacterial cells
- 1,000 ml or 500 ml centrifuge pots and lids
- Benchtop vortexer
- 100 ml beaker and stir bar to fit
- Magnetic stirrer
- 25 ml disposable plastic pipettes
- Selection of single channel pipettes (1,000 µl, 100 µl, 20 µl, 10 µl)
- Pipette tips for above
- Refrigerated centrifuge and rotor capable of spinning ~50 ml tubes at 18k x g.
- Refrigerated ultracentrifuge, rotor and polycarbonate ultracentrifuge tubes capable of handling ~30 - 50 ml volumes
- French Press (Thermo Electron Corp, catalogue number: FA-078 )
- Standard pressure cell (40 kpsi; 35 ml capacity) (Thermo Electron Corp, catalogue number: FA-031 )
- 50 ml syringes (for filtering solutions)
- 0.22 µm sterile filters to fit 50 ml syringe
- 1 L and 500 ml Duran bottles with lids
- 1.5 ml Eppendorf tubes
- Medical wipes (Kimwipes®)
- Parafilm
- UV/vis spectrophotometer and 10 mm pathlength quartz cuvette
- 10 x 4 mm, 1,400 µl volume quartz cuvette for fluorescence spectroscopy (Hellma, catalog number: 104F-QG )
- Small magnetic stir bar to fit inside quartz cuvette
- Fluorometer e.g. Fluoromax-4 (Horiba) capable of performing time-based acquisition measurements and fitted with a temperature controlled cuvette holder and stirrer
- Source of compressed air (either aerosol can or fixed supply) for drying cuvette
Procedure
- Growth and harvesting of bacterial cultures and preparation of inverted membrane vesicles
- Due to typically low levels of endogenous transporter expression, this method necessitates that the transporter of interest be overexpressed from a multicopy plasmid. We therefore begin the protocol with a description of the conditions used specifically for overexpression of MdtM from the pBAD/Myc-His A expression vector; these conditions (for example, the vector type, inducer concentration, and growth temperature) will vary depending upon the transporter that requires to be overexpressed. Furthermore, although the protocol described here pertains to measurement of electrogenic bile salt (sodium cholate) transport by MdtM specifically, it can be readily adapted for measurement of the electrogenicity of transport by other secondary active antiporters. Typically, a 1 L culture of bacterial cells will provide sufficient material for these experiments.
- Perform a fresh transformation with the pBAD/Myc-His A plasmid DNA encoding wild type or mutant transporter (or your vector of choice encoding your transporter of interest) into chemically competent E. coli TO114 cells. Plate the cells onto Luria Bertani Potassium (LBK) agar containing 100 μg/ml carbenicillin for selection. LBK agar must be used instead of regular Luria Bertani (LB) agar because the latter contains NaCl and the TO114 strain is sensitive to this salt due to deletion of the chromosomally encoded Na+/H+ antiporter NhaA. Incubate the plate overnight (~15 to ~18 h) at 37 °C.
- Pick a few single colonies from the agar plate and use to inoculate 2 x 250 ml conical flasks, each containing 100 ml of LBK liquid medium supplemented with 100 μg/ml carbenicillin. Grow the cultures overnight (~15 h) at 30 °C with 250 rpm shaking in a temperature-controlled shaking incubator. Measure the optical density at 600 nm (OD600) of the culture using a spectrophotometer; it should be between 3.0-3.3. If the OD600 is less than 3.0, then incubate for a further 0.5-1.0 h and re-measure. If the OD600 is greater than 3.3, the cultures have overgrown and fresh cultures will need to be prepared.
- Inoculate a 5 L conical flask containing 1,000 ml of LBK liquid medium supplemented with 100 μg/ml carbenicillin with 15 ml of overnight culture. Incubate at 32 °C with 270 rpm shaking for ~2.5 hrs in a temperature-controlled shaking incubator. The OD600 should be ~0.6. Drop the temperature to 25 °C and grow with 270 rpm shaking until the OD600 is 1.0. This usually takes between 0.5 to 1.0 h.
- Induce overexpression of MdtM by addition of 0.1% w/v L-(+)-arabinose [5 ml of 20% w/v L-(+)-arabinose] to the culture. After addition of arabinose, grow the cells for a further 1.5 h at 25 °C with 270 rpm shaking prior to harvesting.
- Transfer the E. coli TO114 cells that contain overexpressed transporter into 500 ml or 1,000 ml capacity centrifuge pots that have been pre-chilled on ice for at least 15 min. Pre-cool the centrifuge to 4 °C and harvest cells by centrifugation at 5,000 x g for 20 min.
- Decant the supernatant and wash the pelleted cells by resuspending (either by gentle vortexing using a benchtop vortexer or by gentle aspiration using a 25 ml sterile plastic pipette) in chilled (4 °C) TSDS buffer. Use 30 ml of TSDS buffer for each one litre of cell culture that was pelleted. Maintain the cells on ice during this procedure. Harvest the washed cells by centrifugation as described in step A1 (above) and repeat the washing procedure. Resuspend the resultant cell pellet in 30 ml of chilled TSDS buffer containing 2 mM PMSF (Which should be made up as a 100 mM stock in ethanol and stored at -20 °C; ensure the solution is thawed thoroughly and vortexed vigorously before use.) and 5 µM DNase and maintain the mixture on ice.
- Decant the resuspended cells into a 100 ml beaker containing a stir bar, place on a magnetic stirrer and stir in a cold room at 4 °C for 20 min. Alternatively, place the beaker on ice in an ice bucket, place on the magnetic stirrer and stir for 20 min.
- The method for production of inverted vesicles relies on a combination of the fluid shear forces and decompression created as the cell mixture passes through the needle valve of a French pressure cell. Generate inverted membrane vesicles by a single passage of the resuspended cell mixture through a French pressure cell at a minimum of 4,000 psi. If the pressure is too low, inverted vesicles will not be formed. The pressure cell should be chilled on ice for ~30 min prior to use. The resulting inverted vesicle mixture should be collected in a 100 ml conical flask kept on ice.
- Decant the mixture into a pre-chilled ~50 ml centrifuge tube and remove any unbroken cells and cell debris by centrifugation at 18,000 x g for 10 min at 4 °C. Carefully decant the supernatant containing the cell membrane vesicles (do not disturb the pellet that contains unbroken cells and cell debris) into a pre-chilled 30-50 ml volume polycarbonate ultracentrifuge tube on ice.
- Harvest the inverted vesicles by ultracentrifugation at 100,000 x g for 1 h at 4 °C. Carefully decant the supernatant and retain the pellet. Place the ultracentrifuge tube containing the pelleted vesicles on ice.
- Thoroughly resuspend the inverted vesicle pellet in 1 ml of ice-cold TSDS buffer by gentle aspiration using a 1,000 µl pipette. Transfer the resuspended vesicles to a pre-chilled 1.5 ml Eppendorf tube on ice for use in the transport assay. In our experience, vesicles stored on ice are stable for several hours.
- Quantify the total membrane protein content of the inverted vesicles by UV absorbance spectroscopy at 280 nm. Blank the spectrophotometer using a 10 mm pathlength quartz cuvette containing 1,000 µl of TSDS buffer. The buffer should be at room temperature to prevent frosting of the cuvette faces. Clean the faces of the cuvette using a fresh paper wipe prior to measurement. Once the spectrophotometer is blanked, remove 5 µl of buffer from the cuvette using a 10 µl pipette and replace with 5 µl of vesicles. Cover the opening of the cuvette with a square of Parafilm and invert the cuvette a few times to ensure a homogeneous distribution of vesicles. Record the absorbance of the vesicle mixture at 280 nm and calculate the total membrane protein concentration assuming that an A280 of 1.0 is equivalent to a protein concentration of 1.0 mg/ml. Remember to multiply the 280 nm absorbance value you obtain by a factor of 200 to calculate the concentration of the undiluted vesicle mixture in mg/ml.
- Due to typically low levels of endogenous transporter expression, this method necessitates that the transporter of interest be overexpressed from a multicopy plasmid. We therefore begin the protocol with a description of the conditions used specifically for overexpression of MdtM from the pBAD/Myc-His A expression vector; these conditions (for example, the vector type, inducer concentration, and growth temperature) will vary depending upon the transporter that requires to be overexpressed. Furthermore, although the protocol described here pertains to measurement of electrogenic bile salt (sodium cholate) transport by MdtM specifically, it can be readily adapted for measurement of the electrogenicity of transport by other secondary active antiporters. Typically, a 1 L culture of bacterial cells will provide sufficient material for these experiments.
- Measurement of electrogenic transport
- In this section we describe the set-up parameters for a Fluoromax-4 fluorometer. These parameters, however, can form the basis for the set-up of other fluorometers. Once the instrument is switched on and the software booted up, set the temperature of the cuvette holder to 25 °C. Open the instrument software and select for time-based data acquisition with excitation and emission wavelengths of 599 nm and 634 nm, respectively. Set the excitation and emission slit widths to 10 nm and 20 nm, respectively.
- Add an aliquot of inverted vesicles (which should be maintained on ice in TSDS buffer) to room temperature transport assay buffer containing the Oxonol V probe in a 10 mm x 4 mm quartz cuvette to a final concentration of 0.5 mg/ml membrane protein in a total volume of 1,500 µl. The longest pathlength of the cuvette should face the excitation light source. Place a small magnetic flea into the cuvette and stir the contents gently. Allow the vesicles and assay buffer to equilibrate for ~200 sec.
- Start recording the fluorescence emission.
- After approximately 50 sec, add 15 µl of 200 mM stock solution of sodium DL-lactate to the cuvette contents to give a final sodium DL-lactate concentration of ~2.0 mM. Addition of lactate initiates respiration-dependent generation of ∆Ψ that causes a dequench of the Oxonol V fluorescence signal (see Figure 1).
- If the fluorescence signal does not quench, or enhances, the vesicles have either not maintained integrity or are not inverted and their preparation needs to be repeated.
- Following the establishment of ΔΨ, monitor the Oxonol V fluorescence dequench for a further ~150 sec until it stabilises. Initiate MdtM-mediated antiport by adding substrate (in this case the bile salt sodium cholate) to the inverted vesicle mixture. We added 12.5 µl of 250 mM stock solution made up in high purity water to give a final concentration of sodium cholate in the cuvette of ~2.0 mM. For other substrates we suggest testing a range varying from 1 mM to 100 mM to establish the concentration that gives the best dequench signal. If the transport reaction is electrogenic there should be an immediate dequench (enhancement) of the Oxonol V fluorescence emission signal as the established membrane potential, ΔΨ, is consumed by the transport reaction (see Figure 1a).
- Record the fluorescence dequench signal for ~60 s to allow the antiport reaction to achieve a steady state (as observed by a plateauing of the fluorescence signal). Addition of the protonophore CCCP to a final concentration of ~100 µM (1.6 µl of a 100 mM stock made up in ethanol) in the assay mixture should be performed to abolish transport by collapsing both the membrane potential and ΔpH. Addition of CCCP should result in a further dequench of the fluorescence signal (see Figure 1a). Record the fluorescence signal for a further ~40 sec then terminate the acquisition and save the electronic data.
- Decant the cuvette contents into a suitable waste container and wash the cuvette thoroughly with ethanol then high-purity water. Dry the cuvette carefully with compressed air.
- As a further control, and to provide evidence that the inverted vesicles retain integrity and are therefore able to maintain an electrochemical potential across the membrane during the assay lifetime, the fluorescence response of Oxonol V upon addition of the ionophore nigericin (which at low concentrations selectively consumes ΔpH in the presence of potassium ions via electroneutral K+/H+ exchange) in place of substrate should be measured. For this experiment, inverted vesicles of TO114 cells containing the transporter of interest are incubated in assay buffer that contains 50 mM potassium D-gluconate. After addition of sodium DL-lactate to initiate respiration as described in step B3 above, 1.5 µl of 1 mM nigericin stock solution made up in ethanol (to give a final concentration of ~1 µM) is added to the cuvette. Concentrations of nigericin > 1 µM must be avoided to prevent electrogenic exchange that can occur under conditions of high nigericin concentrations and abolish both components of the electrochemical gradient. Record the fluorescence dequench signal for a further ~60 s then add the ionophore valinomycin (to selectively abolish ΔΨ) to a final concentration of ~5 μM by adding ~1.6 μl of 5 mM stock made up in ethanol (see Figure 1d). Record data for a further 40 sec prior to termination of the acquisition.
- In this section we describe the set-up parameters for a Fluoromax-4 fluorometer. These parameters, however, can form the basis for the set-up of other fluorometers. Once the instrument is switched on and the software booted up, set the temperature of the cuvette holder to 25 °C. Open the instrument software and select for time-based data acquisition with excitation and emission wavelengths of 599 nm and 634 nm, respectively. Set the excitation and emission slit widths to 10 nm and 20 nm, respectively.
Representative data
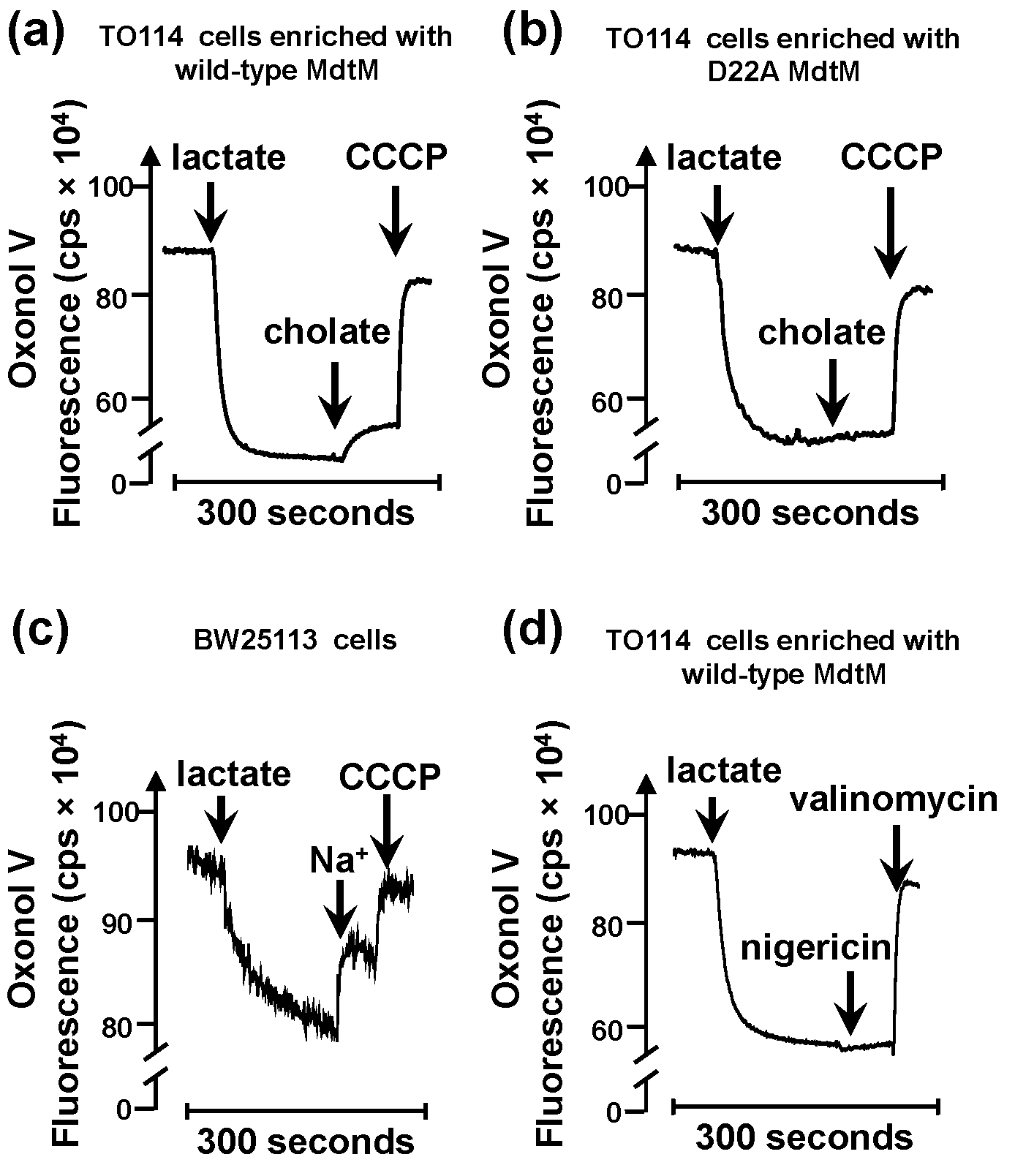
Figure 1. Representative data illustrating typical changes in the fluorescence signal of the ΔΨ-sensitive fluorophore Oxonol V in response to MdtM-catalysed electrogenic transport in inverted membrane vesicles. Addition of lactate to energise vesicles results in generation of a respiratory ΔΨ, as evidenced by a rapid quench of the Oxonol V fluorescence signal. (a) Addition of 2.5 mM cholate to inverted vesicles generated from TO114 cells enriched with wild-type MdtM results in a partial depolarization of ΔΨ, represented as a dequenching of the Oxonol V fluorescence, as the ΔΨ was consumed by the MdtM-mediated bile salt/H+ transport reaction. (b) Addition of cholate to negative control vesicles generated from TO114 cells enriched with dysfunctional MdtM D22A results in a small but perceptible dequench arising from residual electrogenic Na+/H+ antiport activity of the mutant. (c) Positive control assay in which Na+ ions are added to inverted vesicles that contain a full complement of electrogenic antiporters in order to measure electrogenic Na+/H+ activity of NhaA transporter. (d) Response of Oxonol V fluorescence to addition of the nigericin. In the presence of K+ ions, this ionophore selectively dissipates ΔpH and converts it into ΔΨ, resulting in a further quench of the fluorescence signal. Valinomycin collapses the ΔΨ. In assays (a, b and c), addition of the protonophore CCCP at the time indicated resulted in almost complete dissipation of ΔΨ. The fluorescence intensity is measured in counts per second (cps). The fluorescence intensity you measure may differ depending upon how your instrument is set up.
Notes
- Resuspended inverted vesicles from step A8 can be transferred to tubes in aliquots of 25-100 µl, snap-frozen in liquid nitrogen and stored at -80 °C for subsequent use. Vesicles frozen in this way will retain their integrity for several months. However, if frozen vesicle stocks are used for the subsequent transport measurements, the vesicles must be thawed very slowly on ice prior to use to prevent their fracture.
- To ensure the suitability of the experimental conditions for detection of electrogenic antiport, a positive control should first be performed using inverted vesicles produced from E. coli cells e.g. strain BW25113 that contain a full complement of electrogenic antiporters. This control experiment should be performed by following the protocol below except that the transport assay buffer pH should be adjusted to pH 8.5 with H2SO4, and sodium gluconate (to a final concentration in the cuvette of ~100 mM by adding 80 µl of a 2 M stock of sodium gluconate) should be added in place of sodium cholate to specifically enable detection of electrogenic Na+/H+ exchange catalysed by the NhaA transporter (Figure 1c).
- To ensure reproducibility, the assays should be performed in triplicate on at least two separate preparations of inverted vesicles.
- As with all assays that rely on detection of fluorescence, robust controls must be in place to ensure that any detected transport activity can be attributed unambiguously to the protein of interest. In our experiments, we used inverted vesicles that overexpressed MdtM D22A, a dysfunctional point mutant of MdtM, as a negative control (see Figure 1b).
- If the method is to be used for detection of metal ion/H+ antiport activity, the use of inverted vesicles generated from the antiporter-deficient TO114 strain of E. coli is important because at least four other transporters (NhaA, NhaB, ChaA and MdfA) present in the bacterium can catalyse a monovalent metal cation/H+ exchange.
- Because chloride ions can depolarise the membrane potential, all buffer systems must be maintained as chloride-free. Therefore, pH of buffers should be adjusted using H2SO4 instead of HCl.
- Finally, if this protocol is used for comparison of antiport activities of wild type and mutant transporters, the amount of target protein present in the inverted vesicle membranes must be quantified (usually by immunodetection methods) to ensure that any measured differences in H+ uptake are due solely to differences in the activity of the transporters and not to differences in expression levels.
Recipes
- LBK agar (100 ml)
1.0 g tryptone
0.5 g yeast extract
0.745 g KCl
1.5 g agar
Make up to 100 ml with high purity water then autoclave
Add 100 μg/ml carbenicillin for selection when the solution is still liquid and warm to the touch. Add to Petri dishes under sterile conditions - LBK liquid medium (1 L)
10 g tryptone
5 g yeast extract
7.45 g KCl
Make up to 1,000 ml with high purity water then autoclave - Tris/sorbitol/dithiothreitol/sucrose (TSDS) buffer (1 L)
Consisting of 10 mM Tris (pH 7.5), 280 mM sorbitol, 0.5 mM dithiothreitol and 250 mM sucrose
1 M Tris (pH 7.5) 10 ml
1 M sorbitol 280 ml
1 M sucrose 250 ml
Make up to 999.5 ml with high purity water then check pH and adjust if necessary with H2SO4 Sterile filter and store at 4 °C
Add 0.5 ml dithiothreitol from frozen 1 M stocks immediately before use - Transport assay buffer (100 ml)
Consisting of 10 mM BisTris propane (pH 7.2) (or pH 8.5 for the initial control experiment), 5 mM MgSO4, 5 μM Oxonol V
100 mM BisTris propane (pH 7.2 or pH 8.5) 10 ml
2 M MgSO4 0.25 ml
1 mM Oxonol V (made up in ethanol) 0.5 ml
Make up to 100 ml with high purity water then sterile filter.
For control experiments that use nigericin, potassium D-gluconate (2.5 ml of 2 M stock) to a final concentration of 50 mM should be added to the assay buffer
Oxonol V is light sensitive so both the transport assay buffer and Oxonol V stocks should be stored either in amber bottles or in a container protected from light - 2 M sodium D-gluconate stock solution (10 ml)
4.36 g made up to 10 ml with high purity water
Sterile filter and store at 4 °C - 2 M potassium D-gluconate stock solution (10 ml)
6.68g made up to 10 ml with high purity water
Sterile filter and store at 4 °C
Acknowledgments
This work was supported by BBSRC grant BB/K014226/1 (to CJL). The protocol described above is adapted from one reported previously (Resch et al., 2010).
References
- Paul, S., Alegre, K. O., Holdsworth, S. R., Rice, M., Brown, J. A., McVeigh, P., Kelly, S. M. and Law, C. J. (2014). A single-component multidrug transporter of the major facilitator superfamily is part of a network that protects Escherichia coli from bile salt stress. Mol Microbiol 92(4): 872-884.
- Resch, C. T., Winogrodzki, J. L., Patterson, C. T., Lind, E. J., Quinn, M. J., Dibrov, P. and Hase, C. C. (2010). The putative Na+/H+ antiporter of Vibrio cholerae, Vc-NhaP2, mediates the specific K+/H+ exchange in vivo. Biochemistry 49(11): 2520-2528.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Holdsworth, S. R. and Law, C. J. (2014). Measurement of the Electrogenicity of Bile Salt/H+ Antiport in Escherichia coli. Bio-protocol 4(21): e1279. DOI: 10.21769/BioProtoc.1279.
Category
Microbiology > Microbial biochemistry > Other compound
Microbiology > Microbial metabolism > Nutrient transport
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









