- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Plastic Embedding of Arabidopsis Stem Sections
(*contributed equally to this work) Published: Vol 4, Iss 20, Oct 20, 2014 DOI: 10.21769/BioProtoc.1261 Views: 14601
Reviewed by: Elias BassilRu ZhangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Direct Plant Regeneration From Immature Male Inflorescence of Banana (Musa spp.)
Pradeep Chand Deo
Oct 20, 2025 1461 Views

Live-Cell Monitoring of Piecemeal Chloroplast Autophagy
Masanori Izumi [...] Shinya Hagihara
Nov 5, 2025 1716 Views
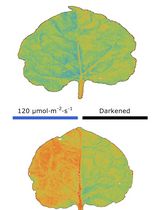
Chloroplast Movement Imaging Under Different Light Regimes With a Hyperspectral Camera
Paweł Hermanowicz [...] Justyna Łabuz
Dec 20, 2025 789 Views
Abstract
The inflorescence stem of the flowering plant Arabidopsis thaliana (thale cress) is an excellent model system to investigate plant vascular tissue patterning and development. Plant vasculature is a complex conducting tissue arranged in strands called vascular bundles, formed by xylem (tissue that carries water) and phloem (tissue that carries photosynthates and signaling molecules). Xylem and phloem are originated from cell division of the meristematic cells of the vascular cambium. In Arabidopsis the flowering stem elongates about three weeks after germination. At this stage it is possible to visualize defects in its development and morphology. Here we describe a protocol to embed in plastic (resin) stem segments either freshly dissected from living plants or previously assayed for β-glucuronidase. This protocol provides an excellent cellular morphology ideal to visualize stem cell types including those of vascular bundles using high-resolution light microscopy.
Keywords: ArabidopsisMaterials and Reagents
- 30 day-old Arabidopsis stem fragments
- 90% acetone
- 4% glutaraldehyde
- 0.1% Tween 20
- Absolute ethanol
- Historesin standard kit (Reichert-Jung, catalog number: 7O-2218-500)
Components:- Basic resin liquid (Figure 2A)
- Activator powder
- Hardener
- Basic resin liquid (Figure 2A)
- Cresyl violet acetate (Sigma-Aldrich, catalog number: C-5042)
- Preinfiltration solution (see Recipes)
- Infiltration solution (see Recipes)
- Embedding medium (see Recipes)
- Cresyl violet staining medium (see Recipes)
Equipment
- Vacuum pump (Figure 1)
- Vacuum jar (Figure 1)
- Histomold trays (6 x 8 mm) (Leica Microsystems, catalog number: 14702218311)
- Histomold trays (13 x 19 mm) (Leica Microsystems, catalog number: 14702218313)
- Specimen Holder adapter (Leica Microsystems, catalog number: 14702218310)
- Drying oven DO-90 (Raypa)
- Motorized microtome Leica RM2155 (Leica Microsystems)
- Water/ethanol bath HI 1210 (Leica Microsystems)
- Tungsten carbide disposable blades TC-65 (Leica Microsystems, catalog number: 140216266379)
- Glass slides Superfrost ultra plus (Menzel-Glasër, catalog number: J3800AMNZ)
- Surgipath micromount (Leica Microsystems, catalog number: 3801731)
- Coverslip (24 x 60) (Menzel-Glasër, catalog number: BB024060A1)
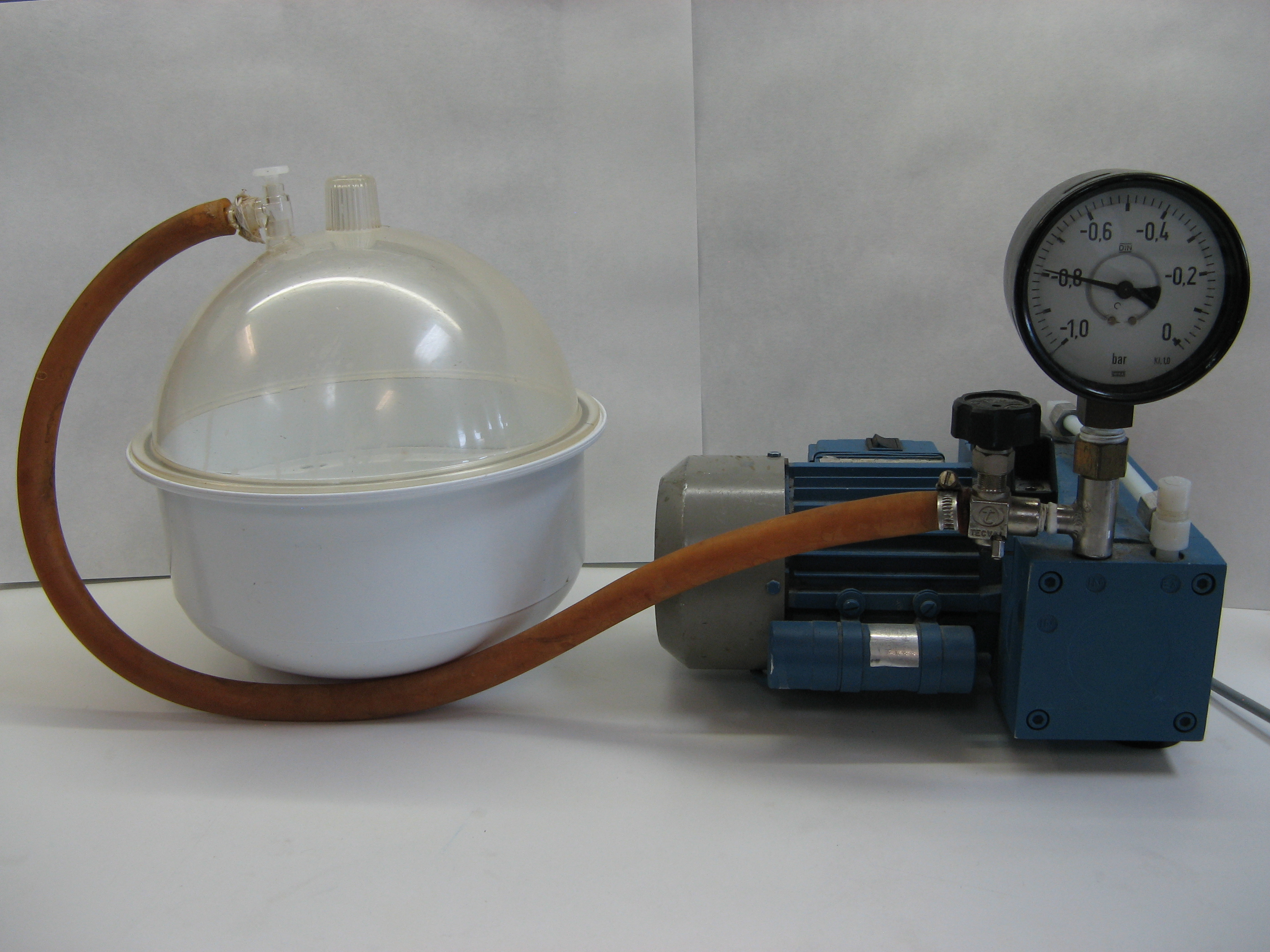

Figure 1. Vacuum jar and vacuum pump used for tissue infiltration
Procedure
- Collect stem fragments (5 to 10 mm) of 30 day-old Arabidopsis plants of the main inflorescence from immediately above the uppermost rosette leaf in a 15 ml falcon (Figure 2A).
- Incubate for 30 min in 90% acetone.
- Wash twice with water by emptying the recipient and filling it with fresh water.
- Cover with 4% glutaraldehyde, 0.1% Tween 20 and vacuum infiltrate for 15 min.
- Leave at 4 °C overnight or longer. Stems can be stored in this fixative.
- Wash twice with water.
- Dehydrate in ethanol series up to 100% ethanol:
30% ethanol (3 h)
50% ethanol (3 h)
75% ethanol (3 h)
95% ethanol (3 h)
100% ethanol (overnight) - Incubate the stems in Preinfiltration solution overnight (minimum) at 4 °C.
- Incubate the stems in Infiltration solution at 4 °C (overnight minimum, up to two weeks).
- Fill the bottom of HISTORESIN MOLD TRAYS with embedding medium (Figures 2B, 2C).
- Immerse and orient the specimens in the mold trays (Figures 2D, 2E).
Put the block holder adapter on the mold before polymerization so that it forms a single piece together with the block (Figures 2F, 2G). - Allow to polymerize overnight at room temperature.
- Take out the block plus adaptor from the mold (Figures 2H-J).
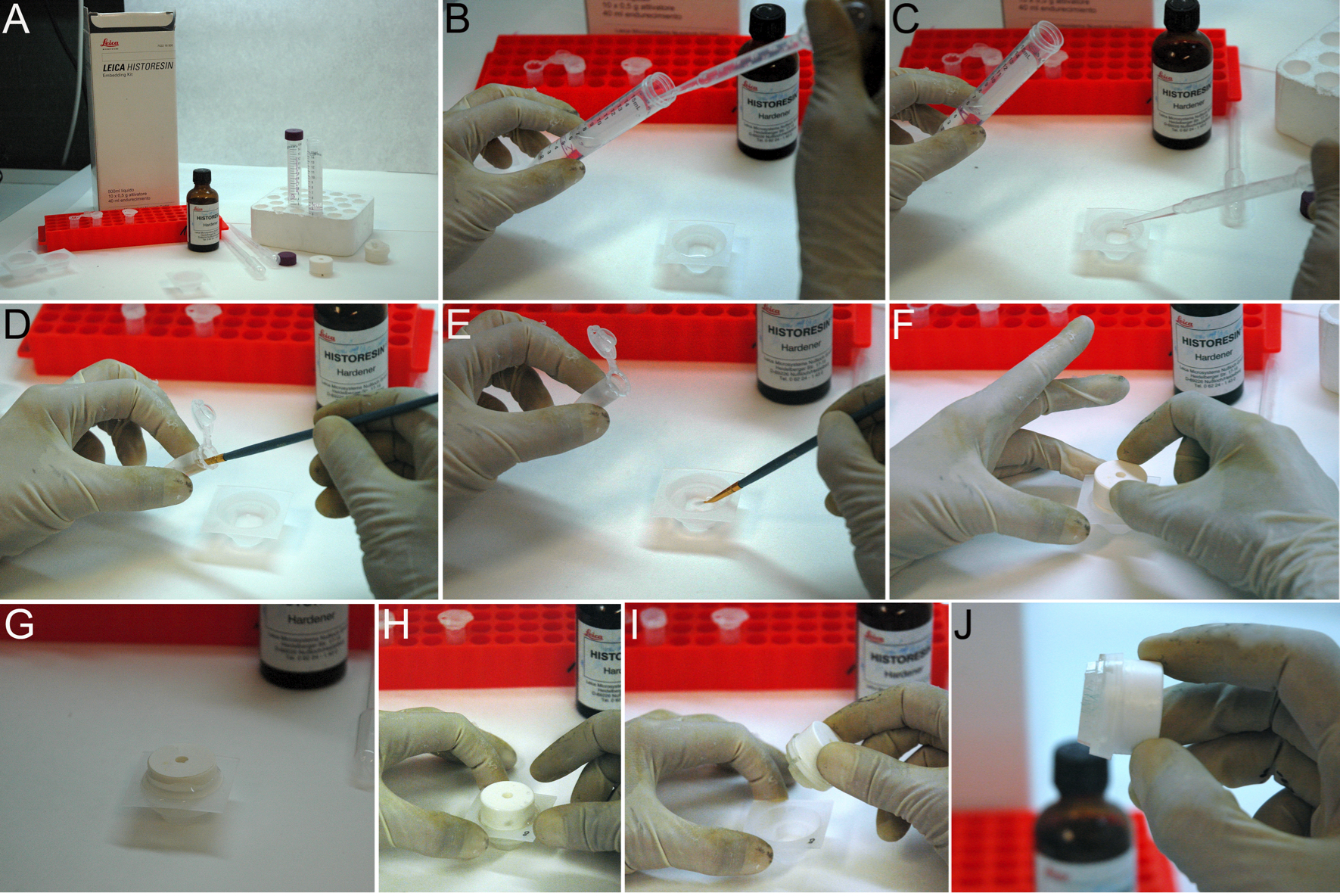

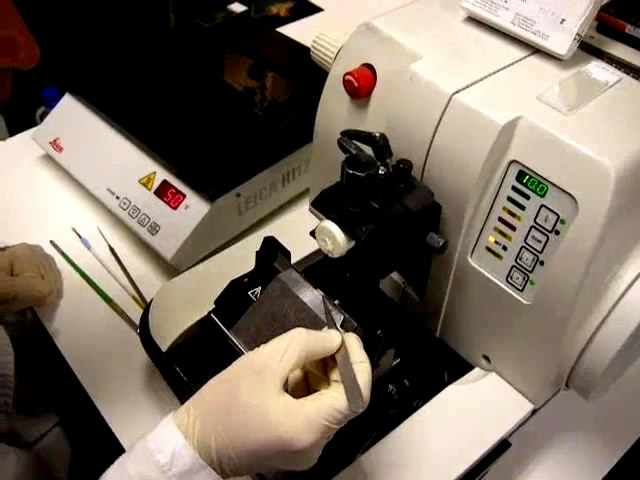
Figure 2. Making a resin block. A. Reactives needed for block making. B, C. The mold trays are filled with embedding medium. D, E. The specimens are immersed and orient in the mold trays. F, G. The block holder adapter is set on the mold before polymerization. H-J. The block plus adaptor is taken out of the mold. - Fix it to the mobile head of the microtome (Figure 3A-3C).
- Cut 3-8 µm sections of resin-embedded specimens using a motorized microtome Leica RM2155 with tungsten carbide blades TC-65 (Figures 3C, 3D, Video 1).
Video 1. Cutting and slides - Float sections in a 50 °C water/ethanol bath for a few seconds until the section flattens (Figures 3E, 3F, Video 1). About 40 ml of 100% ethanol per liter of water are added to restrict the rapid movements of the historesin sections occurring when floating in water.
- Collect sections on a glass slide (Figure 2G, Video 1) and allow to dry in an oven overnight at 50 °C.
- Optional staining step (only when a contrast staining solution is needed).
- Immerse the slides in cresyl violet staining medium 2-5 min.
- Wash with distilled water 2 min.
- Dry the excess of moisture from the slides using a kimwipe.
- Dry in oven for 15 min at 50 °C.
- Mount sections in Surgipath micromount.
- Cover with a coverslip.

Figure 3. Sectioning a resin block. A-C. The block is fixed to the mobile head of the microtome. D. 3-8 µm sections of resin-embedded specimens are cut using a motorized microtome. E. Sections are collected with tweezers. F. Sections are put to float in a 50 °C water bath for a few seconds until the section flattens. G. Sections are collected on a glass slide.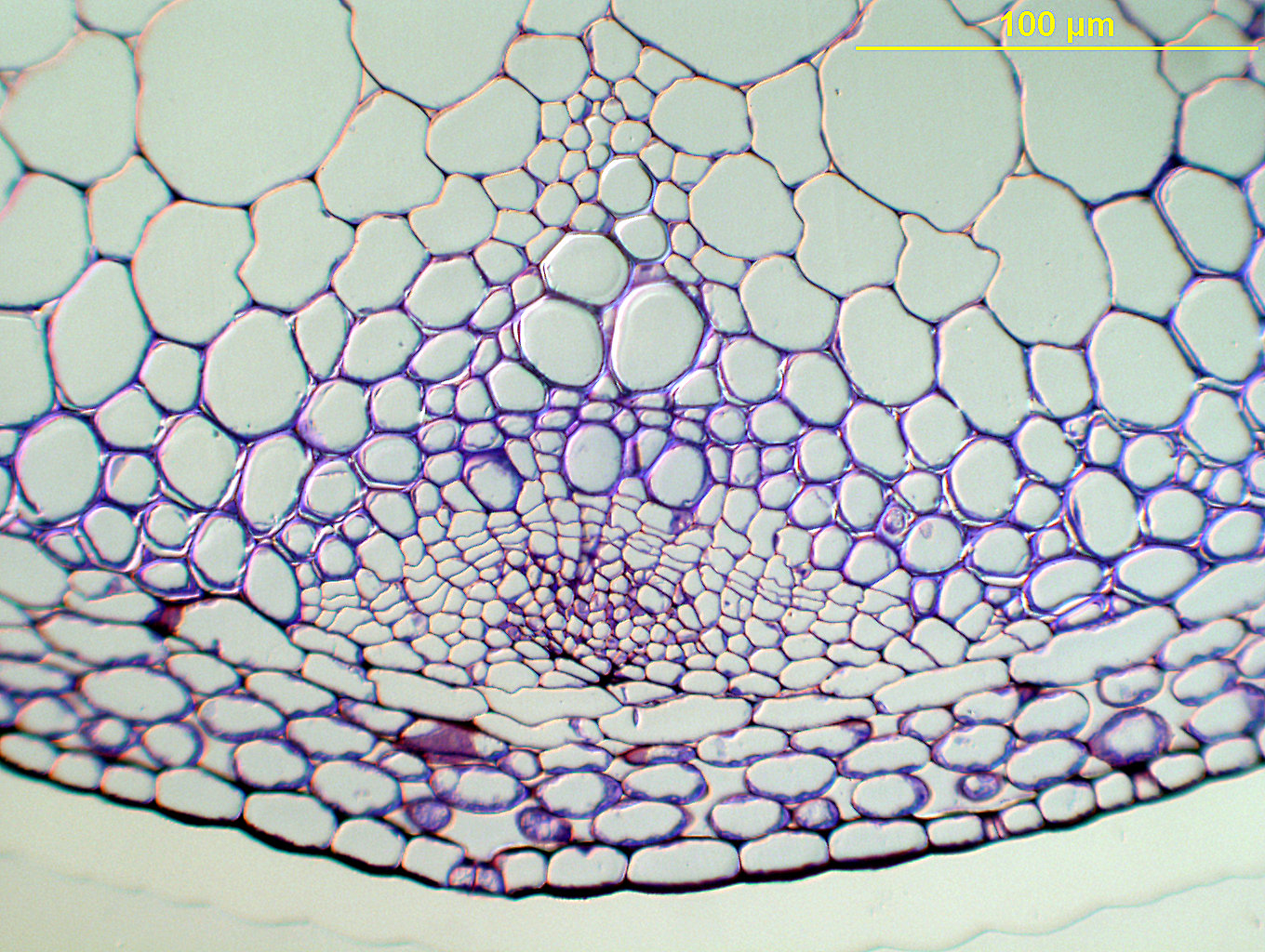
Figure 4. Arabidopsis stem section stained with cresyl violet, prepared as described in this protocol
Notes
- To obtain transversal stem sections a 2-step process is used. First, the stems are horizontally laid down in the mold, at the bottom of the surface, and a first block is made. Then this block is trimmed to leave only a piece of resin containing the stem. This block is oriented vertically in a second mold, which is used to produce a second block where the stem will be properly oriented for sectioning.
- This procedure can be applied to other samples, both from animal and plant origins. Generally, plant tissues require longer infiltration times (minimum 2-3 days), as compared to animal tissues (where normally an overnight infiltration time is sufficient).
Recipes
- Preinfiltration solution
50% absolute ethanol
50% infiltration solution - Infiltration solution
50 ml basic resin/liquid
1 packet (0.5 g) activator
Stir with a magnetic stirrer until completely dissolved
It can be stored at 4 °C for several weeks. - Embedding medium
15 ml infiltration solution
1 ml hardener
Freshly made
Mix and use immediately - Cresyl violet staining medium
1% cresyl violet acetate in distilled water
100 mg of cresyl ciolet acetate
10 ml of distilled water
Acknowledgments
This work was supported by the Ministerio de Ciencia y Tecnología (BIO2011-25687). This protocol was adapted from Giménez et al. (2001) and Chevalier et al. (2014). We thank Inés Poveda for photographic work.
References
- Chevalier, F., Nieminen, K., Sanchez-Ferrero, J. C., Rodriguez, M. L., Chagoyen, M., Hardtke, C. S. and Cubas, P. (2014). Strigolactone promotes degradation of DWARF14, an alpha/beta hydrolase essential for strigolactone signaling in Arabidopsis. Plant Cell 26(3): 1134-1150.
- Gimenez, E., Giraldo, P., Jeffery, G. and Montoliu, L. (2001). Variegated expression and delayed retinal pigmentation during development in transgenic mice with a deletion in the locus control region of the tyrosinase gene. Genesis 30(1): 21-25.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Chevalier, F., Iglesias, S. M., Sánchez, Ó. J., Montoliu, L. and Cubas, P. (2014). Plastic Embedding of Arabidopsis Stem Sections. Bio-protocol 4(20): e1261. DOI: 10.21769/BioProtoc.1261.
Category
Plant Science > Plant cell biology > Cell imaging
Plant Science > Plant developmental biology > Morphogenesis
Cell Biology > Cell imaging > Fixed-tissue imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











