- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Karyotype Analysis
Published: Vol 4, Iss 10, May 20, 2014 DOI: 10.21769/BioProtoc.1129 Views: 29496
Reviewed by: Lin FangFanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
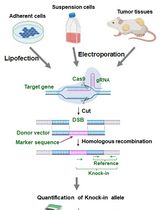
Assay for Site-Specific Homologous Recombination Activity in Adherent Cells, Suspension Cells, and Tumor Tissues
Yuki Yoshino [...] Natsuko Chiba
Apr 5, 2025 2499 Views
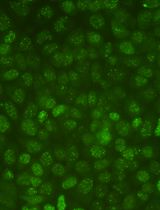
Protocol for Quantifying γH2AX Foci in Irradiated Cells Using Immunofluorescence and Fiji Software
Lu Deng [...] Lingying Wu
Aug 20, 2025 2519 Views
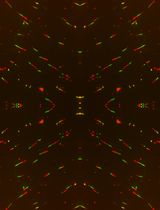

A Quantitative DNA Fiber Assay to Monitor Replication Fork Progression, Protection, and Restart
Debanjali Bhattacharya and Ganesh Nagaraju
Feb 5, 2026 331 Views
Abstract
A chromosome is the structure that organizes DNA and protein in cells. It is a single piece of coiled DNA containing coding and non-coding sequences. Human cells have 23 pairs of chromosomes including 22 pairs of autosomes and one pair of sex chromosome, giving a total of 46 per cell. In tumor cells, chromosomal instability has been considered to be one of the hallmarks of tumor formation. Here we use the karyotype analysis to separate the chromosomes and observe the chromosomes in tumor cells with a microscope.
Materials and Reagents
- Cell lines: FaDu (ATCC, catalog number: HTB-43TM )
- Trypsine (Life Technologies, Gibco®, catalog number: 15400-054 )
- 10% fetal bovine serum (Thermo Fisher Scientific, catalog number: SH30071.03 )
- 1% Penicillin-Streptomycin (10,000 U/ml; 10,000 μg/ml) (Life Technologies, Gibco®, catalog number: 15140-122 )
- Colcemid (10 μg/ml) (Life Technologies, Gibco®, catalog number: 15212-012 )
- Hypotonic solution (0.075 M KCl) (J.T.Baker®, catalog number: 7447-40-7 )
- Glacial acetic acid (J.T.Baker®, catalog number: 64-19-7 )
- Fixative (see Recipes)
- Growth medium (see Recipes)
Equipment
- 10 cm cell culture dishes (Trueline Valve Corpotation, catalog number: TR4002 )
- Centrifuge tube
- Microscope (OLYMPUS, model: DP71 )
- CO2 incubator
- Centrifuge at room temperature (using “relative centrifugal force, rcf”)
- Water bath
Procedure
Day 1
- Seed 5 x 105 cells on 10 cm cell culture dishes for attachment.
- Cells were incubated at 37 °C, 5% CO2 for 48 h.
Day 3
- Add 0.1 ml colcemid, which can collapse mitotic spindles and prevent the completion of mitosis, to each dish and mix gently. Incubated at 37 °C, 5 % CO2 for 2 h.
- Transfer medium to centrifuge tubes from the cell culture dishes. Use PBS to wash the dishes, and remove PBS. Then, add 1 ml 0.1% Trypsine into the dishes at 37 °C, 5% CO2 for 2 min. Also, transfer the mixture (Trypsine and cells) into the centrifuge tubes and mix with the medium which is transferred to centrifuge tubes before we use PBS to wash the dishes. Further, centrifuge at 100 rcf for 10 min at room temperature.
- Discard the supernatant and leave 0.5 ml medium to mix the pellet gently.
- Resuspend the pellet in 5-7 ml 37 °C hypotonic solution and mix thoroughly. Incubate in water bath at 37 °C for 10 min.
- Centrifuge at 100 rcf for 10 min at room temperature.
- Discard the supernatant and leave 0.5 ml solution to mix the pellet gently.
- Resuspend the pellet in 5 ml cold fixative (drop by drop and flick the tube between drops to prevent cell clumping).
- Put the centrifuge tube on ice at least 20 min.
- Centrifuge at 100 rcf for 10 min at room temperature.
- Discard the supernatant and add 3-5 ml cold fixative.
- Repeat steps 9-10 three times or until the supernatant is clear and the pellet become white.
- After the final centrifugation, suspend the cells in a few drops of cold fixative to give a slightly opaque suspension.
- Drop 1-2 drops onto wet and clean slides (Before that, the slides have to be rinsed by 1-2 drops of cold fixative.)
- Dry the slides at room temperature.
- Observe the chromosomes with the microscope (Olympus DP71 with 200x magnification).
- Count the number of chromosomes about 50 cells (A normal human cell have 23 pairs of chromosomes.).
Note: Chromosomes center in a scope, we define that “one cell”.
Representative data
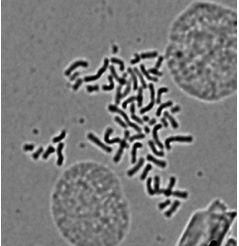
Figure 1. Representative pictures of karyotype analysis in FaDu cell line (see Chou et al., 2013)
Recipes
- Fixative
Methanol and glacial acetic acid (3:1) to be made fresh and chilled before using
- Growth medium
RPMI 1640 supplemented with:
10% fetal bovine serum
1% Penicillin Streptomycin (10,000 U/ml; 10,000 μg/ml)
Acknowledgments
This protocol has been adapted from Chou et al. (2013).
References
- Chou, C. H., Yang, N. K., Liu, T. Y., Tai, S. K., Hsu, D. S., Chen, Y. W., Chen, Y. J., Chang, C. C., Tzeng, C. H. and Yang, M. H. (2013). Chromosome instability modulated by BMI1-AURKA signaling drives progression in head and neck cancer. Cancer Res 73(2): 953-966.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Chou, C. and Yang, M. (2014). Karyotype Analysis. Bio-protocol 4(10): e1129. DOI: 10.21769/BioProtoc.1129.
Category
Cancer Biology > Genome instability & mutation > Cell biology assays > DNA structure and alterations
Cell Biology > Cell imaging > Fixed-cell imaging
Cell Biology > Cell structure > Chromosome
Share
Bluesky
X
Copy link