- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Construction of Deletion-knockout Mutant Fowlpox Virus (FWPV)
Published: Vol 4, Iss 10, May 20, 2014 DOI: 10.21769/BioProtoc.1126 Views: 8955
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
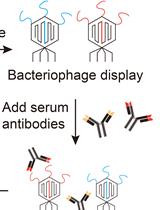
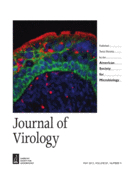
VirScan: High-throughput Profiling of Antiviral Antibody Epitopes
Ellen L. Shrock [...] Stephen J. Elledge
Jul 5, 2022 8659 Views
A Novel Method of Inducible Directed Evolution to Evolve Complex Phenotypes
Ibrahim S. Al’Abri [...] Nathan Crook
Oct 20, 2022 3080 Views
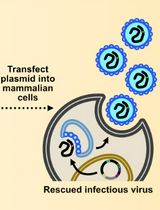
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3072 Views
Abstract
The construction of deletion-knockout poxviruses is a useful approach to determining the function of specific virus genes. This protocol is an adaptation of the transient dominant knockout selection protocol published by Falkner and Moss (1990) for use with vaccinia virus. The protocol makes use of the dominant selectable marker Escherichia coli guanine phosphoribosyltransferase (gpt) gene (Mulligan and Berg, 1981), under the control of an early/late poxvirus promoter. The deletion viruses that are produced no longer contain a selectable marker, which may be preferable for the production of vaccines.
Keywords: KnockoutMaterials and Reagents
- Primary chick embryo fibroblast cells (CEFs)
Notes:- Prepared from specific pathogen free quality embryos (10 days old)
- For a protocol to prepare CEFs see http://cshprotocols.cshlp.org/content/2006/2/pdb.prot4475.full
- Prepared from specific pathogen free quality embryos (10 days old)
- Fowlpox virus (FWPV) (e.g. FP9 strain) (GenBank accession number: AJ581527 )
- Gene specific oligonucleotides
- High fidelity taq polymerase (e.g. Q5, New England Biolabs, catalog number: M0491S )
- Qiaquick PCR purification kit (QIAGEN, catalog number: 28104 )
- T4 DNA ligase (New England Biolabs, catalog number: M0202T )
- Restriction enzymes (New England Biolabs)
- 199 media (Life Technologies, catalog number: 31150-022 )
- DMEM (Life Technologies, catalog number: 11995-065 )
- 10% tryptose phosphate broth (Sigma-Aldrich, catalog number: T8159 )
- Penicillin/Streptomycin (Life Technologies, catalog number: 15140122 )
- Nystatin (Sigma-Aldrich, catalog number: N1638 )
- NewBorn bovine serum (Life Technologies, catalog number: 16010-167 )
- Poxvirus recombination vector e.g. pGNR (as described in Reference 1)
- Transfection reagent e.g. Polyfect (QIAGEN, catalog number: 301105 )
- Mycophenolic acid (Sigma-Aldrich, catalog number: M5255 )
- Xanthine (Sigma-Aldrich, catalog number: X3627 )
- Hypoxanthine (Sigma-Aldrich, catalog number: H9636 )
- Low melting point agarose (Sigma-Aldrich, catalog number: A4018 )
- 10x MEM (Life Technologies, catalog number: 21430-020 )
- L-glutamine (Life Technologies, catalog number: 25030-024 )
- Sodium bicarbonate (Life Technologies, InvitrogenTM, catalog number: 25080-060 )
- Wizard® SV genomic DNA purification kit (Promega Corporation, catalog number: A2361 )
- Taq DNA polymerase (Life Technologies, InvitrogenTM, catalog number: 10342053 )
- MXH media (see Recipes)
- Overlay medium (see Recipes)
- Complete 199 media (see Recipes)
Equipment
- Thermal Cycler
- Microcentrifuge
- T25 cell culture flask
- -80 °C freezer
- Cell culture 6 well plates
- Marker pen
- 37 °C 5% CO2 cell culture incubator
- Sterile screw cap microfuge tubes
- Pipette tips (ART® 200 G)
Procedure
- Construction of recombination plasmid encoding deletion of the gene of interest (GoI)
- Design four oligonucleotide primers
- Primer 1: Forward primer to bind 500 bp upstream of GoI and including 5’ recombination vector compatible restriction site.
- Primer 2: Reverse primer to bind 20 bp downstream of GoI start codon and including a 5’ tail complementary to primer 3.
- Primer 3: Forward primer to bind 30-50 bp upstream of GoI stop codon and designed to disrupt the open reading frame (ORF), by introduction of a frameshift, and including a 5’ tail complementary to primer 2.
- Primer 4: Reverse primer to bind 500 bp downstream of GoI and including a 5’ recombination vector compatible restriction site.
Note: Care should be taken to ensure primers 2 and 3 do not disrupt any putative early/late transcription initiation or termination sites.
- Primer 1: Forward primer to bind 500 bp upstream of GoI and including 5’ recombination vector compatible restriction site.
- Using PCR and a high fidelity polymerase, separately amplify PCR products (using manufacturers' recommendations) from purified FWPV template DNA using primer pairs 1 & 2 and 3 & 4.
- Purify the PCR products from step A2 using Qiaquick PCR purification (according to the manufacturers' recommendations).
- Combine equimolar amounts of the PCR products from step A3 and carry out a further PCR using primers 1 & 4. This will result in a single PCR product of approximately 1.1 kb consisting of a region of the FWPV genome deleted for the GoI.
- Purify the PCR product from step A4 using Qiaquick PCR purification.
- Ligate the PCR product into the recombination vector (pGNR) using the selected/designed restriction enzymes (for guidance see Reference 4).
- Design four oligonucleotide primers
- Making the recombinant fowlpox virus
Due to matching sequences between the recombination vector and fowlpox virus genome homologous recombination occurs resulting in recombinant viruses (A scheme of the recombination events can be found in Figure 4, Reference 2.).
- Seed a T25 tissue culture flask with 3.9 x 106 CEFs (to achieve 70% confluency the following day) in 5 ml 10% serum 199 media.
- Remove media and infect CEFs with FWPV (to give a multiplicity of infection of 3 to 5) in 0.5 ml of serum-free DMEM.
- Incubate for 2 h at 37 °C, 5% CO2 with occasional rocking.
- Add 3 ml 2% serum 199 media and leave for a further 2 h.
- Prepare transfection mix [DNA (from step A6): 2 µg; serum-free DMEM: 200 µl; Polyfect: 10 µl], pipette up and down a few times and leave at room temperature for 15-20 min (http://www.qiagen.com/knowledge-and-support/resource-center/resource-download.aspx?id=bf924409-51f9-4bd0-b63f-a8369c70a331&lang=en).
- Remove media from cells and wash with serum free DMEM media.
- Add 3 ml 2% serum 199 media to cells.
- Add transfection mix directly to flask and rock a few times to mix.
- Incubate 37 °C, 5% CO2 overnight.
- The following day, replace the media in the T25 with 5 ml 2% serum 199 MXH media and incubate 37 °C, 5% CO2 for a further 3 days.
- Release progeny virus from cells by freeze thawing three times using a -80 °C freezer. The protocol can be stopped at this stage and resumed when purification of the virus is required. It is not necessary to centrifuge the freeze-thawed mixture to remove cell debris.
- Seed a T25 tissue culture flask with 3.9 x 106 CEFs (to achieve 70% confluency the following day) in 5 ml 10% serum 199 media.
- Purification of recombinant virus
- Seed a 6 well plate with 1.2 x 107 CEFs (2 x 106 cells in 2 ml/well) (to achieve 100% confluency the following day) in 10% 199 media.
- Prepare 10 fold serial dilutions in serum-free DMEM (10-1 to 10-6) of the freeze-thawed transfected/infected cell supernatant containing progeny virus.
- Remove media from the cells and add 1 ml of the progeny virus dilutions to the wells of the 6 well plate.
- Incubate for 2 h at 37 °C, 5% CO2 with occasional rocking. Remove virus and overlay with 1% agarose overlay media containing MXH. For the preparation of this medium incubate 2x MEM (containing the MXH solutions) at 37 ºC and 2% low gelling temperature agarose at 42 °C. Upon removal of virus from infected CEFs combine the 2x MEM (containing the MXH solutions) and 2% agarose solution in the tissue culture cabinet and overlay cells (3 ml per well of a 6 well plate). Allow agarose to set before incubating at 37 °C, 5% CO2 until plaques are visible (5 to 6 days).
Note: Importantly and unusually, FWPV plaques are visible as opaque, not clear, areas! Their presence can be confirmed by broad field microscopy, preferably under phase contrast. Note also that MXH causes FWPV to replicate more slowly, resulting in smaller plaques so do not just select large plaques.
- By eye mark the location of isolated plaques on the bottom of the plates with a fine marker pen, then pick individual plaques using cut-off filter tips (e.g. ART 200 G max, volume 200 µl) into 500 µl serum free DMEM.
- Freeze thaw the isolated plaque suspension three times (as previously).
- Repeat steps C2-6 three times in order to ensure the virus is sufficiently homogenous.
Note: The above steps isolate single-crossover intermediate recombinant FWPV, each carrying two copies of the GoI, one of which is mutant, the other parental (A scheme of the recombination events can be found in Figure 4 in Reference 2.). In order to resolve the intermediate virus to derivatives lacking gpt and containing either parental or mutant GoI, further plaque purification in the absence of MXH selection is needed. Assuming the GoI is not essential or required for FWPV replication and plaque production in CEF, resolved derivatives containing parental or mutant GoI will be isolated at equal frequencies, as long as primers 1 and 4 are placed equidistant from the GoI. Equivalence of isolation frequency is therefore conversely a test of whether a GoI plays a significant role in FWPV replication and plaque production in CEF.
- Repeat plaque purification three further times in the absence of MXH.
- Test plaques to determine if intermediate recombinant FWPV have resolved by infecting CEFs in 6 well plates with 100 µl freeze thawed plaque purified virus (diluted in 2 ml 2% serum 199 media) and incubating 37 °C, 5% CO2 for 4 days.
- Remove media from infected CEF and wash cells with PBS.
- Using a Wizard® SV genomic DNA purification kit lyse cells using 300 µl of supplied lysis buffer and isolate FWPV DNA (as per manufacturers' instructions - http://www.promega.co.uk/resources/protocols/technical-bulletins/101/wizard-sv-genomic-dna-purification-system-protocol/).
- Using 2 µl of extracted genomic DNA carry out PCR using the original flanking primers (1 & 4) and separate by agarose electrophoresis. The resultant amplicons will represent either parental (GoI + 1,100 bp) or mutant (in this case deleted) genotype (1,100 bp) for the GoI. To ensure that the recombinant is homogeneous and lacking the gpt gene, a further PCR can be carried out using a flanking primer (1 or 4) in combination with an appropriate gpt internal primer. This PCR assay should result in no amplicon.
Note: It may be necessary to carry out further plaque purifications in the absence of MXH until the virus resolves. If it is suspected that the GoI might be essential it would be prudent to perform parallel plaque purifications from 10 to 20 intermediate recombinant plaques. Should PCR screening indicate that all of the derivatives contain parental GoI, then it is probable that the GoI is essential to FWPV replication and plaque production in CEF.
- Seed a 6 well plate with 1.2 x 107 CEFs (2 x 106 cells in 2 ml/well) (to achieve 100% confluency the following day) in 10% 199 media.
Recipes
- MXH media
Stock solutions:
Mycophenolic acid 10 mg/ml in 0.1 M NaOH
Xanthine 10 mg/ml in 0.1 M NaOH
Hypoxanthine 6 mg/ml in 0.1 M NaOH
Filter sterilise all solutions
To 100 ml (final) of medium:
0.25 mlMycophenolic acid (final conc: 0.025 mg/ml)
2.5 ml Xanthine (final conc: 0.25 mg/ml)
0.25 mlHypoxanthine (final conc: 0.015 mg/ml)
Note: Mycophenolic acid is toxic, therefore wear gloves and rinse glassware thoroughly after use.
- Overlay medium
Note: Overlay medium is prepared by the mixing of 2x MEM and 2% low gelling temperature agarose. For media that requires the addition of MXH, this is added to the 2x MEM, before mixing with the agarose.
2x MEM (50 ml):
10 ml 10x MEM
2.3ml 7.5% sodium bicarbonate
1 ml L-Glutamine
0.1 ml Penicillin/Streptomycin
2.5 ml Nystatin
2 ml newborn bovine serum
32.1 ml water
Mix and then hold medium at 37 °C
2% low gelling temperature agarose:
1 g agarose in 50 ml distilled water, autoclaved and held at 42 °C
Note: Stocks of 2% agarose may be prepared in advance and melted just prior to use in a boiling water bath.
- Complete 199 media
1x 199 media:
10% tryptose phosphate broth
1% Penicillin-Streptomycin
2.5% Nystatin
10% (for cell growth) or 2% (for cell maintenance) newborn bovine serum
Acknowledgments
This protocol was adapted from Boulanger et al. (1998); Falkner and Moss (1990); and Laidlaw et al. (2013). This work was funded by the Biotechnology and Biological Sciences Research Council (BBSRC), grants BBS/B/00115/2, BB/E009956/1, and BB/G018545/1.
References
- Boulanger, D., Green, P., Smith, T., Czerny, C. P. and Skinner, M. A. (1998). The 131-amino-acid repeat region of the essential 39-kilodalton core protein of fowlpox virus FP9, equivalent to vaccinia virus A4L protein, is nonessential and highly immunogenic. J Virol 72(1): 170-179.
- Falkner, F. G. and Moss, B. (1990). Transient dominant selection of recombinant vaccinia viruses. J Virol 64(6): 3108-3111.
- Fowlpox virus FP9, GenBank accession number: AJ581527.
- He,F. L. (2011). Standard DNA cloning. Bio-protocol 1(7): e52.
- Laidlaw, S. M., Robey, R., Davies, M., Giotis, E. S., Ross, C., Buttigieg, K., Goodbourn, S. and Skinner, M. A. (2013). Genetic screen of a mutant poxvirus library identifies an ankyrin repeat protein involved in blocking induction of avian type I interferon. J Virol 87(9): 5041-5052.
- Mulligan, R. and Berg, P. (1981). Selection for animal cells that express the Escherichia coli gene coding for xanthine-guanine phosphoribosyltransferase. Proc Natl Acad Sci U S A 78(4): 2072-2076.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Laidlaw, S. M. and Skinner, M. A. (2014). Construction of Deletion-knockout Mutant Fowlpox Virus (FWPV). Bio-protocol 4(10): e1126. DOI: 10.21769/BioProtoc.1126.
Category
Microbiology > Microbial genetics > Mutagenesis
Molecular Biology > DNA > Mutagenesis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link