- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vivo Lineage-tracing Studies in a Cancer Stem Cell Population in Neuroblastoma
Published: Vol 4, Iss 8, Apr 20, 2014 DOI: 10.21769/BioProtoc.1104 Views: 10380
Reviewed by: Lin FangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
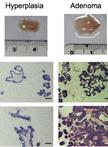
Isolation and Immortalization of Fibroblasts from Different Tumoral Stages
Fernando Calvo [...] Erik Sahai
Apr 5, 2014 20347 Views
Cell-derived Matrix Assays to Assess Extracellular Matrix Architecture and Track Cell Movement
Kendelle J. Murphy [...] David Herrmann
Dec 20, 2022 3411 Views
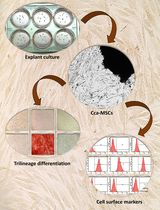
Isolation and Characterization of Cervical Cancer-Associated Mesenchymal Stem Cells From Primary Tumors Using Explant Culture
Surbhi Singla [...] Shalmoli Bhattacharyya
Jun 20, 2025 2811 Views
Abstract
Tumors are comprised of heterogeneous subpopulations that may exhibit differing capacity for differentiation, self-renewal, and tumorigenicity. In vivo lineage-tracing studies are a powerful tool for defining the role of tumor subpopulations in tumor growth and as targets for therapeutic agents. This protocol describes using a neuroblastoma cancer cell line transduced with two different fluorescent proteins (GFP and tdTomato) to track the specific contributions of cells expressing the GCSF receptor (CD114+) or not (CD114-) on tumor growth in vivo.
Materials and Reagents
- Human neuroblastoma cell lines (NGP, NB-1691, IMR-32)
Note: A cell line is transduced with two different fluorescent proteins, for example GFP (Clontech, catalog number: 632370 ) and tdTomato (Clontech, catalog number: 632534 ), such that there is a GFP positive line and a tdTomato line of the same cell type. In this manner, subpopulations of the same cell type (i.e. GCSF-R positive and GCSF-R negative cells) can be traced. If in vivo monitoring of tumor growth via bioluminescent imaging is desired, cell lines should also be transfected or virally transduced with commercially available vectors, e.g. pGL2-Control Vector (Promega Corporation) to express a luminescent reporter gene. For a detailed description and protocol of in vivo bioluminescent imaging, please refer to Reference 2.
- 4-6 week old female non-obese diabetic/severe combined immunodeficient (NOD/SCID) mice (Taconic, model number: NODSCF , http://www.taconic.com/NODSC)
- 293T cells
- RPMI medium 1640 (Life Technologies, catalog number: 11875-101 )
- 10% (v/v) fetal bovine serum (FBS) (Life Technologies, catalog number: 16000044 )
- 1% (v/v) penicillin/streptomycin (Life Technologies, catalog number: 15140122 ) (100x P/S; final concentration 100 units/ml penicillin and 100 mg/ml streptomycin)
- 1% (v/v) of 100x L-glutamine (Life Technologies, catalog number: 25030081 ) (final concentration 2 mM)
- Phosphate-buffered saline (PBS) (sterile) (Life Technologies, catalog number: 70011069 )
- 0.25% trypsin/EDTA (Life Technologies, catalog number: 25200056 )
- Collagenase I (Sigma-Aldrich, catalog number: C0130 ) [prepare a solution containing 10,000 Collagenase Digestion Unites (CDU/ml) in PBS]
- Dispase II (Roche Diagnostics, catalog number: 04942078001 ) (prepare a solution containing 32 mg/ml Dispase II in PBS)
- DNase I (EMD Millipore, Calbiochem®, catalog number: 260913 ) (prepare a solution containing 5 MU/ml DNase I)
- FuGENE 6 (Promega Corporation, catalog number: E2691 )
- Opti-MEM Reduced Serum Medium (Life Technologies, catalog number: 31985-062 )
- PE conjugated anti-CD 114 (GCSFR) antibody (BD Biosciences, catalog number 554538 )
- Cell culture medium (see Recipes)
- Sterile FACS buffer (see Recipes)
- PEB Buffer (see Recipes)
Equipment
- Fluorescence-activated cell sorter (e.g. DAKO Cytomation MoFlo 9-color cell sorter)
- 37 °C, 5% CO2 tissue culture incubator
- Refrigerated centrifuge
- Class 2 biological safety cabinet with laminar flow hood
- 70 μm cell strainer (Thermo Fisher Scientific, catalog number: 22-363-548 )
- T-75 culture flask or 10 cm dish
- Anesthesia machine/chamber with nose cone appropriate for mice (Surgivet or VetEquip)
- Fluorescent microscope
- Surgical instruments
- 5.5-in Mayo-Hegar or similar surgical needle holder (Millennium Surgical or Roboz Surgical Instrument)
- Sterile gloves (Thermo Fisher Scientific)
- Disposable sterile scalpel blade (#10) (Millennium Surgical or Roboz Surgical Instrument)
- 27-G needle
- Sterile 1-cc slip tip syringe
- Polysorb 4-0 sutures with RB-1 tapered needle (U.S. Surgical)
- 9-mm wound clips (VWR International)
- Rodent ear tags (National Band & Tag Company)
- 5.5-in Mayo-Hegar or similar surgical needle holder (Millennium Surgical or Roboz Surgical Instrument)
Procedure
Note: All steps must be performed sterilely under tissue culture hood since the cells will be injected into immunodeficient mice.
- From a given cell line, establish two lines that express two different fluorescent proteins, e.g. NGP/GFP+ and NGP/tdTomato+.
Note: Numerous transduction protocols exist. We used a lentiviral transduction protocol, using FuGENE 6 and Opti-MEM Reduced Serum Medium to transfect 293T cells with the viral packaging plasmids and our construct of interest. Viral supernatant was collected at 48 and 72 h. Viral supernatant was then used to transduce our neuroblastoma cell line of interest. Cells were incubated with viral supernatant for 24 h and then selected with antibiotic according to the antibiotic resistant gene contained in the plasmid until all non-transduced cells died (4-5 days).
- Harvest cells transduced with fluorescent protein using 0.05% Trypsin.
- For a T-75, use 1.5 ml Trypsin and 8.5 ml media.
- For a 10 cm dish, use 0.5 ml Trypsin and 4.5 ml media.
- For a T-75, use 1.5 ml Trypsin and 8.5 ml media.
- Transfer to a 15 ml tube and spin down at 250 x g for 5 min.
- Resuspend cells in either sterile FACS Buffer or PBS.
- Resuspend in 5 ml for a 10 cm dish or 10 ml for a T-75.
- Count cells.
Note: Total number of cells desired depends on how many cells are planned for injection into each mouse and how many mice are being injected for each experiment. For neuroblastoma cell lines, the number of cells injected per mouse can range from 1,000 to 1.0 x 106. For the in vivo lineage-tracing studies performed in Reference 1, 1,000 cells were injected into each mouse.
- Label FACS tubes.
- Put desired number of cells in FACS tubes and spin down to wash.
- Any cells not used for sorting can be put back into culture at this point.
- Aspirate the wash. Be very careful not to aspirate the cells. Sometimes they do not adhere as a pellet very well, and they slide around.
- If you have to leave a little bit of volume in the tube in order to save the cells, you can add an extra wash to be sure all media/Trypsin has been removed.
- Add 2 ml sterile FACS buffer to tubes, vortex, and spin down to wash again. At least 2 washes are necessary.
- Aspirate the FACS buffer.
- Resuspend cells in FACS buffer in a final concentration of 1 x 107 cells/ml for incubation with primary antibodies. To detect the GCSF-R (CD114), we used PE conjugated anti-CD 114 (GCSFR) antibody. We used 1 μg of antibody per 1.0 x 106 cells in a total volume of 100 µl, however the concentration of antibody varies based on the particular antibody used and the antigen being detected.
- For 1 x 106 cells, resuspend in 100 µl total (subtract out the volume of antibody that will be added). If staining more than 1 x 106 cells, you can scale up.
- Add antibody, mix well, and immediately keep away from light.
- If adding 5 µl of antibody, first resuspend cells in 95 µl of FACS buffer for a total volume of 100 µl.
- Incubate cells with antibody for 30 min on ice in the dark.
- Can use an ice bucket with a lid in the tissue culture hood.
- If staining a large number of cells, it is necessary to vortex them a few times during this incubation because the cells will start to settle to the bottom of the tube.
- During this time, put FACS buffer on ice or back in the 4 °C. This should be kept cold.
- Can use an ice bucket with a lid in the tissue culture hood.
- Add 2 ml FACS buffer to tubes and vortex. Spin down to wash.
- Aspirate the wash. Be very careful not to aspirate the cells. Sometimes they do not adhere as a pellet very well, and they slide around.
- If you have to leave a little bit of volume in the tube in order to save the cells, you can add an extra wash to be sure all of the extra stain has been removed.
- Repeat steps 13-14.
- Resuspend cells in FACS buffer for sorting.
- For 1 x 106 cells, resuspend in 0.5 ml. You can scale up with a larger number of cells.
- Run cells through a 40 µm filter just before sorting so clumps of cells won’t clog the sorting machine.
- Sort cells of interest using fluorescence-activated cell sorter, e.g. DAKO Cytomation MoFlo 9-color cell sorter, for example tdTomato+/GCSFR+ cells and GFP+/GCSFR- cells.
- Mix cells of differing lineages in desired ratio for injection. For example, NGP/tdTomato+/GCSFR+ cells mixed with NGP/GFP+/GCSFR- cells mixed in a 1:1 ratio.
- Inject cells sterilely into immunodeficient mice. For a detailed protocol, see Patterson et al. (2011).
- After tumors have grown for desired time period, sacrifice mice and harvest tumors.
Note: Time to tumor growth will depend on the number of cells and the cell type injected. If cells have been transduced with a luciferase gene, bioluminescence can be used to monitor tumor growth. Otherwise, mice can be examined and tumors palpated and measured by calipers weekly to detect tumor growth. For an average neuroblastoma cell line, if 1.0 x 106 cells are injected, palpable tumors are typically present by 4 weeks. For more detailed information, please refer to Hsu et al. (2013) and Patterson et al. (2011).
- Mice may be sacrificed by CO2 asphyxiation. A necropsy is performed and the tumor resected. Fresh tumors should be kept on ice and in the dark until after gross examination under fluorescent microscope. Tumors should be examined immediately after resection to maintain cell viability.
- Tumors may be examined grossly under fluorescent microscope to determine dominant cell lineage, e.g. if tumor is predominately GFP+ or tdTomato+.
- To quantify contribution of each lineage to total tumor make-up, prepare tumor cells for flow cytometry.
- Place tumor sample immediately in cell culture media (DMEM or RPMI-1640). Mince tumor into very small pieces (approximately 5 mm) using a sterile scalpel blade or sterile scissors in a 10 cm petri dish with a small amount of media.
- Gently mechanically disrupt tumor as follows: Place a 70 μm cell strainer in a new petri dish, put small amount of media in dish (enough to coat bottom of dish). In batches, pass media containing tumor sample (from step 1) through cell strainer (gently push tumor through the cell strainer using the back end of 10 ml syringe plunger).
- Collect cell suspension in 15 ml tube. Centrifuge. Resuspend in 5 ml media without serum or PBS.
- Add 150 μl Collagenase I (10,000 CDU/ml) and 150 μl Dispase II (32 mg/ml) and 2 μl DNase I to solution.
- Incubate sample for 20-30 min at 37 °C. Shake or vortex the tube every 10 min during incubation period or place on tube rotator.
- Centrifuge sample, aspirate supernatant.
- Resuspend the sample in 5 ml of PEB buffer and apply to a cell strainer (70 μm mesh size) placed in a 50 ml tube. Wash the cell strainer with 5 ml PEB buffer.
- Discard cell strainer and add PEB buffer to a final volume of 50 ml.
- Centrifuge cell suspension at 300 x g for 10 min. Aspirate supernatant completely.
- Resuspend cells in FACS buffer to analyze. For 1.0 x 106 cells, resuspend in 0.5 ml of FACS buffer.
- Analyze cells on fluorescence activated flow cytometer to specifically quantify each lineage, i.e. percentage of GFP+ versus tdTomato+ cells in each tumor.
Recipes
- Cell culture medium
Serum-free medium (e.g., RPMI medium 1640)
Supplemented with 10% (v/v) fetal bovine serum
1% (v/v) penicillin/streptomycin
1% (v/v) of 100x L-glutamine (final concentration 2 mM)
- Sterile FACS buffer
Sterile PBS with 1% sterile FBS
No sodium azide
Must keep at 4 °C
Do not keep longer than 1 week
- PEB Buffer
PBS (pH 7.2)
0.5% BSA
2 mM EDTA
Keep buffer cold (2-8 °C)
Acknowledgments
This work was conducted with support from the American Cancer Society (J.M. Shohet), Alex's Lemonade Stand Foundation (J.M. Shohet), Gillson-Longenbaugh Foundation (J.M. Shohet), Children's Neuroblastoma Research Foundation (J.M. Shohet), St. Baldrick's Foundation (E.S. Kim), Texas Children's Hospital Department of Surgery Seed Grant (E.S. Kim), and Texas Children’s Hospital Department of Surgery institutional support (E.S. Kim). We would also like to acknowledge the assistance of the Texas Children’s Cancer Center Flow Cytometry Core.
References
- Hsu, D. M., Agarwal, S., Benham, A., Coarfa, C., Trahan, D. N., Chen, Z., Stowers, P. N., Courtney, A. N., Lakoma, A., Barbieri, E., Metelitsa, L. S., Gunaratne, P., Kim, E. S. and Shohet, J. M. (2013). G-CSF receptor positive neuroblastoma subpopulations are enriched in chemotherapy-resistant or relapsed tumors and are highly tumorigenic. Cancer Res 73(13): 4134-4146.
- Patterson, D. M., Shohet, J. M. and Kim, E. S. (2011). Preclinical models of pediatric solid tumors (neuroblastoma) and their use in drug discovery. Curr Protoc Pharmacol Chapter 14: Unit 14 17.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Hsu (Maiden name: Danielle M. Patterson), D. M., Shohet, J. M. and Kim, E. S. (2014). In vivo Lineage-tracing Studies in a Cancer Stem Cell Population in Neuroblastoma. Bio-protocol 4(8): e1104. DOI: 10.21769/BioProtoc.1104.
Category
Cancer Biology > Invasion & metastasis > Tumor microenvironment > Tissue isolation
Cancer Biology > Proliferative signaling > Tumor formation > Self-renewal
Stem Cell > Adult stem cell > Cancer stem cell
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










