- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Electrophoresis Mobility Shift Assay
Published: Vol 4, Iss 7, Apr 5, 2014 DOI: 10.21769/BioProtoc.1099 Views: 18658
Reviewed by: Ru ZhangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
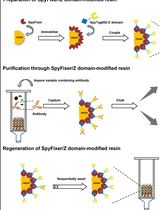
Split-Chloramphenicol Acetyl Transferase Assay to Study Protein-Protein Interactions and Ubiquitylation in Escherichia coli
Amir Florentin [...] Gali Prag
Sep 5, 2022 3416 Views
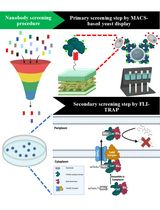
Isolation of Antigen-Specific Nanobodies From Synthetic Libraries Using a Protein Selection Strategy That Combines MACS-Based Screening of YSD and FLI-TRAP
Apisitt Thaiprayoon [...] Dujduan Waraho-Zhmayev
Jan 20, 2026 519 Views
Abstract
Protein (transcription factors and/or transcription cofactors)-binding to DNA is a critical event in regulation of transcription. Electrophoresis Mobility Shift Assay (EMSA), also known as gel shift assay, is a useful tool to detect protein- or protein complex-DNA/RNA interaction and to evaluate DNA binding specificity of transcription factors in vitro. Here we describe a simple method for EMSA with fluorescent dye-bound oligo DNA probes and recombinant protein expressed in bacterial cells. Using fluorescent dye instead of radioisotope enables easy handling and long-term storage of labelled-probes without reduction of detection sensitivity.
Materials and Reagents
- Oligo DNA 5’ end-labeled with IRDye 700 or IRDye 800 (sense strand) (Integrated DNA Technologies)
- Non-labelled oligo DNA (both sense and antisense strands)
- Non-labelled mutated oligo DNA (both sense and antisense strands)
- Recombinant DNA-binding proteins expressed in Escherichia coli (E. coli) (10 ng/µl in protein storage buffer)
- Sterile distilled water (SDW)
- Odyssey infrared EMSA kit (LI-COR, catalog number: 829-07910 )
- Poly(dI-dC) (Sigma-Aldrich, catalog number: 4929 )
- Tris
- Boric acid
- EDTA 2Na
- NaCl
- HCl
- Acrylamide
- N,N’-Methylene-bisacrylamide
- Glycerol
- Triton X-100
- Phenylmethylsulfonyl fluoride (PMSF)
- β-mercaptoethanol
- Ammonium persulfate (APS)
- N,N,N',N'-tetramethylethylenediamine (TEMED)
- 10x TBE( see Recipes)
- 4% native polyacrylamide gel (see Recipes)
- Native-PAGE running buffer (see Recipes)
- Protein storage buffer (see Recipes)
Equipment
- Odyssey CLx Infrared Imaging System (LI-COR)
- A set of devices for polyacrylamide gel electrophoresis
- Power supply
- Refrigerator or cold room
- Heat block
Procedure
- Suspend lyophilized oligo DNA with dH2O and mix them as follows (final concentration is 50 µM each).
Probe: labelled and complementary non-labelled oligo DNAs
Competitor: non-labelled and complementary non-labelled oligo DNAs
Mutated competitor: mutated non-labelled and complementary non-labelled oligo DNAs
- Heat at 100 °C for 10 min on heat block for denature.
- Turn off heat block and leave denatured DNAs on the block until room temperature.
- Dilute probes to 50 nM with dH2O.
- Prepare 4% native polyacrylamide gel and 0.5x TBE.
- Pre-electrophoresis for 30 min at 150 V at 4 °C.
- Prepare reaction mixtures as follows during pre-electrophoresis: Note: Tube numbers indicate the vial numbers in Odyssey Infrared EMSA kit.
Negative Control
Probe
Competitor
Mutated competitor
Protein (10 ng/µl)
-
1 µl
1 µl
1 µl
Probe (50 nM)
1 µl
1 µl
1 µl
1 µl
Competitor (50 µM)
-
-
1 µl
-
Mutated competitor (50 µM)
-
-
-
1 µl
10 x buffer (tube 1)
2 µl
2 µl
2 µl
2 µl
25 mM DTT/2.5% Tween-20 (tube 2)
2 µl
2 µl
2 µl
2 µl
1 µg/µl Poly (dIdC) (tube 3)
1 µl
1 µl
1 µl
1 µl
50% Glycerol (tube 5)
2 µl
2 µl
2 µl
2 µl
100 mM MgCl2 (tube 8)
1 µl
1 µl
1 µl
1 µl
dH2O
11 µl
10 µl
9 µl
9 µl
- Place at room temperature for 20 min in dark.
Note: Avoid light during reaction not to reduce signal intensity.
- Add 2 µl of 10x Orange Dye (tube 10) to reaction mixture after reaction.
- Wash gel wells with electrophoresis buffer (0.5x TBE) after pre-electrophoresis.
- Load the samples on gel and run the gel at 150 V for 2.5 h at 4 °C in dark until the Orange Dye migrates to the bottom of the gel.
- Remove gel from glass plate and place it directly on Odyssey.
- Adjust focus offset of Odyssey to 1/2 of gel thickness and scan.
Note: Carefully remove air bubbles between gel and Odyssey.
Look at supplemental Figure 19 of Reference 1 as a representative EMSA result.
Recipes
- 10x TBE
Consisting of 890 mM Tris, 890 mM boric acid and 20 mM EDTA 2Na (pH 8.3)
Mix:
108 g of Tris base
55 g of boric acid
3.7 g of EDTA 2Na
Add dH2O to 1 L
Autoclave and stored at room temperature
- 4% native polyacrylamide gel
Consisting of 4% acrylamide, 0.5x TBE, 2.5% glycerol, 0.1% APS, 0.1% TEMED
Mix:
2.67 ml of 30% acrylamide (acrylamide: bisacrylamide = 29:1)
1 ml of 10x TBE
1 ml of 50% glycerol
0.2 ml of 10% APS
20 µl of TEMED
Add dH2O to 20 ml
- Native-PAGE running buffer
0.5x TBE
- Protein storage buffer
Consisting of 10 mM Tris-HCl (pH7.5), 150 mM NaCl, 0.1% Triton X-100, 1 mM PMSF, 0.05% β-mercaptoethanol, 50% glycerol
Mix:
1 ml of 1 M Tris-HCl (pH 7.5)
3 ml of 5 M NaCl
1 ml of 10% Triton X-100
1 ml of 100 mM PMSF
50 µl of β-mercaptoethanol
50 g of glycerol
Add dH2O to 100 ml
Acknowledgments
This protocol is adapted from Nakata et al. (2013).
References
- Nakata, M., Mitsuda, N., Herde, M., Koo, A. J., Moreno, J. E., Suzuki, K., Howe, G. A. and Ohme-Takagi, M. (2013). A bHLH-type transcription factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, acts as a repressor to negatively regulate jasmonate signaling in Arabidopsis. Plant Cell 25(5): 1641-1656.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Nakata, M. and Ohme-Takagi, M. (2014). Electrophoresis Mobility Shift Assay. Bio-protocol 4(7): e1099. DOI: 10.21769/BioProtoc.1099.
Category
Molecular Biology > DNA > DNA-protein interaction
Biochemistry > Protein > Interaction > EMSA
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link