- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measuring Homologous Recombination Frequency in Arabidopsis Seedlings
Published: Vol 4, Iss 7, Apr 5, 2014 DOI: 10.21769/BioProtoc.1094 Views: 13321
Reviewed by: Masahiro MoritaRu ZhangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

CRISPR-Cas9 Protocol for Efficient Gene Knockout and Transgene-free Plant Generation
Enzo A. Perk [...] Ignacio Cerrudo
Jun 5, 2024 3506 Views

Optimized Protocol for DNA Extraction in Three Theobroma Species
Angie F. Riascos-España [...] Pedro A. Velasquez-Vasconez
May 5, 2025 2135 Views
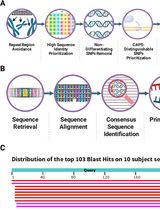
CAPS-Based SNP Genotyping for Nitrogen-Response Phenotypes in Maize Hybrids
Jannis Jacobs [...] Peter K. Lundquist
Dec 20, 2025 586 Views
Abstract
Somatic homologous recombination (SHR) is a major pathway of DNA double-strand break (DSB) repair, in which intact homologous regions are used as a template for the removal of lesions. Its frequency in plants is generally low, as most DSB are removed by non-homologous mechanisms in higher eukaryotes. Nevertheless, SHR frequency has been shown to increase in response to various chemical and physical agents that cause DNA damage and/or alter genome stability (reviewed in March-Díaz and Reyes, 2009). We monitor the frequency of SHR in transgenic Arabidopsis seedlings containing recombination substrates with two truncated but overlapping parts of the β-glucuronidase (GUS) reporter gene (Orel et al., 2003; Schuermann et al., 2005). Upon an SHR event, a functional version of the transgene can be restored (Figure 1A). A histochemical assay applicable to whole plantlets allows the visualization of cells in which the reporter is restored, as the encoded enzyme converts a colorless substrate into a blue compound. This type of reporter has been extensively used to identify gene products required for regulating SHR levels in plants. We analyze plants stimulated for SHR by treatments with DNA damaging agents (bleocin, mitomycin C and UV-C) and compare them to non-treated plants.
Keywords: Homologous recombinationMaterials and Reagents
- Arabidopsis thaliana seeds of reporter lines IU.GUS-8 and DGU.US-1 (Figure 1) (Orel et al., 2003) [introgressed into various genetic backgrounds and genotyped for homozygosity of reporters and mutations (Rosa et al., 2013). As an example, we use wild type Columbia-0, and mutants arp6-3 and swc6-1. Both mutants lack subunits of the Arabidopsis homolog of the SWR1 complex and, additional to being sensitive to DNA damaging agents (Rosa et al., 2013), have pleiotropic developmental defects (for more information see Schuermann et al., 2005)]
- Sodium hypochlorite (Sigma-Aldrich, catalog number: 425044 )
- Tween-80 (Sigma-Aldrich, catalog number: 4780 )
- Sterile H2O
- Solid growth medium
- Liquid plant growth medium (same as solid growth medium but without agar)
- Hydroxyurea
- 70% ethanol
- Seed sterilization solution (see Recipes)
- Bleocin (commercial name for bleomycin) (EMD Millipore, catalog number: 203408 ) (see Recipes)
- Mitomycin C (Duchefa Biochemie BV, catalog number: M0133 ) (see Recipes)
- GUS staining solution (see Recipes)
- 1 M Sodium Phosphate Buffer (see Recipes)
Equipment
- Plastic petri dishes for plant culture (see Notes) (round 200 x 15 mm, with 20 ml of solid growth medium)
- Sterile hood, preferably a biological safety cabinet to avoid exposure to the genotoxins
- Bench top block shaker (e.g. Eppendorf, Thermomixer®)
- UV crosslinker (254-nm UV light bulbs, 15 watts each) (Stratagene, model: Stratalinker 2400 )
- Forceps
- Box or aluminum foil
- Parafilm
- 1.5 ml Eppendorf tubes
- 50 and 15 ml Falcon tubes
- Vacuum applicator (e.g. desiccator)
- Plant growth facilities with 16-h-light/8-h-dark cycles, at 21 °C
- Incubator at 37 °C
- Stereomicroscope
Procedure
- Wash the seeds in 1.5 ml Eppendorf tubes with 1 ml of the sterilization solution, shake in a benchtop block shaker at 300 rpm for 6 min at room temperature. Allow seeds to set in the bottom of the Eppendorf and remove supernatant. Wash with 1 ml of sterile water, 5 times for 5 min in the shaker. After the final wash, remove supernatant and dry overnight in the closed sterile hood, leaving the tubes open.
Note: Sterilize approximately 200 seeds per genotype and treatment. All subsequent steps involving seed or plantlet manipulation should be carried in the sterile hood. - Plate seeds on 200 x 15 mm petri dishes with growth medium (≈ 75 to 100 per plate and 2 plates per genotype and treatment), using flame-sterilized and cooled forceps. Stratify in the dark (in a box or wrapped in aluminum foil) at 4 °C for 2 to 4 days and transfer to the growth chambers with illumination.
Note: Any other sterile plating techniques are suitable. - Treatments
- Bleocin and mitomycin C: Grow plants until day 7. Add 10 ml of drug-free liquid media, or media with 0.1 μg/ml of bleocin or 10 μg/ml of mitomycin C. Seal with parafilm and return to normal growth conditions until analysis. Shaking is not necessary.
Note: Grow until day 7 = 7 days after transfer to the growth chambers. - Hydroxyurea: Plate seeds on solid growth plates, without or with 1 mM of the chemical, and grow at standard conditions until analysis.
- UV-C: Grow seedlings until day 4 of growth and treat with 8 kJ/m2 UV-C in a Stratalinker. Return to standard conditions until analysis.
Note: Before the treatment sterilize the interior of the Stratalinker with 70% Ethanol and run it for 3 min with the maximum dose of UV-C. Then place plates inside and remove the lid, keeping it inside the Stratalinker. Quickly close the door, to avoid contamination, and run the desired dosage of UV-C. When the treatment is finished close the plates still inside the Stratalinker and return them to normal growth conditions.
- Bleocin and mitomycin C: Grow plants until day 7. Add 10 ml of drug-free liquid media, or media with 0.1 μg/ml of bleocin or 10 μg/ml of mitomycin C. Seal with parafilm and return to normal growth conditions until analysis. Shaking is not necessary.
- GUS histochemical assay
- With forceps, carefully collect the 12-day-old plants of each plate in 15 ml Falcon tubes and submerge them in 10 ml of GUS staining solution. Be careful not to break the roots, cotyledons or leafs. Vacuum infiltrate for 15 min and incubate overnight at 37 °C.
- De-stain the plants in 70% ethanol overnight at 37 °C. Continue to change the ethanol twice a day until most plants tissue is white.
- Under a stereomicroscope score the number of GUS positive blue sectors indicating recombination events per whole plant (Figure 1B). To analyze the data, generate histograms with the frequency of seedlings with different numbers of blue sectors, and the total frequency in each line and treatment (Figure 2).
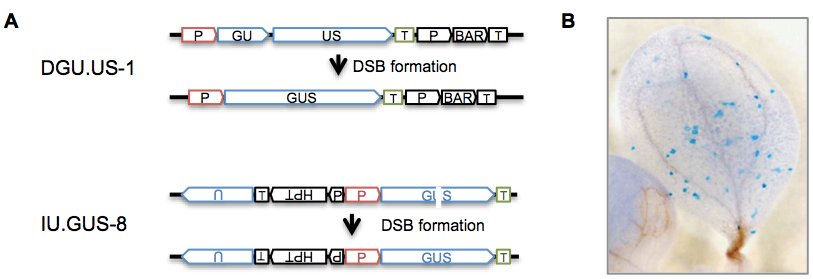
Figure 1. Principle of the assay. (A) Recombination substrates in reporter lines for SHR DGU.US-1 and IU.GUS-8 (adapted from Reference 2). These lines were designed to distinguish different somatic recombination pathways (single-strand annealing and synthesis-dependent strand annealing) upon DSB repair after expression of the endonuclease I-SceI (Orel et al., 2003; for more information about different SHR pathways see Reference 1). However, they are used here to measure SHR without I-SceI but after different exogenous DNA damage treatments. The DGU.US-1 substrate contains two fragments of the GUS ORF with homologous parts (GU, US) in direct orientation, harboring an overlap of 557 bp. In IU.GUS-8 , two fragments of the GUS ORF (“GU”, “US”; blue) in direct orientation are separated, rendering the gene not functional. A fragment of 1087 bp partially homologous to “GU” and “US”, and containing “U” is located upstream in inverted orientation. SHR events between the repeats can restore a functional gene (GUS) in different ways: in line DGU.GUS-1, the sequence in common between the two repeats is deleted, while HR between indirect repeats in IU.GUS-8 results in the restoration of the GUS gene. P: constitutive 35S promoter of cauliflower mosaic virus; T: nopaline synthase terminator; HPT: hygromycin resistance gene, BAR: phosphinothricin acetyltransferase resistance gene, GUS: β -glucuronidase gene. (B) Histochemical GUS staining allows to visualize cells in which the GUS gene was restored by SHR (from Orel et al., 2003).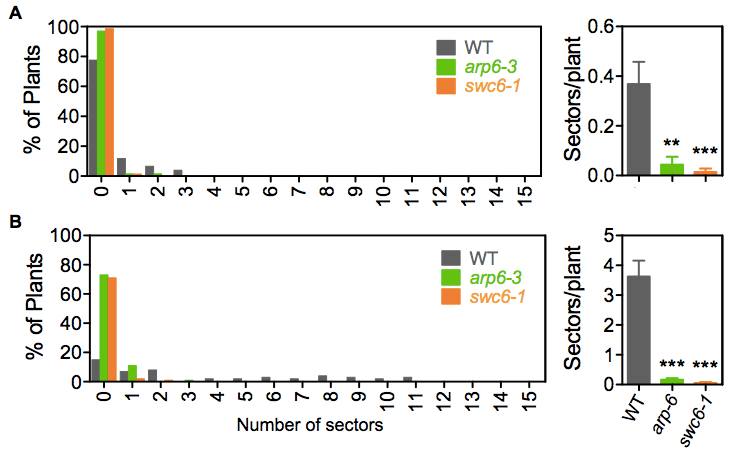
Figure 2. Examples of SHR frequency measurements. Distribution of seedlings with different numbers of blue sectors and total frequency in line IU.GUS-8. (A) Mock and (B) 1 μg/ml Bleocin treatment. Sectors/plant refers to the average number of blue sectors, corresponding to recombination events, in the analyzed seedling population. Error bars correspond to the SE. SHR frequencies in each mutant population were significantly different to their wild-type (WT) counterpart, with a P value < 0.001. Asterisks indicate the significance between treated and mock populations according to P values from unpaired t tests: ***P < 0.001 and **0.001 < P < 0.01. - With forceps, carefully collect the 12-day-old plants of each plate in 15 ml Falcon tubes and submerge them in 10 ml of GUS staining solution. Be careful not to break the roots, cotyledons or leafs. Vacuum infiltrate for 15 min and incubate overnight at 37 °C.
Notes
- For all treatments, always keep the plates belonging to one experiment (mock + treatment) growing side-by-side. For example, take plates for mock and UV-C treatment out of the growth chambers at the same time.
- In this experiment we used Germination Media (GM) described in Reference 6 (detailed protocol available at http://www.gmi.oeaw.ac.at/research-groups/ortrun-mittelsten-scheid/research-groups/ortrun-mittelsten-scheid/gm-medium-protocol). We have not tested the use of regular MS plates but have no reason to believe the media will be a determining factor as long as the plates used in each experiment come from the same batch of media.
Recipes
- Seed sterilization solution
Consisting of 5% sodium hypochlorite and 0.05% Tween-80
Mix 500 µl of sodium hypochlorite and 5 µl of Tween-80 in 10 ml of sterile H2O - Bleocin (stock 10 mg/ml)
Dissolve 100 mg of Bleocin in 10 ml of H2O
Make 0.2 ml aliquots and keep at -20 °C until use
Avoid thawing/freezing cycles
Dilute to the desired concentration in liquid media, using disposable 50 ml falcon tubes - Mitomycin C (stock 0.5 mg/ml)
Dissolve 2 mg of Mitomycin C in 4 ml of H2O immediately before use
Dilute to the desired concentration in liquid media, using disposable 50 ml falcon tubes - GUS staining solution
Consisting of 1 mM sodium phosphate buffer (pH 7); 10 mM EDTA, 0.1% Triton-X, 100 mg/ml chloramphenicol; 2 mM potassium ferrocyanide; 2 mM potassium ferricyanide; 0.5 mg/ml X-glucuronide
Mix:
10 ml of 1M Sodium Phosphate Buffer Solution (pH 7) (see below)
4 ml of 0.25 M EDTA solution pH 8
0.5 ml 20% (w/v) Triton X-100
0.2 ml of 50 mg/ml Chloramphenicol
1 ml of 0.2 M Potassium Ferricyanide
1 ml of 0.2 M Potassium Ferrocyanide
2.5 ml 20 mg/ml X-Gluc (kept in the dark at -20 °C)
Bring up to 100 ml with MilliQ water, filter-sterilize (0.2 micron filter) and keep at 4 °C wrapped in aluminum foil. If stored under these conditions, this solution will be stable for up to one month. - 1 M Sodium Phosphate Buffer (final pH 7)
Make 200 ml of 1 M Sodium Dihydrogen Phosphate (this is acid, about pH 4)
Put in a beaker and gradually add to this 1 M Di-Sodium Hydrogen Phosphate (pH about 9), until the mix reaches a pH of 7 at 20 °C
Autoclave
Acknowledgments
We thank Holger Puchta for kindly providing the recombination substrate lines and Gudrun Böhmdorfer for helpful discussions on DNA damaging agents. Work was supported by Grants GEN-AU GZ 200.140-VI/1/2006 from the Austrian Federal Ministry of Science and Research and FWF P18986-B17 from the Austrian Science Fund. This protocol was used in Rosa et al. (2013). The GUS histochemical assay was originally established by Pecinka et al. (2009).
References
- March-Díaz, R. and Reyes, J. C. (2009). The beauty of being a variant: H2A. Z and the SWR1 complex in plants. Mol Plant 2(4): 565-577.
- Masson, J. and Paszkowski, J. (1992). The culture response of Arabidopsis thaliana protoplasts is determined by the growth conditions of donor plants. Plant J 2(5): 829-833.
- Orel, N., Kyryk, A. and Puchta, H. (2003). Different pathways of homologous recombination are used for the repair of double-strand breaks within tandemly arranged sequences in the plant genome. Plant J 35(5): 604-612.
- Rosa, M., Von Harder, M., Cigliano, R. A., Schlogelhofer, P. and Mittelsten Scheid, O. (2013). The Arabidopsis SWR1 chromatin-remodeling complex is important for DNA repair, somatic recombination, and meiosis. Plant Cell 25(6): 1990-2001.
- Pecinka, A., Rosa, M., Schikora, A., Berlinger, M., Hirt, H., Luschnig, C. and Mittelsten Scheid, O. (2009). Transgenerational stress memory is not a general response in Arabidopsis. PLoS One 4(4): e5202.
- Schuermann, D., Molinier, J., Fritsch, O. and Hohn, B. (2005). The dual nature of homologous recombination in plants. Trends Genet 21(3): 172-181.
- Waterworth, W. M., Drury, G. E., Bray, C. M. and West, C. E. (2011). Repairing breaks in the plant genome: the importance of keeping it together. New Phytol 192(4): 805-822.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Rosa, M. and Scheid, O. M. (2014). Measuring Homologous Recombination Frequency in Arabidopsis Seedlings. Bio-protocol 4(7): e1094. DOI: 10.21769/BioProtoc.1094.
Category
Plant Science > Plant physiology > Tissue analysis
Plant Science > Plant molecular biology > DNA > DNA recombination
Molecular Biology > DNA > DNA recombination
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











