- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In situ Chemotaxis Assay in Caenorhabditis elegans (for the Study of Circadian Rhythms)
Published: Vol 4, Iss 3, Feb 5, 2014 DOI: 10.21769/BioProtoc.1040 Views: 14337
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Measuring Spatiotemporal Dynamics of Odor Gradient for Small Animals by Gas Chromatography
Akiko Yamazoe-Umemoto [...] Koutarou D. Kimura
Apr 5, 2018 7426 Views

Time-off-pick Assay to Measure Caenorhabditis elegans Motility
Alyssa C. Walker [...] Daniel M. Czyż
Jun 20, 2022 3214 Views
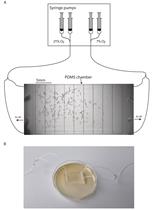
Aerotaxis Assay in Caenorhabditis elegans to Study Behavioral Plasticity
Qiaochu Li [...] Karl Emanuel Busch
Aug 20, 2022 2273 Views
Abstract
Olfaction is a well-studied sensory mechanism in Caenorhabditis elegans (C. elegans). The nematodes respond to a wide range of chemicals by either attraction, repulsion or a mixture thereof (Bargmann et al., 1993). We have used olfaction to characterize behavioural and molecular circadian rhythms in C. elegans. The circadian clock is a biological oscillator that provides an endogenous temporal structure that approximately matches the 24-hour periodicity in the environment (due to the rotational movement of the Earth). Circadian rhythms are present in most organisms from cyanobacteria to humans and they typically regulate sensory functions among many other processes. Olfaction is under circadian control in many animals (Granados-Fuentes et al., 2006; Granados-Fuentes et al., 2011; Tanoue et al., 2008; Krishnan et al., 1999). This protocol was designed to allow the assessment of olfaction for a population of worms within a short time interval, in the same plate where the worms grew (to avoid washing steps that may disturb the rhythms), and in the presence of food.
Keywords: CircandianMaterials and Reagents
- Egg preparation
100 eggs are used for each petri dish. Assays are performed in triplicate. - LB broth
- Cholesterol
- Ethanol
- CaCl2
- MgSO4
- KPO4
- 3.7% (v/v) 1-octanol (Merck KGaA, catalog number: 820931 ) in Ethanol
This concentration was established after a titration assay to find the lowest concentration needed for a fast response (within the minutes range) (Olmedo et al., 2012) - NGM plates (see Recipes)
Note: For one experiment, always use plates that were poured from the same batch and were allowed to dry for the same amount of time. - Escherichia coli (E. coli) OP50 culture at 100 g wet bacterial weight/liter (see Recipes)
Equipment
- Template for labeling plate (Figure 1)
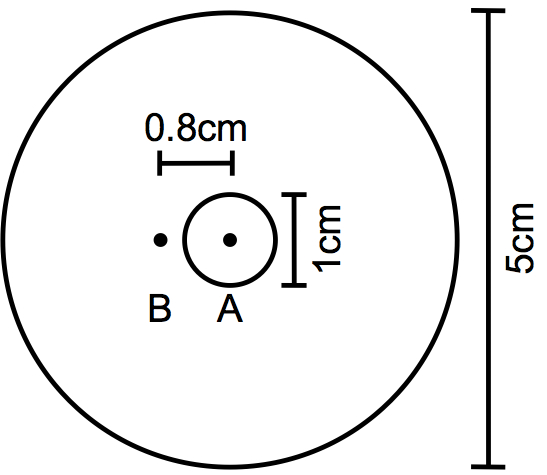
Figure 1. Template for plate labeling. On a piece of paper or plastic, draw a 5 cm-diameter circle in which to place the petri dish and a concentric 1 cm-diameter circle to mark the spot for the E. coli drop. Mark the center of this circle (A) with a dot. Draw another dot (B) at 0.8 cm from the center of the circle. - 5 cm Petri plates
- Stereomicroscope equipped for picture acquisition (we use a Stereo Discovery V8 from Zeiss at a 10x magnification.)
- Centrifuge
Procedure
- Add 15 µl of the 100 g wet weight/L E. coli OP50 culture to the center of the NGM plate (inside A).
Note: You may need to optimize the size of the drop for your stereomicroscope so that the pictures taken show the entire drop. A 15 µl drop of E. coli will form a circle of approximately 1 cm in diameter. - Let the drop dry and allow the bacteria to grow O/N at room temperature.
- Add about 100 eggs on the E. coli drop. Grow the worms under the desired conditions until the developmental stage for which you want to score olfaction.
Note: For circadian experiments, we grow the worms under temperature cycles of 16 h at 13 °C and 8 h at 16 °C. This 24-hour cycle is repeated 5 times until the worms become young adults. Then, the chemotaxis assay is performed every hour for 24 h to look for circadian rhythms in the response to 1-octanol. - Chemotaxis assay: Place the NGM plate over the template, and center the E. coli drop with the circle of the template.
- Open the lid and add 1 µl of 1-octanol at dot B. Take care that the 1-octanol does not touch with the E. coli drop. If it does, discard the plate (since the worms that are in direct contact with the 1-octanol/ethanol solution will die).
- Close the lid of the plate and incubate for 15 min at the desired temperature.
Note: For circadian experiments the assay can be performed over 24 h in constant conditions or under the same cycle used to entrain the worms. In both cases, the plates are incubated for 15 min at the same temperature at which they were before adding the 1-octanol. - Take a picture of the final position of the worms. Always keep the dot of 1-octanol at the same position of the picture (Figure 2).
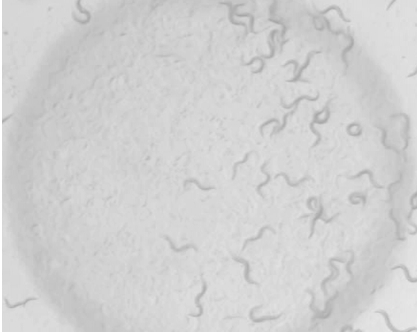
Figure 2. Final picture of the E. coli drop of a plate showing high response to 1-octanol - On the picture, draw a vertical line dividing the 1 cm circle in two equal parts (Figure 3). Count the number of worms in the proximal (p) and distal (d) sides of the circle. Worms that intersect the middle line are not counted.
- Calculate the chemotaxis index (CI=d-p/p+d).
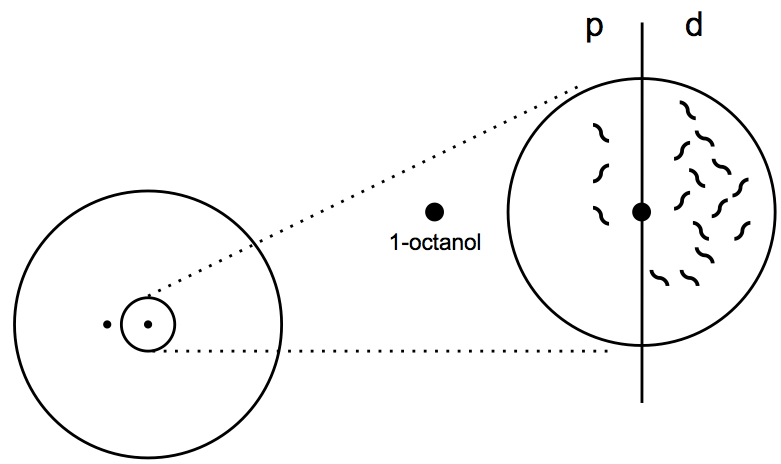
Figure 3. Calculation of chemotaxis index. The chemotaxis index is calculated by counting the worms in the proximal (p) and distal (p) sides of the circle and applying the formula CI = d - p/p + d.
Recipes
- NGM plates
Prepare NGM media:
Note: It is very important to prepare all the plates for the experiment in the same batch and allow them to dry for the same time. Variability in the level of humidity of the plates will introduce variability in the assay.- NGM media: Mix 3 g NaCl, 17 g agar, and 2.5 g peptone. Add 975 ml dH2O and stir to mix. Pour half content into a second flask and autoclave. Cool media in a 55 °C water bath or incubator for at least 30 min.
- Using sterile conditions, add 1 ml 1 M CaCl2, 1 ml 5 mg/ml cholesterol in ethanol, 1 ml 1 M MgSO4 and 25 ml 1 M KPO4 buffer and mix well.
Dispense 10 ml of NGM solution into 5 cm Petri plates using a pump. Leave plates at room temperature for 2 days before use to allow excess moisture to evaporate.
- NGM media: Mix 3 g NaCl, 17 g agar, and 2.5 g peptone. Add 975 ml dH2O and stir to mix. Pour half content into a second flask and autoclave. Cool media in a 55 °C water bath or incubator for at least 30 min.
- 100 g wet weight/L E. coli OP50 culture
Pick one single colony of OP50 bacteria and inoculate in 200 ml of LB broth. Grow O/N at 37 °C. Centrifuge the culture 10 min at 5,000 x g. Remove the supernatant, calculate the wet weight of bacteria and adjust concentration using LB broth.
Acknowledgments
This method is based on work that was originally published in Olmedo et al. (2013). This work was supported by the Dutch Science Foundation (NWO; VICI Program).
References
- Bargmann, C. I., Hartwieg, E. and Horvitz, H. R. (1993). Odorant-selective genes and neurons mediate olfaction in C. elegans. Cell 74(3): 515-527.
- Granados-Fuentes, D., Tseng, A. and Herzog, E. D. (2006). A circadian clock in the olfactory bulb controls olfactory responsivity. J Neurosci 26(47): 12219-12225.
- Granados-Fuentes, D., Ben-Josef, G., Perry, G., Wilson, D. A., Sullivan-Wilson, A. and Herzog, E. D. (2011). Daily rhythms in olfactory discrimination depend on clock genes but not the suprachiasmatic nucleus. J Biol Rhythms 26(6): 552-560.
- Krishnan, B., Dryer, S. E. and Hardin, P. E. (1999). Circadian rhythms in olfactory responses of Drosophila melanogaster. Nature 400(6742): 375-378.
- Olmedo, M., O'Neill, J. S., Edgar, R. S., Valekunja, U. K., Reddy, A. B. and Merrow, M. (2012). Circadian regulation of olfaction and an evolutionarily conserved, nontranscriptional marker in Caenorhabditis elegans. Proc Natl Acad Sci U S A 109(50): 20479-20484.
- Tanoue, S., Krishnan, P., Chatterjee, A. and Hardin, P. E. (2008). G protein-coupled receptor kinase 2 is required for rhythmic olfactory responses in Drosophila. Curr Biol 18(11): 787-794.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Merrow, M. and Olmedo, M. (2014). In situ Chemotaxis Assay in Caenorhabditis elegans (for the Study of Circadian Rhythms). Bio-protocol 4(3): e1040. DOI: 10.21769/BioProtoc.1040.
Category
Neuroscience > Behavioral neuroscience > Chemotaxis
Developmental Biology > Morphogenesis > Motility
Neuroscience > Sensory and motor systems
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











