- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
ADCC Assay Protocol
Published: Vol 4, Iss 2, Jan 20, 2014 DOI: 10.21769/BioProtoc.1029 Views: 44720
Reviewed by: Lin FangFanglian HeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Detachment Procedure of Bacteria from Atmospheric Particles for Flow-cytometry Counting
Carolina M. Araya [...] Isabel Reche
Jun 20, 2019 5629 Views
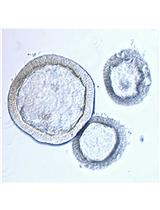
Intestinal Enteroid Culture for Human Astroviruses
Irene A. Owusu [...] Abimbola O. Kolawole
Jul 20, 2020 4796 Views
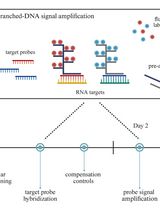
Flow Cytometric Quantification of HIV-1-Infected Cells Expressing Either Abortive or Elongated HIV-1 Transcripts Using Flow-FISH
Shirley Man [...] Neeltje A. Kootstra
Jul 20, 2025 2177 Views
Abstract
Antibody-dependent cell-mediated cytotoxicity (ADCC) bridges innate and adaptive immunity, and it involves both humoral and cellular immune responses. ADCC has been found to be a main route of immune protection against viral infections and cancers in vivo. Here we developed a flow cytometry based protocol for ADCC assay using human peripheral blood mononuclear cells (PBMCs) as effector cells. Using this protocol, we determined the ADCC activity of convalescent plasma IgGs from six H1N1-infected human subjects in China, and identified two dominant ADCC epitopes, designated E1 [amino acid (AA) 92-117] and E2 (AA 124-159), on haemagglutinin of pandemic H1N1 influenza virus by epitope mapping of the convalescent plasma IgGs with different levels of ADCC activity. Our study may aid in designing immunogens that can elicit antibodies with high ADCC activity. Vaccine immunogens designed to include the structural determinants of potent broadly neutralizing antibodies and ADCC epitopes may confer a comprehensive immune protection against viral infections.
Materials and Reagents
- Raji cells (ATCC)
- A/California/04/2009 (H1N1) influenza virus (Reference Laboratory)
- Turkey red blood cells (from animal)
- Tissue culture medium with serum (complete medium) RPMI 1640 (Life Technologies) containing 10% heat-inactivated fetal calf serum (FCS) and 2% L-glutamine
- Serum, albumin, or other system-compatible protein (FBS) (Life Technologies, catalog number: 10099141 )
- PKH67 cell labelling dye (Sigma-Aldrich, catalog number: MIDI67 )
- 7-AAD Stain (Life Technologies, catalog number: A1310 )
- PBS (Calcium Magnesium free) (Life Technologies, catalog number: 10010-023 )
- RPMI 1640 media (Life Technologies, catalog number: 12633-020 )
- Pen/Strep (Life Technologies, catalog number: 10378016 )
- Triton X-100 (Sigma-Aldrich, catalog number: T8787 )
Equipment
- Round bottomed 96-well plate
- Temperature controlled centrifuge
- Polypropylene conical bottom centrifuge tubes (4-15 ml)
- Bio-safety Cabinet
- Haemocytometer or cell counter
- Slides and coverslips
- Instrument (s) for fluorescence analysis (Flow cytometer)
- 37 °C, 5% CO2 incubator
Procedure
- Preperation of Target Cells
- Infect Raji cells at a MOI 2, 48 h before assay with A/California/04/2009 (H1N1) influenza virus in BSL-3 facility at a multiplicity to give about 80-95% infected cells. Detail procedure in Note 4.
- Take a sample of target cells 48 h after infection in order to assess the incidence of infected cells through its ability to produce hem-agglutination with Turkey red blood cells and binding with polyclonal antibodies. Detail procedure in Note 5.
- 2 x 106 cells washed twice with PBS.
- PKH67 cell labelling performed as per the protocol for General Cell Membrane Labelling by Sigma-Aldrich.
- Infect Raji cells at a MOI 2, 48 h before assay with A/California/04/2009 (H1N1) influenza virus in BSL-3 facility at a multiplicity to give about 80-95% infected cells. Detail procedure in Note 4.
- Preparation of Effector Cells (Isolation of PBMCs)
- Isolate PBMCs from Blood samples of healthy volunteers, according to method given by Panda and Ravindran (2013).
- Wash twice isolated PBMCs with PBS.
- Isolate PBMCs from Blood samples of healthy volunteers, according to method given by Panda and Ravindran (2013).
- Antibody-dependent cell-mediated cytotoxicity assays (ADCC)
Antibody-dependent cell toxicity was determined using the flow-cytometry based method employing the principle of live and dead cell discrimination (Srivastava et al., 2013).- Dispense 5.0 x 104 labelled target cells in 50 μl RPMI 1640 media in each well as per the layout given bellow in round-bottomed 96 well plate in duplicate.
- Add 50 μl of antibodies (total IgGs from plasma of infected patients 1μg/ml and/or 5 μg/ml resulting in 0.5 μg/ml and 2.5 μg/ml) to the wells as per layout.
- Incubate for 15 min at 37 °C in CO2 incubator.
- Add unlabelled normal PBMC effector cells in 100 μl of RPMI 1640/0.5% Pen/Strep at a concentration of 2.5 x 107 cells/ml to each well as per the layout.
- Incubate cells for 2 h at 37 °C in CO2 incubator.
- After 2 h add 1 µl of the fluorescent dead cell dye 7-amino-actinomycin-D (7-AAD).
- Incubate at 4 °C in dark for 20 min.
- Analyse cells on FACS machine. A total of 5,000 target cells acquire.
- Determine percentage cell death by software analysis of four identifiable cell populations, live effector cells (no dye), dead effector cells (7-AAD only), live target cells (PKH-67 only) and dead target cells (PKH-67 and 7-AAD).
1 2 3 4 5 6 7 Unstained Target Cells Unstained Effector Cells PKH67 Stained Target Cells 7AAD Stained Target Cells PKH67 Stained Target Cells + media + 7AAD Stain PKH67 Stained Target Cells + 5 µl 1% Triton X-100 + 7AAD Stain PKH67 Stained Target Cells + Antibody + 7AAD Stain
- Dispense 5.0 x 104 labelled target cells in 50 μl RPMI 1640 media in each well as per the layout given bellow in round-bottomed 96 well plate in duplicate.
- FACS analysis step
- Identify total (Live and dead) target cell zone (FSC Vs SSC) (5,000) (Well no 1).
- Identify effector cell zone (FSC Vs SSC) (5,000) (Well no 2).
- Acquire PKH67 positive cells (5,000) (Well no 3).
- Acquire 7AAD positive cells (5,000) (Well no 4).
- Acquire all wells 5, 6 and 7 (5,000 PKH67 positive cells).
- Locate double positive population (PKH67 and 7AAD) by gating PKH67 cells and then gating 7AAD positive cells.
- Identify total (Live and dead) target cell zone (FSC Vs SSC) (5,000) (Well no 1).
Notes
- Assay controls will be used to define cell populations included target cells without antibody (background cell death) and target cells with 5 µl 1% Triton X-100 (maximum fluorescence).
- Data presented as % increase in ADCC = (% cell death in presence of IgG - % cell death in absence of IgG)/(% Cell death in maximum lysis - % cell death in absence of IgG) x100.
- Experiments should be repeated for each IgG using pooled PBMCs from three healthy volunteers.
- Infection of Raji cells (briefly)
- 10 ml of 2 x 105 cells/ml wash 3 times with 5 ml of RPMI1640 containing 2 μg/ml of TPCK-trypsin.
- Remove media and inoculate A/California/04/2009 (H1N1) influenza virus at a MOI2.
- Allow inoculum to adsorb for 60 min at 37 °C.
- Gently wash with 6 ml of media (RPMI1640) containing 2 μg/ml of TPCK trypsin without serum.
- Add 5 ml of media (RPMI1640) containing 2 μg/ml of TPCK trypsin with 2% calf serum to T-25 flasks.
- 10 ml of 2 x 105 cells/ml wash 3 times with 5 ml of RPMI1640 containing 2 μg/ml of TPCK-trypsin.
- Hem-agglutination Assay (HA)
- Prepare culture supernatant, serially dilute virus in PBS.
- Take 50 μl of each dilution and place in pre-labeled U bottom 96-well plate.
- Add 50 μl of a 0.5% turkey Red Blood Cells suspension to each tube and mix by agitation.
- Allow tubes to sit undisturbed at room temperature for 30 min.
- After 30 min, record results. A negative tube is seen as a perfectly outlined round "button" of cells settled at the bottom of the tube. Intermediately positive results are seen with irregular clumps of cells at the bottom of the tube. Positive results are seen with a uniform film that lacks a distinct shape covering the bottom of the tube.
- Prepare culture supernatant, serially dilute virus in PBS.
Acknowledgments
We wish to thank Kwok-Yung Yuen and Kelvin KW To for providing the patient plasmas, Hong-Lin Chen and Zhiwei Chen for influenza virus H1N1 strains and the recombinant plasmid containing the full-length H1N1 HA gene, Dimiter S Dimitrov and Zhongyu Zhu for pYD7 yeast plasmid, and Martial Jaume for Raji cell line. We thank Kwok-Yung Yuen, Kelvin KW To, Linqi Zhang, Qi Zhao, Li Liu and Jia Guo for helpful discussions, and Yanyu Zhang and Jingjing Li for technical assistance. This work was supported by China 12th 5-year Mega project (# 2012ZX10001006) and Small Project Funding (# 201109176176 and # 201007176258) from the University of Hong Kong to M-Y. Z.
References
- Panda, S. and Ravindran, B. (2013). Isolation of Human PBMCs. Bio-protocol 3(3): e323.
- Srivastava, V., Yang, Z., Hung, I. F., Xu, J., Zheng, B. and Zhang, M. Y. (2013). Identification of dominant antibody-dependent cell-mediated cytotoxicity epitopes on the hemagglutinin antigen of pandemic H1N1 influenza virus. J Virol 87(10): 5831-5840.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Srivastava, V., Yang, Z., Hung, I. F. N., Xu, J., Zheng, B. and Zhang, M. (2014). ADCC Assay Protocol. Bio-protocol 4(2): e1029. DOI: 10.21769/BioProtoc.1029.
Category
Immunology > Immune cell function > Cytotoxicity
Cell Biology > Cell-based analysis > Flow cytometry
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









