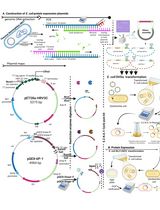
Bacteria are commonly known to secret proteins in large amounts into the surrounding environment in high concentrations via various pathways. These proteins can be involved in numerous processes like cell-cell communication, exopolymer formation but also metabolic active enzymes are secreted that are interesting for industrial production of proteins. One of the most regularly used organisms for industrial protein production is the Gram-positive bacterium Bacillus subtilis (B. subtilis). Here we describe a protocol that can be used to quantitatively and qualitatively analyze secreted proteins from B. subtilis.