- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Single-cell Gene Expression Profiling of Mouse Stem Cells With Fluidigm BiomarkTM Dynamic Array
Published: Vol 3, Iss 9, May 5, 2013 DOI: 10.21769/BioProtoc.692 Views: 19159

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
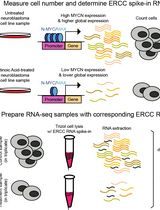
Accurate Measurement of Cell Number–Normalized Differential Gene Expression in Cells Treated With Retinoic Acid
Nina Weichert-Leahey [...] Brian J. Abraham
Nov 5, 2024 1811 Views
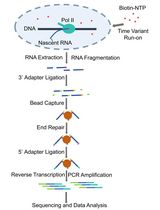
Fast TV-PRO-seq: Accelerated and Streamlined Protocol for Timing RNA Polymerase Pausing
Jie Zhang [...] Shaohui Zhang
Jul 20, 2025 2419 Views
Efficient Fluorescent Labeling of Human Trophoblast Stem Cells via a CRISPR/Cas9-Mediated Knock-In Approach in a Safe Harbor Locus
Hengshan Zhang [...] Danny J. Schust
Jan 5, 2026 293 Views
Abstract
This protocol describes how to use Fluidigm BiomarkTM 96.96 dynamic arrays for high-throughput expression profiling from single mouse stem cells, assaying up to 96 independent samples with up to 96 quantitative PCR (qPCR) probes (equivalent to 9,216 reactions) in a single experiment. This Dynamic Array contains a network of microfluidic channels, chambers and valves that automatically assemble all these individual PCR reactions. Single-cell profiling can provide definitive evidence of stem cell heterogeneity. Modifications are most likely needed if users intend to use BiomarkTM 48.48 Dynamic array or experimenter-designed primers in conjunction with DNA-binding dyes such as EvaGreen (Biotium 31000).
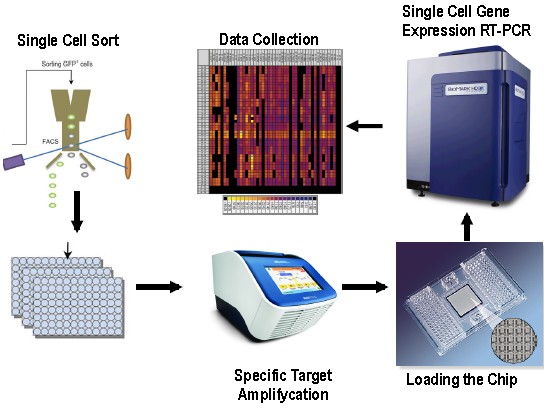
Figure 1. Workflow for Single-Cell Gene Expression Profiling in Mouse Stem Cells using Fluidigm BiomarkTM Dynamic Array. The main basic steps of this protocol are: Single cell sorting on 96 well plates, specific target amplifycation, chip loading, single cell gene expression RT-PCR and data collection. This Fluidigm Dynamic Array integrated fluidic circuit chip (IFCs) contains a network of microfluidic channels, chambers and valves that automatically assemble individual PCR reactions. More specific information about this particular chip can be obtained from http://www.fluidigm.com/single-cell-gene-expression.html.
Materials and Reagents
- Mouse ES cells (E14 Cell Line) (ATCC®, catalog number: SCRC-1040TM )
- Low-EDTA (0.1 mM EDTA) TE buffer (Teknova, catalog number: T0227 )
- CellsDirectTM One-Step qRT-PCR kit (Life Technologies, InvitrogenTM, catalog number: 11753-100 )
- Taqman Gene Expression Assays of the genes of interest for your study (Applied Biosystems)
Note: TaqMan® Assays are probe and primer sets based on 5’ nuclease chemistry using TaqMan® MGB (minor groove binder) probes.Search your Taqman gene expression assays here: https://bioinfo.appliedbiosystems.com/genome-database/gene-expression.html- Select the type of experiment you are conducting. Click In gene expression
- Select the type of assay you want. click in all gene expression assays
- Select what specie you want to target. Click on mouse since our samples are Mouse ES cells.
- Enter the target information. Add the gene symbol or entrez gene symbol or entrez gene ID of the gene you want to analyze its expression.
- 96.96 Dynamic Array Chip for Gene Expression (Fluidigm, catalog number: BMK-M-96.96 )
- Gene expression 96.96 Dynamic Array sample & loading reagent kit (Fluidigm, catalog number: 85000802 ), which contains:
- TaqMan® universal PCR master mix (Applied Biosystems, catalog number: 4304437 )
- Dulbecco's Phosphate-Buffered Saline (DPBS) (Life Technologies, catalog number: 14190-136 )
- TryPLE express solution (Life Technologies, catalog number: 12605-010 )
- 4',6-diamidino-2-phenylindole (DAPI) (Life Technologies, catalog number: D1306 )
- Propidium iodide (PI) (Life Technologies, catalog number: P-3566 )
- 2x Assay Loading Reagent (Fluidigm, catalog number: 85000736)
- 2x TaqMan®Universal PCR Master Mix (Applied Biosystems, catalog number: 4304437)
- 20x GE Sample Loading Reagent (Fluidigm, catalog number: 85000746 )
Equipment
- 96-well semi-skirted plate, 0.2 ml wells, straight raised sides (USA scientific, catalog number: 1402-9200 )
- Optical adhesive film (Applied Biosystems, catalog number: 4311971 )
- PCR-8-Tube Strips & Strip Camps, natural (USA Scientific, catalog number: 1402-2500 )
- TipOne 0.1–10 μl extended-length filter tips (USA Scientific, catalog number: 1120-3810 )
- TipOne 101–1,000 μl extended-length filter tips (USA Scientific, catalog number: 1122-1830 )
- TipOne 1–200 μl graduated filter tips (USA Scientific, catalog number: 1120-8810 )
- Fluorescence-activated cell sorter (FACS Aria II; BD Biosciences, catalog number: 643756 )
- Multichannel pipettes (10-100 μl Eppendorf Research®, catalog number: 38-3122000043 ; 0.5-10 μl catalog number: 38-3122000019 )
- Thermal cycler (VeritiTM 96-well Applied Biosystems, catalog number: PN 4375786 )
- The Integrated Fluidic Circuit IFC-Controllers
- The MX model primes and loads 48.48 chips
- The HX model primes and loads the 96.96 chips.
- Centrifuge (Eppendorf, catalog number: 5810 )
Procedure
- Reverse transcription-specific target amplification (RT-STA) master mix preparation.
- Thaw the CellsDirect 2x Reaction Mix from the CellsDirectTM One-Step qRT-PCR kit. All steps should be performed on ice, and reagents should be chilled and/or thawed immediately prior to use.
- Transfer 65 μl of the 2x Reaction Mix to each tube of the PCR-8-Tube Strip.
- Add 5 μl of 2x Reaction Mix from the PCR-8-Tube Strip to each well of the 96 well-plate with the multichannel pipette.
Since the amount of RNA from a single cell is few pictograms, we don’t perform a traditional RNA isolation protocol. Instead, the sorted cells are directly introduced into the 2x Reaction Mix. To ensure optimal sample preservation, this master mix contains CellDirect 2x reaction Mix (CellsDirect One-Step qRT-PCR kit) and SUPERase-In. This enzyme protects the RNA extracted from a lysed cell from any RNAases that could be present in the RT-STA master mix.
- Cell sorting
- Detach the mouse ES cells with TryPLE Express solution. Gently aspirate away cell media culture and wash the cells with 5-6 ml of fresh 1x DPBS to remove serum that could inhibit TryPLE Express solution activity.
- Aspirate away the 1x DPBS.
- Add 1 ml of pre-warmed TryPLE Express solution directly to the cell culture plate, and incubate the plate at 37 °C, 5% CO2 for 3-5 min.
- Take the cell culture plate out, gently tap or shake it, and then add 10 ml of fresh cell culture medium with 10% (v/v) FBS to stop trypsin activity. Mix cells with a pipette and transfer 1 x 106 cells into a 15 ml conical tube. Centrifuge at 300 x g, 4 °C for 5 min.
- Aspirate the cell supernatant and resuspend 1 x 106 cells in 1 ml of 1x DPBS supplemented with 5% (v/v) FBS.
- Keep 1 x 106 cells on ice before sorting or label the cells with the suitable surface markers according to the cell population you are interested to study.
- Add 10 μl of DAPI (> 200 ng/ml) or 10 μl of PI (5 μg/ml) per 1 x 106 cells to the single-cell suspension that contains 1 x 106 cells suspended in 1 ml of 1x DPBS supplemented with 5% (v/v) FBS just before cell sorting to distinguish the live cell population.
- Perform cell sorting in FACS Aria II and data analysis with FACS Diva software.
- When sorting cells gate for singlet’s in your sorting parameters.
- Sort single cells directly into the wells of a 96 well-plate containing a 5 μl mixture of CellsDirect 2x reaction mix (component CellsDirect One-Step qRT PCR Kits, Invitrogen). We estimate that the final volume of the reaction after cells are sorted into the wells will be 6 μl.
Note: Sort the cells into the same plate that is used for the RT-STA PCR. Always keep the well-plate on ice. - Centrifuge at 300 x g, 4 °C for 5 min the 96 well-plate to let the cells go into the CellsDirect 2x reaction mix.
- Seal the plate with the optical adhesive film.
- Store the plate at -80 °C. This overnight freezing step allows a better lysis efficiency.
- Preparation of the Specific Target Amplification (RT- STA) master mix.
- Make an excel file template for the 96 well-plate and label each cell with the specific 20x Taqman assay probe.
- Thaw the 20x Taqman assays and deposit them in the 96 well-plate according to the location establish in the excel file template for each 20x Taqman assay probe.
- Thaw the plate with the sorted cells in the CellsDirect 2x reaction mix on ice.
- Prepare the assay mix in a 1.5 ml eppendorf tube. Pool all Taqman real-time assays that will be analyzed in the single-cell real-time PCR that you previously deposited on a 96 well-plate in step b. Final concentrations of the Taqman assays used in the mix will be 0.2x. Use TE buffer to dilute the Taqman assays. This is the 0.2x assay mix.
Component Volume to add per reaction (μl) Final concentration 20x Taqman gene expression assay1 1.5 0.2x 20x Taqman gene expression assay2 1.5 0.2x … … … TE buffer Up to 150 μl TOTAL 150 μl - Prepare the sample RT-STA Master Mix by combining the following reagents:
Note: It is important to create a bulk solution for all the samples being tested to overcome pipetting limitations.Component Volume (μl) 0.2x Assay Mix 2.5 SuperScriptTM RT/Platinum® Taq Mix 0.2 TE Buffer 1.3 TOTAL 4 - Add 4 μl of the RT-STA master mix to each well that already contain the sorted cell immersed into the CellsDirect 2x reaction mix.
- Seal the plate with the optical adhesive film. Leave the plate on ice for 5 min meantime you program the thermal cycler.
Before you run the Reverse transcription and specific target amplification verify that each well from the 96 well plate has the components and the volumes according to the table below. Your final volume has to be 9 μl.Component Volume (μl) CellsDirect 2x Reaction Mix (This reagent is already in the plate with the sorted cell) 5.0 0.2x Assay Mix 2.5 SuperScriptTM RT/Platinum® Taq Mix 0.2 TE Buffer 1.3 TOTAL 9
- Make an excel file template for the 96 well-plate and label each cell with the specific 20x Taqman assay probe.
- Reverse transcription and specific target amplification.
- Place the 96 well plate with the RT-STA master mix in a thermal cycler.
50 °C for 15 min (Step to reverse transcribe the RNA to cDNA) 95 °C for 2 min (Step to inactivate the RT enzyme and activate the Taq) 22 Cycles of 95 °C for 15 sec and 60 °C for 4 min (STA Specific Target Amplification) Hold at 4 °C - Run the sample PCR according to these parameters:
Note: We screened different number of cycles and we found that for single cell analysis 22 cycles gave us better results in the Chip resolution. - Dilute the resulting cDNA product 1:5 with TE buffer.
Any dilution between 1:2 and 1:5 is possible.
- Place the 96 well plate with the RT-STA master mix in a thermal cycler.
- Preparing 10x Assays
In a DNA-free hood, prepare a 96 well-plate with 5μl aliquots of 10x assays using the volumes in the table below (scale up appropriately for multiple runs). Add the 20x TaqMan assays, one per well, to the 96 well-plate in the same position as in the excel file template.
Seal the plate with the optical adhesive film.Component Volume per Inlet (μl) 20x Taqman® gene expression assay (Applied Biosystems) 2.5 2x Assay Loading Reagent 2.5 Total volume 5.0 Final concentration (at 10x) Primers: 9 μM: Probe: 2 μM
Important: For unused sample inlets, add 3.3 μl of sample mix and 2.7 μl of DNA-free water per inlet, for unused assay inlets, add 3.0 μl assay loading reagent and 3.0 μl of water.
- Preparing sample mixture
Combine the components in the table below to make the Sample Mixture (scale up appropriately for multiple runs).
Seal the plate with the optical adhesive film.Component Volume per Inlet (μl) TaqMan® Universal PCR Master Mix (2x) 2.5 20x GE sample loading reagent 0.25 cDNA (already diluted 1/5) 2.25 Total volume 5.0
- Priming the 96.96 Dynamic ArrayTM in the Integrated Fluidic Circuit (IFC).
- Take a BioMark 96 x 96 Dynamic Array from its bag. Avoid touching the center of the chip.
Control line fluid on the chip or in the inlets makes the chip unusable. This chip needs to be use within 24 h after opening the package. - Take the 96.96 syringes with 150 μl of control line fluid and inject the 150 μl of control line fluid into each accumulator on the chip by introducing the syringe through the accumulators and pushing down the black O-ring. Use one syringe per accumulator. Avoid bending the pipette tip from the syringe.
- Place the chip into the Integrated Fluidic Circuit (IFC) controller HX, then select the Prime (136x) script program to prime the control line fluid into the chip. This program takes (~20 min).
- Take a BioMark 96 x 96 Dynamic Array from its bag. Avoid touching the center of the chip.
- Chip Loading
- Load the chip within 60 minutes of priming to reduce the loss of pressure within the chip.
- Vortex thoroughly and at 300 x g, 4 °C for 5 min all assay and sample solutions from the 96 well-plates before pipetting into the chip inlets.
Important: It is crucial to avoid forming any bubbles inside the chip inlets when loading the 10x assay mixes (prepared in step 5) on the chip inlets located on the left side and the 10x sample mixes (prepared on step 6) on the chip inlets located on the right side of the chip. To avoid bubbles, press the pipette only until the first stop. If bubbles appeared, pop them with a fresh sterile tip. Take care of cross-contamination or loss of reaction volume. - Place the chip back into the IFC Controller HX, select the Load mix (163x) program to load the samples and assays into the chip. Selecting OK starts the loading. This program takes (~1 h and 30 min).
- While the Load Mix (136x) program is running select the “BioMark data collection Fluidigm” program from the computer.
- Select “double-click to warm up the lamp” from the program. The lamp will be ready in 20 min and the lamp indicator on the software should be in green.
- When the Load Mix (136x) program has finished, remove loaded chip from the IFC Controller HX.
- Load the chip within 60 minutes of priming to reduce the loss of pressure within the chip.
- Using the data collection software.
- Double-click the data collection software icon on the desktop to launch the software.
- Click start a New Run to open the machine door.
- Check the status bar to verify that the camera temperature and the lamp are ready in green.
- Remove the blue plastic film just before placing the chip on the tray. Place the chip with the barcode facing the outside of the machine. Follow the “A” orientation.
- Click Load.
- Verify that the reader identifies properly the chip barcode and recognizes the chip type.
- Click Next.
- Click run file.
- Select new.
- Browse for a file location where you want your data to be stored.
- Click Next.
- Select new.
- Amplification, Reference, Probes:
- Select application type: gene expression.
- Select passive reference: (ROX).
- Select assay: Single probe.
- Select probe types: FAM-MGB.
- Click Next.
- Select application type: gene expression.
- Click Browse to find thermal protocol file: GE 96 x 96 Standard v1.pcl
PCR profile included:95 °C for 10 min 40 cycles 95 °C for 15 sec and 60 °C for 60 sec - Confirm auto expose is selected.
- Click Next.
- Verify the chip run information.
- Click Start Run. This program takes (~2 h).
- Double-click the data collection software icon on the desktop to launch the software.
- Result analysis Single-cell gene expression profiling represented as CT values.
- When the run has been completed, open the program “Fluidigm Real-time PCR analysis”.
- Open the chipRun file. bml to be analyzed.
- Select “Analysis view”.
- With the view of the results, set the following parameters for your analysis: the quality threshold, the baseline correction and CT threshold method.
- Click “Analyze”.
- The results are shown as CT values. The CT values can be seen in the software as a heat- map view, image view or as a table. Export all files with the “.cvs” extension.
- The data can be visualized and fully analyzed using Fluidigm real-time PCR analysis or any program for gene expression arrays.
- When the run has been completed, open the program “Fluidigm Real-time PCR analysis”.
Acknowledgments
We thank all the authors of this paper MacArthur et al. (2012) that participated in the single cell generation profiling as well as the data analysis. We gratefully acknowledge funding support by NIH (GM078465) and NYSTEM (C024410) to IRL. This work was also supported by an EPSRC Doctoral Training Centre grant (EP/G03690X/1) and an EPSRC 2011/12 Institutional Sponsorship Award (EP/J501530/1).
References
- MacArthur, B. D., Sevilla, A., Lenz, M., Muller, F. J., Schuldt, B. M., Schuppert, A. A., Ridden, S. J., Stumpf, P. S., Fidalgo, M., Ma'ayan, A., Wang, J. and Lemischka, I. R. (2012). Nanog-dependent feedback loops regulate murine embryonic stem cell heterogeneity. Nat Cell Biol 14(11): 1139-1147.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Sevilla, A. (2013). Single-cell Gene Expression Profiling of Mouse Stem Cells With Fluidigm BiomarkTM Dynamic Array. Bio-protocol 3(9): e692. DOI: 10.21769/BioProtoc.692.
Category
Stem Cell > Embryonic stem cell > Cell-based analysis
Cell Biology > Single cell analysis > Microfluidics
Molecular Biology > RNA > Transcription
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









