- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
ImmunoPrecipitation of Nuclear Protein with Antibody Affinity Columns
Published: Vol 3, Iss 3, Feb 5, 2013 DOI: 10.21769/BioProtoc.319 Views: 34743

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Individual-nucleotide-resolution UV Cross-linking and Immunoprecipitation (iCLIP) of UPF1
David Zünd and Oliver Mühlemann
Apr 5, 2014 15076 Views
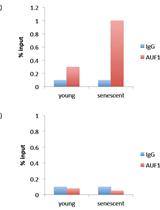
RNA-binding Protein Immunoprecipitation (RIP) to Examine AUF1 Binding to Senescence-Associated Secretory Phenotype (SASP) Factor mRNA
Elise Alspach and Sheila A. Stewart
May 20, 2015 15249 Views
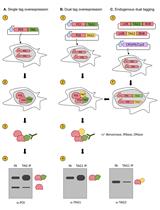
Assessing Self-interaction of Mammalian Nuclear Proteins by Co-immunoprecipitation
Claudia Cattoglio [...] Anders S. Hansen
Feb 20, 2020 9466 Views
Abstract
Co-Immunoprecipitation (Co-IP) is the method used to pull down protein partners of a protein of interest using an antibody that specifically binds to this specific protein in order to test protein-protein interaction. “Pulled down” proteins can be analyzed by western blot for suspected protein partner, or by Mass spectrometry for high throughput protein partner identification. The advantage of this technique is that endogenous protein partners can be identified from cell lines that naturally express these factors.
This protocol is optimized for hard-to-extract nuclear proteins, e.g., that stick to the nuclei inclusion bodies / nucleosome complexes such as TLX1 and TLX3 (Dadi et al., 2012). Most often, these factors are not soluble when using classical protein extraction methods. We used to add nucleases in order to increase solubilization of protein complexes trapped within inclusion bodies; though the efficacy varies depending on the given protein and therefore has to be empirically determined.
Materials and Reagents
- Proteine G agarose beads (Upstate, Millipore Corporation, catalog number: 16-266 )
- Dimethylpimelimidate (Sigma-Aldrich)
- Sodium borate
- Merthiolate
- Ethanolamine
- Merthiolate
- Protease inhibitor cocktail, EDTA free (F. Hoffmann-La Roche, catalog number: 04693159001 )
- Benzonase nuclease (Sigma-Aldrich, catalog number: E1014 )
- 2-mercapthoethanol
- Bromophenol Blue
- NP40
- Sucrose
- CaCl2
- MgOAc
- EDTA
- DTT
- PMSF
- DNase I (Sigma-Aldrich, catalog number: DN25 )
- Laemmli buffer (see Recipes)
- Sucrose buffer (see Recipes)
Equipment
- Wheel in a cold room
- Refrigerating centrifuge 1.5 ml tubes
Procedure
- Antibody affinity columns
- Incubate 5 to 10 mg of the antibody (Ab) per 1 ml of wet washed protein G agarose beads for 1 hour at room temperature with gentle rocking. The quantity of Ab and protein G agarose beads varies and depends on the affinity of the Ab (see note 1).
- Wash the beads twice with 10 volumes of 0.2 M sodium borate (pH 9.0) by centrifugation at 3,000 x g for 30 sec.
- Resuspend the beads in 10 volumes of 0.2 M sodium borate (pH 9.0) and save 1% of the total beads volume (aliquot “before”). Add dimethylpimelimidate (solid) to a 20 mM concentration. The pH is critical and should be above 8.3 after adding the dimethylpimelimidate for efficient coupling and can be checked with pH strips indicator for example.
- Mix for 30 min at room temperature on a rocker or shaker. Save 1% of the total beads volume (aliquot “after”).
- Centrifuge at 3,000 x g for 3 min and discard the supernatant.
- Stop the reaction by washing the beads once with equal volume of 0.2 M ethanolamine (pH 8.0). Centrifuge at 3,000 x g for 3 min and discard the supernatant.
- Repeat the 0.2 M ethanolamine wash one more time and incubate for 2 h at room temperature in with gentle mixing.
- Wash beads with equal volume of PBS. Centrifuge at 3,000 x g for 3 min and discard the supernatant. Repeat the wash with PBS one more time, then store the beads in PBS with 0.01% merthiolate in the desired volume. The beads are stable for over 1 year if stored at 4 °C.
- Check the efficiency of coupling by boiling samples of beads taken before and after coupling in Laemmli buffer. Run on 10% SDS PAGE and stain with Coomassie blue. Good coupling is indicated by heavy-chain bands (55 kDa) in the “before” but not in the “after” lanes. If there are small amounts of heavy chain, on the “after”, prewash the coupled beads with 100 mM glycine (pH 3.0) by centrifugation at 10,000 x g for 30 sec to remove any residual antidodies that are not covalently bound to the beads. Then wash beads with PBS then store the beads in PBS with 0.01% merthiolate in the desired volume.
- Incubate 5 to 10 mg of the antibody (Ab) per 1 ml of wet washed protein G agarose beads for 1 hour at room temperature with gentle rocking. The quantity of Ab and protein G agarose beads varies and depends on the affinity of the Ab (see note 1).
- Nuclear extract preparation
We used this protocol with ALL-SIL and DND41 cell lines derived from patient T cell lymphoblast.- Wash cells with cold 1x PBS then centrifuge for 6 min at 1,200 rpm at 4 °C. From now on, all the steps should be performed on ice.
- Resuspend the cell pellet in the chilled Sucrose buffer (5 μl/1 x 106 cells).
- Add vol/vol Sucrose buffer containing 0.5% NP40 (final concentration 0.25%).
- Mix by pipetting on ice. Save a small aliquot (1 to 5% -aliquot 1).
- Centrifuge 10 min at 1,100 x g at 4 °C. The pellet contains the nuclei and looks nacreous to white. The supernatant contains the cytoplasmic protein extract and can be saved if a cytoplasmic protein is of interest for Co-IP. Save a small aliquot (1 to 5% -aliquot 2).
- Wash the pellet with Sucrose buffer (without NP40) and centrifuge 10 min at 2,000 rpm at 4 °C.
- Depending on the nuclear proteins to be purified, you can lyse nuclei with
- Either the Nuclear Lysis buffer for soluble proteins (5 μl/ 1 x 106 cells);
- Or the Nuclei Lysis buffer for hard-to-extract proteins (5 μl/ 1 x 106 cells) and add DNase 5 U/μl and Benzonase 5 U/μl. Resuspension is hard since it’s very viscous with DNA.
- Either the Nuclear Lysis buffer for soluble proteins (5 μl/ 1 x 106 cells);
- Incubate on a wheel at 4 °C for 45 min to 1 h. Save a small aliquot (1 to 5% -aliquot 3).
- Centrifuge 3 min at 10,000 x g at 4 °C. The supernatant is the protein nuclei extract that will serve for the IP. Save a small aliquot from the supernatant (1 to 5% -aliquot 4; which is also the input of the IP experiment) and the pellet is the insoluble substance such as membrane debris. Resuspend the pellet in 5 μl/ 1 x 106 cells of 150 mM NaCl, 10 mM Tris. Save a small aliquot (1 to 5% -aliquot 5).
- Verify the efficiency of the lysis by analyzing by SDS-PAGE and western-blot the presence of your proteins of interest in the saved aliquots.
- Aliquot 1: total cells
- Aliquot 2: cytoplasm protein extract
- Aliquot 3: total nuclei extract
- Aliquot 4: nuclear protein extract
- Aliquot 5: insoluble nuclei extract
- Aliquot 1: total cells
- Wash cells with cold 1x PBS then centrifuge for 6 min at 1,200 rpm at 4 °C. From now on, all the steps should be performed on ice.
- Co-ImmunoPrecipitation
- Incubate the protein nuclei extract with the Antibody-bound beads. (Ab concentration needs to be determined experimentally.) Optimization is required for efficient IP of your protein of interest and the quantity of Ab may vary according to the quality and affinity of the Ab, the protein G coupling, the protein stability, etc.
- Incubate 2 h at 4 °C with gentle rocking.
- Wash 4 to 6 times the beads in 100 mM NaCl, 15 mM Tris, HCl pH 7.8, by mixing by pipetting and centrifuging 10,000 x g for 30 sec at 4 °C.
- Elute the bound proteins in Laemmli loading buffer and separate by SDS-PAGE and analyze by western blot.
- Incubate the protein nuclei extract with the Antibody-bound beads. (Ab concentration needs to be determined experimentally.) Optimization is required for efficient IP of your protein of interest and the quantity of Ab may vary according to the quality and affinity of the Ab, the protein G coupling, the protein stability, etc.
Recipes
- Laemmli buffer
20 % Glycerol
4% SDS
250 mM Tris (pH 6.8) (stacking buffer for upper gel of SDS PAGE)
1.4 M 2-mercapthoethanol, a pinch of bromophenol blue. - Sucrose buffer
0.32 M Sucrose
3 mM CaCl2
2 mM MgOAc
0.1 mM EDTA
10 mM DTT
0.5 mM PMSF - Nuclei Lysis buffer for hard to extract nuclear factors
50 mM Hepes (pH 7.8)
3 mM MgCl2
300 mM NaCl
1 mM DTT
0.1 mM PMSF
Protease inhibitor complete mini EDTA free tablets 1x - Nuclei Lysis buffer for soluble nuclear factors
50 mM Hepes (pH 7.8)
50 mM KCl
300 mM NaCl
0.1 mM EDTA
10 % Glycerol
1 mM DTT
0.1 mM PMSF
Protease inhibitor complete mini EDTA free tablets 1x
Notes
- We used Protein G agarose beads with mouse IgG1 antibodies. However, the affinity of the isotype antibody of interest needs to be verified accordingly to the protein A or G manufacturer’s recommendations.
- The quantity of the cells used for protein extract may vary according to the expression, the stability of the protein.
- The Benzonase and DNase treatment is used only in case of hard to extract proteins that stick to the nuclei inclusion bodies/ nucleosome. If the proteins of interest are soluble, the use of the Nuclei Lysis buffer for soluble factors is preferred.
Acknowledgments
Work in the PF laboratory is supported by institutional grants from 'Institut National de la Santé et de la Recherche Médicale' (Inserm) and 'Centre National de la Recherche Scientifique' (CNRS), and by dedicated grants from the Commission of the European Communities, the 'Agence Nationale de la Recherche' (ANR), the 'Institut National du Cancer' (INCa), the 'ITMO Cancer Alliance Nationale pour les Sciences de la Vie et de la Santé' (AVIESAN) and the 'Fondation Princesse Grace de la Principauté de Monaco'. S.D. was supported by fellowships from the ‘Ministère de l’Enseignement Supérieur et de la Recherche’, the ‘Fondation pour la Recherche Médicale’ (FRM), and the ‘Société Française d’Hématologie’ (SFH).
References
- Dadi, S., Le Noir, S., Payet-Bornet, D., Lhermitte, L., Zacarias-Cabeza, J., Bergeron, J., Villarese, P., Vachez, E., Dik, W. A., Millien, C., Radford, I., Verhoeyen, E., Cosset, F. L., Petit, A., Ifrah, N., Dombret, H., Hermine, O., Spicuglia, S., Langerak, A. W., Macintyre, E. A., Nadel, B., Ferrier, P. and Asnafi, V. (2012). TLX homeodomain oncogenes mediate T cell maturation arrest in T-ALL via interaction with ETS1 and suppression of TCRalpha gene expression. Cancer Cell 21(4): 563-576.
- Dignam, J. D., Lebovitz, R. M. and Roeder, R. G. (1983). Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res 11(5): 1475-1489.
- Gersten, D. M. and Marchalonis, J. J. (1978). A rapid, novel method for the solid-phase derivatization of IgG antibodies for immune-affinity chromatography. J Immunol Methods 24(3-4): 305-309.
- Schneider, C., Newman, R. A., Sutherland, D. R., Asser, U. and Greaves, M. F. (1982). A one-step purification of membrane proteins using a high efficiency immunomatrix. J Biol Chem 257(18): 10766-10769.
- Simanis, V. and Lane, D. P. (1985). An immunoaffinity purification procedure for SV40 large T antigen. Virology 144(1): 88-100.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Dadi, S., Payet-Bornet, D. and Ferrier, P. (2013). ImmunoPrecipitation of Nuclear Protein with Antibody Affinity Columns. Bio-protocol 3(3): e319. DOI: 10.21769/BioProtoc.319.
Category
Biochemistry > Protein > Immunodetection > Immunoprecipitation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link