- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Rapid IFM Dissection for Visualizing Fluorescently Tagged Sarcomeric Proteins
Published: Vol 7, Iss 22, Nov 20, 2017 DOI: 10.21769/BioProtoc.2606 Views: 11116
Reviewed by: Jyotiska ChaudhuriAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Measurement of Contractile Ring Tension Using Two-photon Laser Ablation during Drosophila Cellularization
Swati Sharma and Richa Rikhy
Mar 20, 2022 2497 Views
Live Imaging of Phagoptosis in ex vivo Drosophila Testis
Diana Kanaan [...] Hila Toledano
Mar 20, 2023 1829 Views
Protocol for Imaging the Same Class IV Neurons at Different Stages of Development
Sonal Shree and Jonathon Howard
Aug 20, 2024 1318 Views
Abstract
Sarcomeres, the smallest contractile unit of muscles, are arguably the most impressive actomyosin structure. Yet a complete understanding of sarcomere formation and maintenance is missing. The Drosophila indirect flight muscle (IFM) has proven to be a very valuable model to study sarcomeres. Here, we present a protocol for the rapid dissection of IFM and analysis of sarcomeres using fluorescently tagged proteins.
Keywords: DissectionBackground
The cytoskeletal structures that enable contractility of striated muscle fibers are hundreds of cables called myofibrils. Myofibrils in turn are an array of serially arranged sarcomeres, all contracting simultaneously. The sarcomere is a perfectly symmetrical structure that contains all the elements required for contraction. At the center of the sarcomere lies the M-line, where myosin thick filaments are anchored. Flanking the sarcomere are the Z-discs, where actin thin filaments are anchored.
Muscular dystrophies are inherited disorders that cause progressive skeletal muscle weakness (Schröder and Schoser, 2009). There is no cure for muscular dystrophy, likely due to an incomplete understanding of the molecular mechanisms that underlie muscular dystrophies (Olive et al., 2013). Drosophila melanogaster is an effective genetic model organism to study muscle biology owing to its short life span, economical maintenance, and abundant available resources (Hales et al., 2015; Wangler et al., 2015).
Flight in Drosophila is powered by the synchronized action of the indirect flight muscles (IFM), the biggest muscles in flies, which are further subdivided into dorsal longitudinal muscles (DLM) and dorsal ventral muscles (DVM). The IFM share many fundamental similarities with human skeletal muscle: contraction mechanism, developmental steps, overall ultrastructure, and protein components (Vigoreaux, 2001). For example, the myopathy-related proteins ZASP and Filamin-C have fly homologs that when mutated develop muscle phenotypes (Liao et al., 2016; Gonzalez-Morales et al., 2017). Despite the advantages of using the IFM for muscle research, IFM dissection can be challenging and time-consuming. Here we present a protocol that combines fast and easy IFM dissection with high-quality imaging of the IFM using fluorescent proteins. We also provide a strategy for analyzing mutant phenotypes and quantifying sarcomeres by semi-automatic detection of sarcomere components.
Materials and Reagents
- Surgical blade (FEATHER Safety Razor, catalog number: No. 23 )
- 1.5 ml microcentrifuge tube
- Pipette tips
- Conventional needles PrecisionGlide 23 G 1 in. (Fisher Scientific, catalog number: 14-826-6B)
Manufacturer: BD, catalog number: 305193 . - BD disposable syringes (BD, catalog number: 309628 )
- Cover Glass No. 1 ½ 22 x 30 mm (e.g., Corning, catalog number: 2850-22 )
- Microscope slides (e.g., Fisher Scientific, catalog number: 12-552-3 )
- Flies expressing sarcomere fluorescent markers (e.g., Zasp52-GFP, Table 1)
- Custom-made 3.5% agar-filled 60 x 15 mm Petri dish plate (BioShop, catalog number: AGR003 )
- Phalloidin-Tetramethylrhodamine B isothiocyanate (TRITC) (Sigma-Aldrich, catalog number: P1951 )
- Phalloidin-Fluorescein Isothiocyanate (FITC) (Sigma-Aldrich, catalog number: P5282 )
- Mounting media ProLong Gold Antifade Mountant (Thermo Fisher Scientific, InvitrogenTM, catalog number: P36934 )
- Magnesium chloride (MgCl2)
- Ethylene glycol-bis-tetraacetic acid (EGTA) (Sigma-Aldrich, catalog number: E3889 )
- Adenosine triphosphate (ATP) (BioShop, catalog number: ATP007 )
- 1,4-Dithiothreitol (DTT) (BioShop, catalog number: DTT001 )
- cOmpleteTM, EDTA-free Protease Inhibitor Cocktail (Roche Diagnostics, catalog number: 11873580001 )
- Glycerol (BioShop, catalog number: GLY001 )
- Triton X-100 (BioShop, catalog number: TRX777 )
- 8% paraformaldehyde aqueous solution glass vial (Electron Microscopy Sciences, catalog number: 157-8 )
- Sodium chloride (NaCl)
- Potassium chloride (KCl)
- Sodium phosphate dibasic (Na2HPO4)
- Potassium phosphate dibasic (K2HPO4)
- Relaxing solution (see Recipes)
- Relaxing-Glycerol solution (see Recipes)
- 8% paraformaldehyde (see Recipes)
- 10x phosphate buffered saline (PBS) (see Recipes)
- 1x PBS, 0.1% Triton X-100 (PBST) (see Recipes)
Equipment
- CO2 Flypad, standard size (8.1 x 11.6 cm) (Genesee Scientific, Flystuff, catalog number: 59-114 )
- Blade handle for surgical blade (FEATHER Safety Razor, catalog number: No. 4 )
- Glass Petri dish; 60 x 15 mm (VWR, catalog number: 89000-770)
Manufacturer: DWK Life Sciences, KIMBLE®, catalog number: 2306-10010 . - Stereo microscope (Leica Microsystems, model: Leica MS5 )
- Dumont #5 forceps (Fine Science Tools, catalog number: 11251-30 )
- Incubator set to 25 °C
- P2, P20, P100, and P1000 Micro Pipettes (e.g., Gilson, catalog numbers: F144801 , F123600 , F123615 and F123602 )
- Platform mixers (e.g., Speci-Mix Test Tube Rocker) (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: M71015Q )
- Medium-sized pointed brush
- Standard fly husbandry equipment
Software
- Fiji (https://fiji.sc/)
- R Statistics package (https://www.r-project.org/)
Procedure
- Prepare flies to dissect
- Anesthetize flies using standard CO2 pads. Use 5-10 days old flies.
- Transfer flies to a custom-made agar plate using a fine brush. Concentrate all flies in the center of the plate.
- Add 100 µl of Relaxing-Glycerol solution (see Recipes) to the flies using a previously cut pipette tip.
- Anesthetize flies using standard CO2 pads. Use 5-10 days old flies.
- Cut half-thoraces
The main goal of this step is to split the thorax into two symmetric halves (Figure 1 and Video 1). This step is relatively easy, however take note not to crush the muscles. If the blade is dull, replace it with a new blade.- Use two fine forceps to remove the abdomen. Leave only the thorax and the head.
- Place and align the thorax with the dorsal side facing your eyes.
- Using a scalpel blade knife and a fast slicing movement cut the thorax in two symmetrical halves. Cut the head out if still attached to the thorax.
- At this point the IFM should be visible under the dissection microscope.
- Take the half-thoraxes with forceps and place them in a 1.5 ml microcentrifuge tube.
- Add 100 µl of Relaxing-Glycerol buffer.
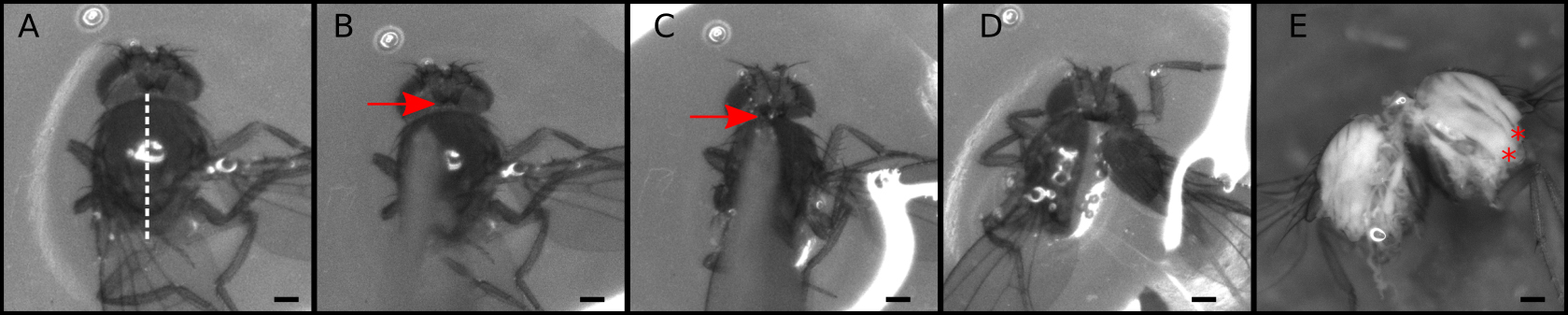
Figure 1. Dissection of the thorax into two halves. A-D. Snapshots from Video 1 showing critical dissection steps. A. Place the thorax with the dorsal side up; B. Gently touch the thorax to make sure the blade is positioned exactly at the middle and that the thorax will not move. C. Aim the tip of the blade at the head/thorax attachment site (arrowhead); D. With a sharp push cut the thorax; E. Representative thorax divided in two. Muscle fibers can be observed at this stage (red asterisks). Scale bars = 200 µm.Video 1. Dissection of the thorax into two halves. Dissection process recorded directly from the stereomicroscope. Snapshots of this video are presented in Figure 1.
- Use two fine forceps to remove the abdomen. Leave only the thorax and the head.
- Remove mitochondria
The mitochondria can be completely removed from the sample by an overnight incubation in Relaxing-Glycerol buffer at -20 °C without agitation (Figure 2). Removal of mitochondria makes imaging and staining easier. If mitochondria must be preserved, skip this section.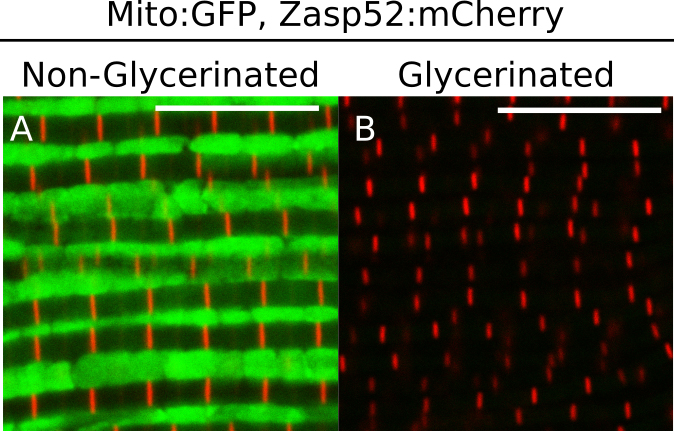
Figure 2. Effect of glycerol extraction on mitochondria. A and B. Representative IFM expressing a mitochondrial:GFP protein and a mCherry-tagged Zasp52. A. In non-glycerinated IFM mitochondria can be detected; B. Mitochondria cannot be detected in glycerinated IFM. Scale bars = 10 µm. - Paraformaldehyde fixation
- Add 150 µl of 8% paraformaldehyde (see Recipes) to the tube that contains the thoraces in relaxing buffer. The final paraformaldehyde concentration should be around 4%. A 4% formaldehyde fixation solution can also be used.
- Incubate at room temperature for 45 min with slow agitation (~50 rpm).
- Wash by replacing fixing solution with PBST (see Recipes). Incubate in PBST for 15 min under slow agitation (~50 rpm). Repeat this step three times. Thoraces quickly sink to the bottom of a microcentrifuge tube.
- Add 150 µl of 8% paraformaldehyde (see Recipes) to the tube that contains the thoraces in relaxing buffer. The final paraformaldehyde concentration should be around 4%. A 4% formaldehyde fixation solution can also be used.
- Dissection of muscle fibers
The main goal of this step is to separate the DLM fibers from the carcass without damaging them (Figure 3 and Video 2). Video 2 shows a detailed real-time dissection, see it before attempting to dissect the muscle fibers to get an idea of the overall process.- Transfer the thoraces to a glass Petri dish using a previously cut pipette tip.
- Add enough PBST to just cover the tissue (excess liquid will make dissection challenging).
- Connect the needles to the syringes to make two dissection needles.
- Use two dissection needles to dissect the muscle fibers from the carcass (for details see Figure 4 and Video 2).
- Separate the fibers as much as possible without tearing the fibers apart.
- Transfer the fibers using a previously cut pipette tip back to a 1.5 ml microcentrifuge tube with PBST.
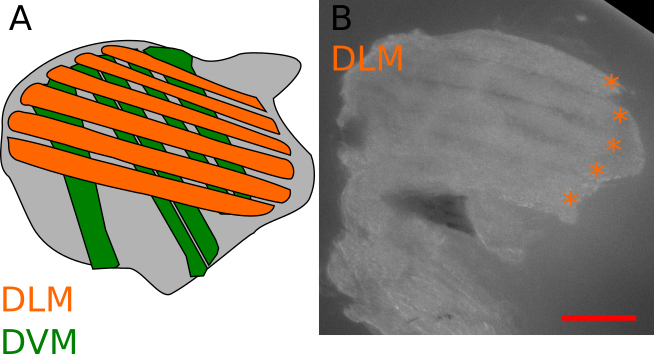
Figure 3. General description of the IFM. A. Schematic view of a longitudinal section of a thorax, showing the position of the IFM. Two types of muscles compose the IFM, the dorsal longitudinal muscles (DLM, orange) and the dorsal ventral muscles (DVM, green). The goal of this protocol is to dissect the DLM. B. Representative isolated DLM, orange asterisks indicate individual DLM fibers (see also Video 2). Scale bar = 200 µm.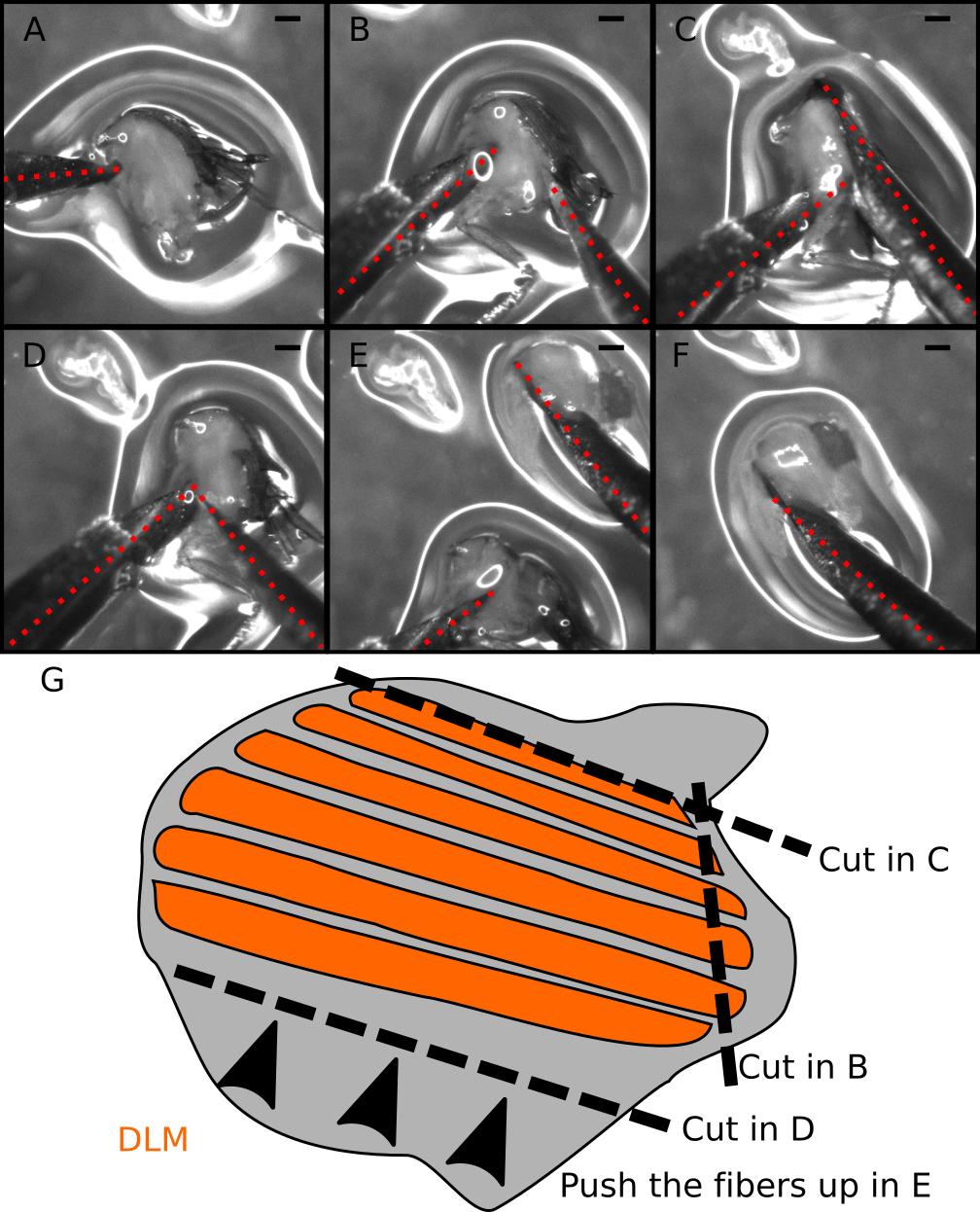
Figure 4. Step-by-step guide to dissect DLM fibers. A-F. Snapshots from Video 2. A. Locate and hold the half-thorax using the left needle. Place the needle just below the DLM and above the legs. B. Make a small incision at the DLM posterior attachment site. C. Make a long cut above the DLM to completely separate the muscle fibers from the dorsal cuticle. D. Move the right needle to the ventral DLM (near the left needle). E. With the right needle push the DLM fibers out of the thorax. F. Isolated DLM can be further separated into individual fibers. G. Cartoon illustrating the dissection process. Dotted red lines mark the dissection needles. Note the small amount of liquid (~100 µl) in the dissection plate to prevent floating of half-thoraces. Scale bars = 200 µm.Video 2. Dissection of the DLM fibers. Stereomicroscope-recorded video showing the steps required for the correct dissection of the DLM fibers. Snapshots of this video are also presented in Figure 4.
- Transfer the thoraces to a glass Petri dish using a previously cut pipette tip.
- Phalloidin staining and mounting
- Replace the PBST solution with detergent-free PBS. Be careful not to aspirate the fibers.
- Add FITC- or TRITC-phalloidin (1:1,250) in triton-free PBS.
- Incubate at room temperature for 1-2 h with slow agitation (~50 rpm).
- Wash three times with PBST under slow agitation (~50 rpm).
- Transfer the muscle fibers to a glass slide.
- Dry the samples using a Kimwipe.
- Add a drop of mounting media, compatible with fluorescent labelling. Cover with coverslip compatible with confocal microscopy.
- Store at 4 °C.
- Replace the PBST solution with detergent-free PBS. Be careful not to aspirate the fibers.
- Confocal microscopy
- Locate the muscle fibers under brightfield at 20x magnification.
- Change the objective to 63x/1.4 NA and adjust the focus using brightfield to protect the sample.
- Scan the muscle fiber using the confocal software, align the fiber and take a full resolution scanning image at 9x magnification 1,024 x 1,024 pixels.
- Take several 9x images at different locations within the muscle fiber to get an accurate representation of the phenotype, especially when dealing with tissue from a mutant.
- Locate the muscle fibers under brightfield at 20x magnification.
Data analysis
- Image analysis
- Images obtained from this protocol can be used to identify sarcomere phenotypes. Direct comparison of images from mutant muscles and control muscles should be enough to detect even subtle sarcomeric phenotypes. As healthy IFM are very regular structures (see Figure 5 for control examples), the mutant phenotypes can be scored by directly counting the number of normal sarcomeres in control and mutant muscles.
- When counting phenotypes exclude all sarcomeres that appear out of focus, do not have well-defined actin staining, or have more than one Z-disc per sarcomere. These are indications of artifacts often caused by overlapping myofibrils. Actin staining should always be used as counterstain when counting phenotypes.
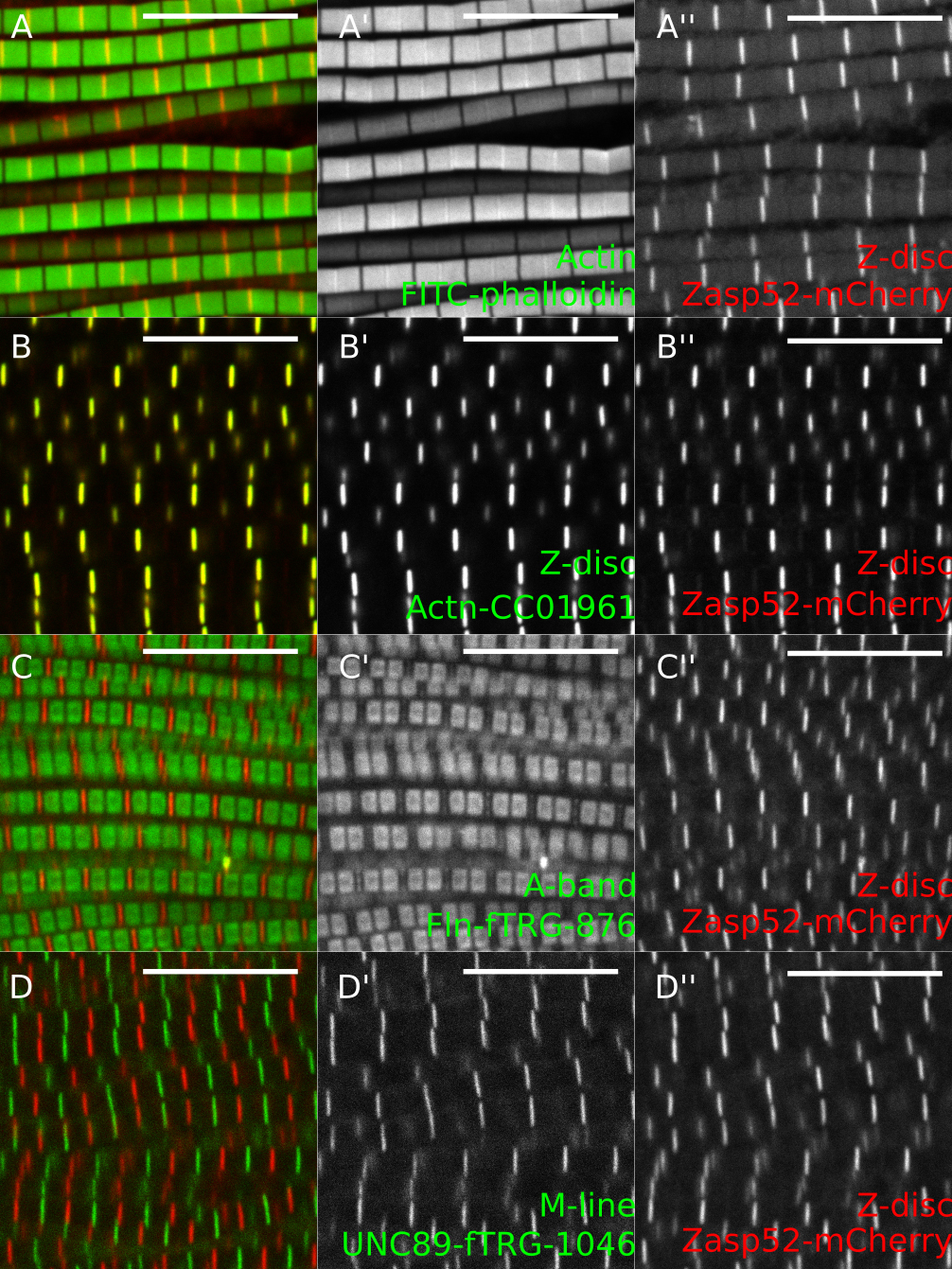

Figure 5. Representative examples of IFM confocal images. A. Zasp52-mCherry stained with phalloidin to visualize actin thin filaments. B. GFP-Actinin (GFP-Actn) and Zasp52-mCherry show a clear Z-disc colocalization. C. Flightin-GFP (Fln-GFP) marks the part of the A-band where the myosin heads are present. Fln does not colocalize with Zasp52-mCherry. D. Obscurin-GFP localizes to the M-line. Scale bars = 10 µm.
- Images obtained from this protocol can be used to identify sarcomere phenotypes. Direct comparison of images from mutant muscles and control muscles should be enough to detect even subtle sarcomeric phenotypes. As healthy IFM are very regular structures (see Figure 5 for control examples), the mutant phenotypes can be scored by directly counting the number of normal sarcomeres in control and mutant muscles.
- Semi-automatic Z-disc or M-line measurements
The fluorescent signal from Z-disc, M-line, or A-band proteins can be used to automatically detect individual particles (Figure 6). This approach is useful when very precise measurements of a large number of sarcomeres are required (e.g., estimating Z-disc size). Video 3 shows a screen-recorded version of this protocol.- Open the image with the Fiji package of ImageJ (Schindelin et al., 2012).
- Open the threshold menu, under the Image/Adjust menu. Select ‘Dark background’ and ‘Auto’.
- Under ‘Analyze’ menu, select ‘Set measurements’. Check the Area, Standard deviation, Bounding rectangle, Shape descriptors, Mean gray value, and Display label boxes.
- Under ‘Analyze’ menu, select ‘Analyze particles’
- Adjust the size value (0.3-1) and/or circularity to exclude undesirable objects (e.g., protein aggregates in mutant muscles).
- Select ‘outlines’ in the drop-down menu.
- The measurement and outline of the image should appear. If the outline contains undesirable objects, adjust the parameters in the ‘Analyze particles’ menu.
- Save the results.
- Analyze the results with any statistical software (e.g., R statistical programming language).
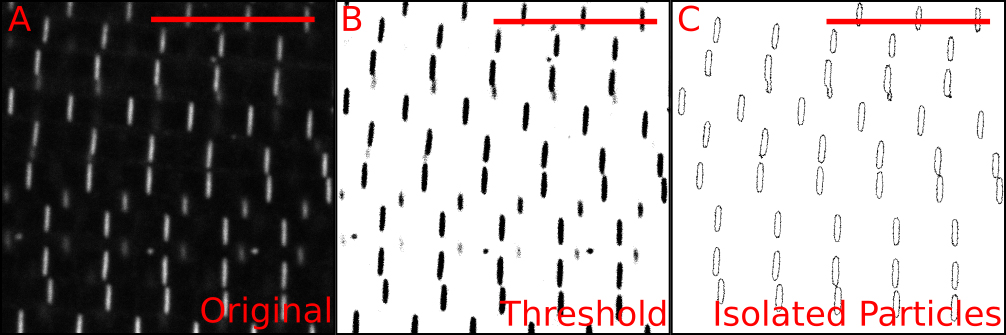
Figure 6. Automatic measurement of individual Z-discs. A. Confocal image of Zasp52-mCherry; B. Thresholded image showing in black the pixels to be included. The thresholding step uses signal intensity as an inclusion criterion. C. Result from running the ‘analyze particles’ step. This step uses size and shape as inclusion criteria. For additional details see Video 3. Scale bars = 10 µm.Video 3. Automatic detection and measurement of individual Z-discs. Screen-recorded video showing the procedure in Fiji to obtain individual Z-discs and measure some properties of the isolated discs. First, a thresholded image is made using the automatic threshold tool. Then, individual Z-disc are obtained using the ‘Analyze particles’ tool. The particle size and the circularity options may be adjusted to optimize the procedure.
- Open the image with the Fiji package of ImageJ (Schindelin et al., 2012).
Notes
- Antibody stainings
Antibody stainings work very well with IFM. However, they do not seem to work perfectly when this protocol is used. If antibodies should be used, the fixation and the dissection steps must be inverted. That is, after the overnight incubation instead of fixing the tissue first dissect the IFM fibers as shown in Video 2. Then wash the samples 3 times with Relaxing solution and then proceed to the fixation step. While this greatly enhances antibody stainings, unfixed DLM muscles are harder to dissect than pre-fixed fibers. - Useful fluorescent markers
This protocol works best when fluorescent markers are available. Many different stocks expressing GFP-tagged proteins are available from common stock centers. Table 1 shows a list of useful GFP stocks for IFM imaging. To complement the existing GFP stocks we made Zasp52MI02908-mCherry by integrating an mCherry cassette in frame with the Zasp52 sequence. This new Zasp52MI02908-mCherry allele recapitulates Zasp52 expression and localization (Figures 2, 5, and unpublished observations).
Table 1. Useful fluorescent markers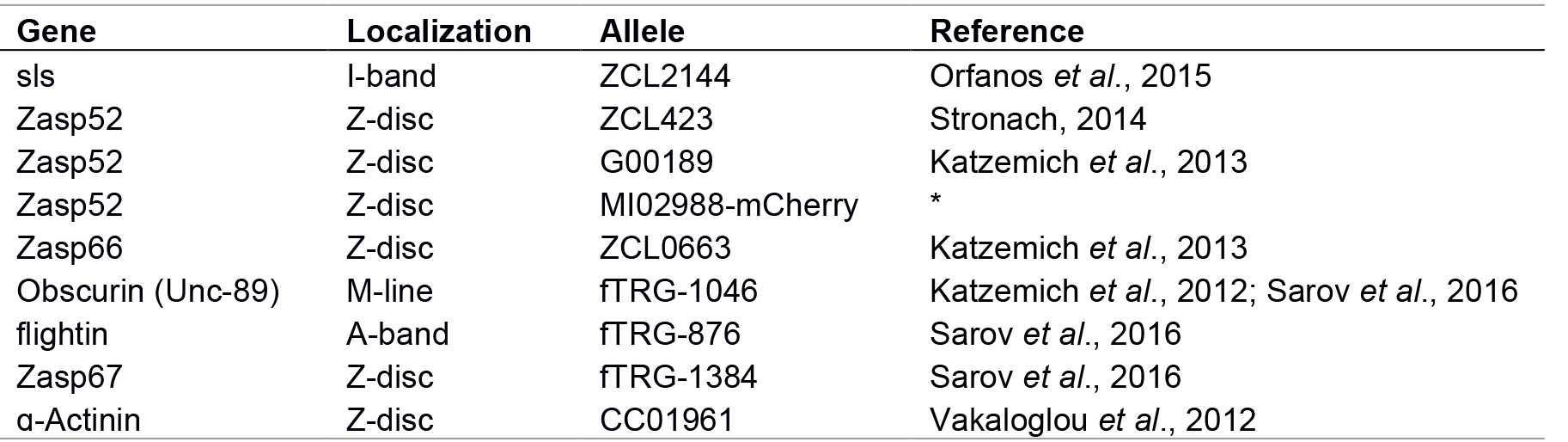
Note: * Not fully described, available upon request. - Confocal images
- All images must be acquired at the same resolution, magnification, and orientation. This allows direct comparison of images and saves time when arranging figures. We recommend a 20 x 20 µm square image, scanned at 1,024 x 1,024 pixel resolution and averaged 3 times.
- Scan the muscle fiber as close to the surface as possible. Phalloidin has limited infiltration into the IFM fibers. Thus, actin staining will be sharp at the surface but fuzzy in the interior. More importantly, IFM phenotypes may slightly vary between the surface and the interior of the muscle fibers.
- All images must be acquired at the same resolution, magnification, and orientation. This allows direct comparison of images and saves time when arranging figures. We recommend a 20 x 20 µm square image, scanned at 1,024 x 1,024 pixel resolution and averaged 3 times.
Recipes
- Relaxing solution
- Dissolve in 40 ml of ddH2O:
1,000 µl (final concentration is 20 mM) of sodium phosphate buffer stock solution 1 M pH 7.0
500 µl (final concentration is 5 mM) of 0.5 M MgCl2
2.5 ml (final concentration is 5 mM) of 0.1 M EGTA
2.5 ml (final concentration is 5 mM) of 0.1 M ATP stock solution
250 µl (final concentration is 5 mM) of 1 M DTT - Add 1 tablet of protease inhibitors (cOmpleteTM, EDTA-free Protease Inhibitor Cocktail, Roche)
- Adjust volume to 50 ml with ddH2O
- Store at -20 °C
- Dissolve in 40 ml of ddH2O:
- Relaxing-Glycerol solution
Note: Prepare Relaxing-Glycerol solution based on York Modified Glycerol solution (Peckham, 1990).- Mix the relaxing solution with glycerol 1:1 (v/v)
- Add 0.5% Triton X-100
- Store in 10 ml aliquots at -20 °C
- Mix the relaxing solution with glycerol 1:1 (v/v)
- 8% paraformaldehyde
- Open the 8% paraformaldehyde aqueous solution glass vial (Electron Microscopy Sciences)
- Divide the vial content into 400 µl aliquots and store them at -20 °C. Aliquots can then be stored for more than 6 months
- Open the 8% paraformaldehyde aqueous solution glass vial (Electron Microscopy Sciences)
- 10x phosphate buffered saline (PBS)
- Dissolve in 800 ml ddH2O
80 g of NaCl
2.0 g of KCl
14.4 g of Na2HPO4
2.4 g of K2HPO4 - Adjust pH to 7.4.
- Adjust volume to 1 L with ddH2O
- Dilute to 1x PBS with ddH2O
- Dissolve in 800 ml ddH2O
- 1x PBS, 0.1% Triton X-100 (PBST)
Add 500 µl of Triton X-100 to 500 ml of 1x PBS
Acknowledgments
This protocol is adapted from the work of others. The original IFM dissection technique was described and distributed by Belinda Bullard and John Sparrow. We thank Anja Katzemich for technical advice, and the CIAN imaging facility for help with confocal microscopy. This work was supported by operating grant MOP-142475 from the Canadian Institutes of Health Research. We have no conflict of interest to declare.
References
- Gonzalez-Morales, N., Holenka, T. K. and Schöck, F. (2017). Filamin actin-binding and titin-binding fulfill distinct functions in Z-disc cohesion. PLoS Genet 13(7): e1006880.
- Hales, K. G., Korey, C. A., Larracuente, A. M. and Roberts, D. M. (2015). Genetics on the Fly: A primer on the Drosophila model system. Genetics 201(3): 815-842.
- Katzemich, A., Kreisköther, N., Alexandrovich, A., Elliott, C., Schöck, F., Leonard, K., Sparrow, J. and Bullard, B. (2012). The function of the M-line protein obscurin in controlling the symmetry of the sarcomere in the flight muscle of Drosophila. J Cell Sci 125(Pt 14): 3367-3379.
- Katzemich, A., Liao, K. A., Czerniecki, S. and Schöck, F. (2013). Alp/Enigma family proteins cooperate in Z-disc formation and myofibril assembly. PLoS Genet 9(3): e1003342.
- Liao, K. A., Gonzalez-Morales, N. and Schöck, F. (2016). Zasp52, a core Z-disc protein in Drosophila indirect flight muscles, interacts with α-Actinin via an extended PDZ domain. PLoS Genet 12(10): e1006400.
- Olive, M., Kley, R. A. and Goldfarb, L. G. (2013). Myofibrillar myopathies: new developments. Curr Opin Neurol 26(5): 527-535.
- Orfanos, Z., Leonard, K., Elliott, C., Katzemich, A., Bullard, B. and Sparrow, J. (2015). Sallimus and the dynamics of sarcomere assembly in Drosophila flight muscles. J Mol Biol 427(12): 2151-2158.
- Peckham, M., Molloy, J. E., Sparrow, J. C. and White, D. C. (1990). Physiological properties of the dorsal longitudinal flight muscle and the tergal depressor of the trochanter muscle of Drosophila melanogaster. J Muscle Res Cell Motil 11(3): 203-15.
- Sarov, M., Barz, C., Jambor, H., Hein, M. Y., Schmied, C., Suchold, D., Stender, B., Janosch, S., K, J. V., Krishnan, R. T., Krishnamoorthy, A., Ferreira, I. R., Ejsmont, R. K., Finkl, K., Hasse, S., Kampfer, P., Plewka, N., Vinis, E., Schloissnig, S., Knust, E., Hartenstein, V., Mann, M., Ramaswami, M., VijayRaghavan, K., Tomancak, P. and Schnorrer, F. (2016). A genome-wide resource for the analysis of protein localisation in Drosophila. Elife 5: e12068.
- Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J. Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P. and Cardona, A. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7): 676-682.
- Schröder, R. and Schoser, B. (2009). Myofibrillar myopathies: a clinical and myopathological guide. Brain Pathol 19(3): 483-492.
- Stronach, B. (2014). Extensive nonmuscle expression and epithelial apicobasal localization of the Drosophila ALP/Enigma family protein, Zasp52. Gene Expr Patterns 15(2): 67-79.
- Vakaloglou, K. M., Chountala, M. and Zervas, C. G. (2012). Functional analysis of parvin and different modes of IPP-complex assembly at integrin sites during Drosophila development. J Cell Sci 125(Pt 13): 3221-3232.
- Vigoreaux, J. O. (2001). Genetics of the Drosophila flight muscle myofibril: a window into the biology of complex systems. Bioessays 23(11): 1047-1063.
- Wangler, M. F., Yamamoto, S. and Bellen, H. J. (2015). Fruit flies in biomedical research. Genetics 199(3): 639-653.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Xiao, Y. S., Schöck, F. and González-Morales, N. (2017). Rapid IFM Dissection for Visualizing Fluorescently Tagged Sarcomeric Proteins. Bio-protocol 7(22): e2606. DOI: 10.21769/BioProtoc.2606.
Category
Developmental Biology > Morphogenesis > Cell structure
Cell Biology > Tissue analysis > Tissue imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









