- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Freeze-fracture-etching Electron Microscopy for Facile Analysis of Yeast Ultrastructure
Published: Vol 7, Iss 18, Sep 20, 2017 DOI: 10.21769/BioProtoc.2556 Views: 14556
Reviewed by: Dennis NürnbergChong HeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Preparation of Candida albicans Biofilms for Transmission Electron Microscopy
Heather T. Taff and David R. Andes
Jul 20, 2013 11173 Views

Metal-tagging Transmission Electron Microscopy for Localisation of Tombusvirus Replication Compartments in Yeast
Isabel Fernández de Castro and Cristina Risco
Apr 20, 2018 7417 Views
Generating ER-TRG and CA-ER-TRG Knock-in Mice and Quantitative in vivo Imaging of ER-phagy
Mengyuan Zhang [...] Qiming Sun
Jan 5, 2026 190 Views
Abstract
We describe a streamlined method that enables the quick observation of yeast ultrastructure by electron microscopy (EM). Yeast cells are high-pressure frozen, freeze-fractured to cut across the cytoplasm, and freeze-etched to sublimate ice in the cytosol and the organelle lumen. The cellular structures delineated by these procedures are coated by a thin layer of platinum and carbon deposited by vacuum evaporation, and this platinum–carbon layer, or replica, is observed by transmission EM. The method differs from the deep-etching of pre-extracted samples in that intact live cells are processed without any chemical treatment. Lipid droplets made of unetchable lipid esters are observed most prominently, but other organelles–the nucleus, endoplasmic reticulum, Golgi, vacuoles, mitochondria–and their mutual relationships can be analyzed readily. It is of note that the entire procedure, from quick-freezing to EM observation, can be performed within a day.
Keywords: Freeze-fractureBackground
Budding yeast (Saccharomyces cerevisiae) is probably the most frequently used model organism, and application of a wide assortment of experimental techniques and the presence of sophisticated genome-wide databases have significantly facilitated research advancement (Botstein and Fink, 2011). The microscopic imaging of yeast, however, is not always carried out in a satisfactory manner due to its relatively small size and round shape. The presence of the cell wall is a problem for conventional electron microscopy (EM) in particular, because it hampers the penetration of reagents used in sample preparation. Freeze-substitution EM of quick-frozen yeast is currently considered the best method for ultrastructural observation (Giddings et al., 2001), but the method has drawbacks, namely, the membrane structures are not clearly visible, some cellular components may not be retained during the substitution process in organic solvents, and the procedure takes at least several days before observation is possible.
In a recent study to examine lipophagy in stationary-phase yeast, we used freeze-fracture-etching to analyze the yeast ultrastructure (Tsuji et al., 2017). Canonical freeze-fracture EM has been used to observe wide areas of membranes in two dimensions. By adding the etching process after freeze-fracturing, the cellular ultrastructure–the lipid droplets and interorganellar relationships in particular–was readily observed. Deep-etching combined with quick-freezing has been used successfully to analyze the cytoskeleton in pre-extracted samples (Heuser and Salpeter, 1979). The method described here is different in that intact live cells are quick-frozen and processed, thereby preserving the membrane organelles. Furthermore, with slight modification, the distribution of proteins and lipids on the nanoscale can also be analyzed (Cheng et al., 2014).
Materials and Reagents
- 200 mesh EM grid (Electron Microscopy Sciences, catalog number: M200-CR )
- 50 mesh EM grid (Electron Microscopy Sciences, catalog number: G50-Cu )
- Polyvinyl formvar (Nisshin EM, catalog number: 602 )
- Copper foil: 20 μm in thickness (Nilaco, catalog number: CU-113213 )
- 10 μl pipette tip (Thermo Fisher Scientific, catalog number: 3510 )
- Aluminum disc: 3 mm in diameter and 0.3 mm in thickness (Engineering Office M. Wohlwend, catalog number: 242 )
- Platinum-carbon (Pt/C) (Leica Microsystems, catalog number: 16771798 )
- Carbon (C) (Leica Microsystems, catalog number: 16771797 )
- Saccharomyces cerevisiae
- Acetone (CH3COCH3) (KANTO KAGAKU, catalog number: 01026-70 )
- Liquid nitrogen
- Household bleach (6% sodium hypochlorite)
Equipment
- 50-100 ml flask (IWAKI, catalog numbers: 4980FK50 , 4980FK100 )
- Razor blade (Feather, catalog number: FAS-10 )
- Forceps (EM Japan, catalog number: T7330 )
- Hole puncher (Carla Craft, catalog number: SD-15-3 )
- Incubator shaker (TAITEC, catalog number: BR-22FP )
- High-pressure freezing machine (Bal-Tec, model: HPM 010 )
- Freeze-fracture apparatus (Balzers, model: BAF400 )
- Electron beam gun (Balzers, model: EK552 )
- Crystal thickness monitor (Balzers, model: QSG 301 )
- Stereoscopic microscope (Leica Microsystems, model: Leica MZ6 )
- Transmission electron microscope (JEOL, model: JEM-1011 )
Procedure
The outline of the method is depicted in Figure 1.
Figure 1. The outline of the method. After ‘quick-freezing’ of yeasts, the cytoplasm is exposed by ‘fracturing’ of frozen cells with cooled knife. The ‘etching’ procedure induces sublimation of water in the cytosol and the organelle lumen. This makes lipid droplets stand out because lipid esters in the lipid droplet core do not sublimate. Vacuum evaporation of platinum and carbon onto the surface makes a ‘replica’ of the cellular ultrastructure.
- Quick freezing
- Prepare round-shaped copper discs of 3 mm in diameter using a hole puncher. Clean the discs by soaking them in acetone (Figures 2A and 2B).
- Concentrate yeast cells by centrifugation at 1,500 x g for 1 min.
- Place yeast cells on an EM grid (200 mesh) by dipping the grid into a pellet or spreading ~0.6 μl of pellet on the grid using a pipette tip (Figure 2B).
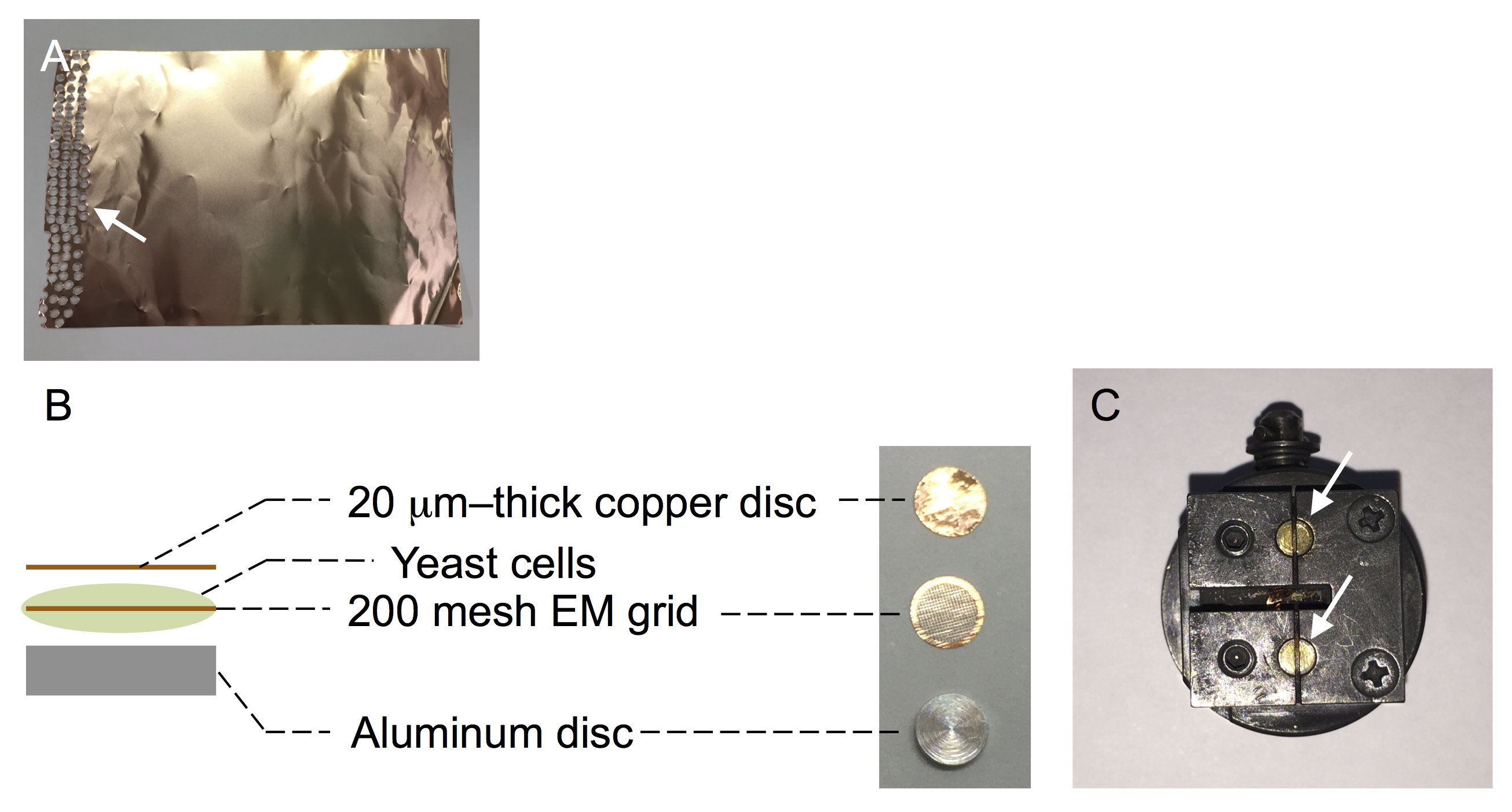

Figure 2. Tools used for quick-freezing and freeze-fracture-etching. A. Copper foil (20 μm thick). The left portion (arrow) has already been used to prepare round-shaped discs. B. The sample sandwich subjected to quick-freezing: a 200 mesh EM grid impregnated with yeast cells is placed between a 20-μm-thick copper disc and an aluminum disc. C. A specimen table of Balzers BAF400. The sample sandwiches are placed in the two round indentations (arrows). - Sandwich the EM grid impregnated with yeast cells between a 20-μm-thick copper foil and a flat aluminum disc (Figure 2B) (see Note 1).
- Quick-freeze yeast cells using an HPM 010 high-pressure freezing machine (or a similar device) according to the manufacturer’s instructions.
- Keep the frozen samples in liquid nitrogen until they are transferred to the cold specimen stage of a freeze-fracture apparatus.
- Freeze-fracturing and etching
- Mount the frozen sample onto a specimen table of the freeze-fracture apparatus (Figures 2C and 3). Here, the procedure for the Balzers BAF400 will be described, but it can easily be adapted to other devices.
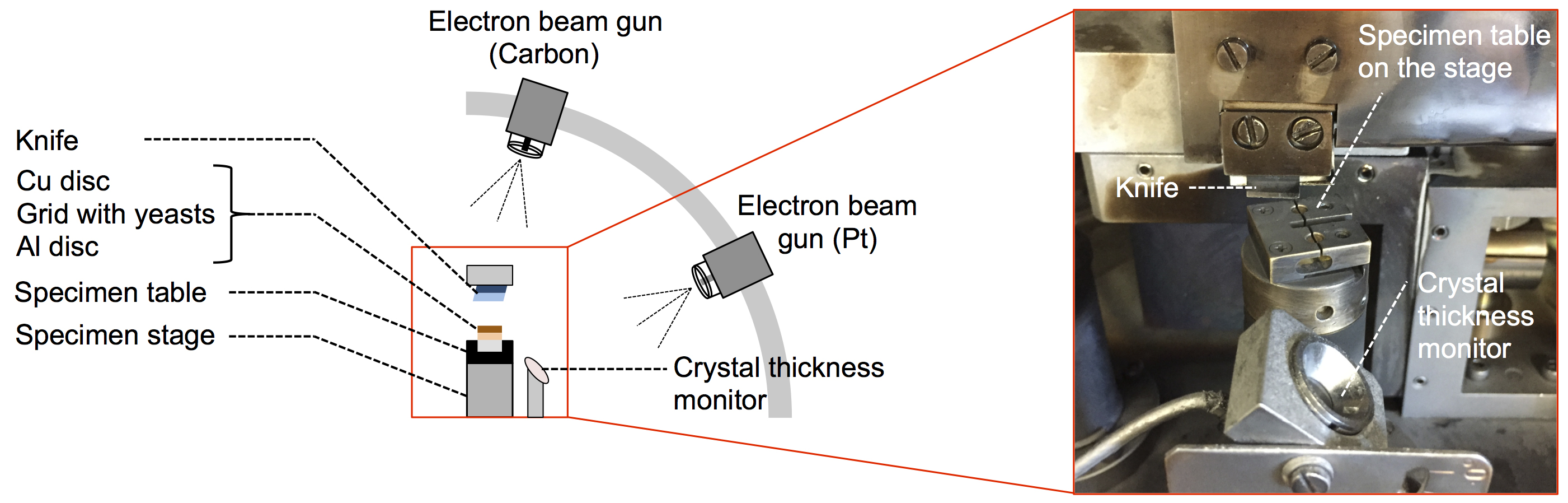
Figure 3. Freeze-fracture apparatus (BAF400) for freeze-fracture-etching. Electron beam guns for carbon and platinum deposition are located at 80° and 20° to the specimen surface, respectively. Specimen table is set on the temperature-controlled specimen stage. - Transfer the specimen table to the pre-cooled specimen stage of the freeze-fracture apparatus (Figure 3). The specimen stage needs to be cooled below -120 °C before this transfer.
- Keep the specimen temperature at -120 °C for 10 min, and then at -102 °C for 3 to 5 min by using the temperature controller of the specimen stage.
- Cool the knife to the lowest-possible temperature (e.g., near the temperature of liquid nitrogen).
- After the vacuum reaches below ~5 x 10-7 mbar, fracture the specimens at -102 °C by separating the copper disc from the aluminum disc using the pre-cooled knife (see Note 2).
- Keep the fractured specimens at -102 °C for another 2 min to induce sublimation of water from the fractured surface.
- Evaporate platinum-carbon (Pt/C) and carbon (C) onto the specimen using the electron beam gun. First, evaporate 2 to 4 nm of Pt/C at an angle of 20° to the specimen surface, then 10 nm of C at an angle of 80°. Rotate the specimen stage at maximum speed during evaporation. Control the thickness of the Pt/C and C deposition using a crystal thickness monitor (Figure 3).
- Break the vacuum, bring the specimen into the atmosphere, and transfer the freeze-fracture replicas to household bleach to digest biological materials for more than 2 h.
- Rinse the replicas with water.
- Mount the replicas on the formvar-coated EM grids under a stereoscopic microscope. The formvar-coated EM grids are prepared by a standard protocol (Slot and Geuze, 2007).
- Observe the replicas by transmission EM (see Note 3).
Data analysis
- Yeast cells are fractured in a random manner. Roughly speaking, approximately 70 to 80% of the yeast cells are freeze-fractured along the plasma membrane. For analysis of cytoplasmic organelles, the rest of the cells showing the cross-fractured cytoplasm are selected. Within the cytoplasm, lipid droplets are invariably cross-fractured and observed as round protruding structures whereas other organelles may be either freeze-fractured along the limiting membrane to show the convex or concave surface, or cross-fractured to reveal the internal structure (Figure 4).
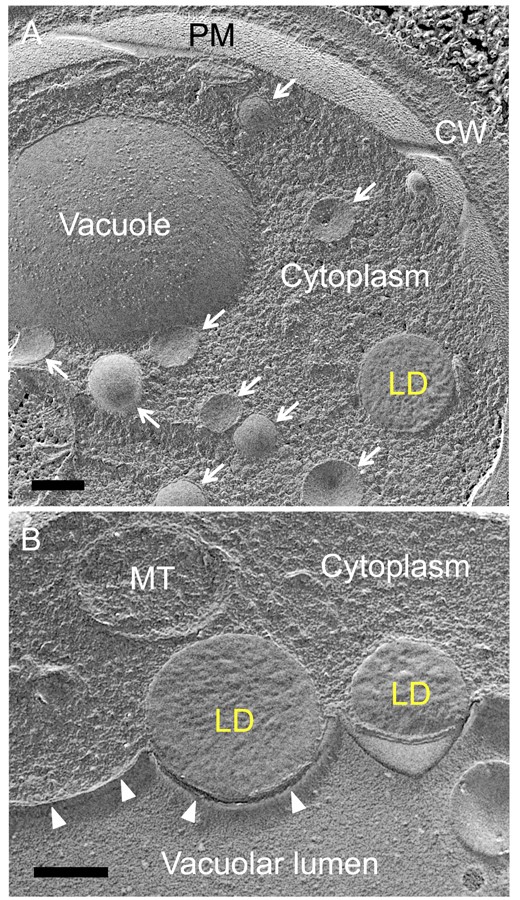
Figure 4. Freeze-fracture-etching EM of yeast in the stationary phase. A. The plasma membrane (PM) and the vacuolar membrane show the two-dimensional freeze-fractured plane whereas the lipid droplet (LD) is cross-fractured. A number of small vesicles (arrows) are also observed. CW: cell wall. B. Two LDs adhere to the vacuolar membrane (arrowheads). A cross-fractured mitochondrion (MT) is observed. Scale bars = 0.2 μm. - Damage to the cellular ultrastructure may occur when ice crystals form due to improper operation of the high-pressure freezing machine or the freeze-fracture apparatus, or by poor handling of frozen samples, for example, failure to keep samples in liquid nitrogen. Suboptimal replicas may form as a result of unsatisfactory vacuum evaporation. Samples exhibiting these problems can be readily identified by EM and should be excluded from further analyses.
- In most experiments, three replicates, prepared by independent quick-freezing and freeze-fracture-etching sessions, are analyzed for each yeast specimen.
Notes
- Liquid (or culture medium) must be kept to a minimum volume so that it will not spread to the outer side of the copper foil, which would result in improper freezing.
- The knife must be cooled to the coldest possible temperature before fracturing samples.
- EM grids must be completely dried before EM observation.
Acknowledgments
The condition of the etching procedure was modified from Heuser and Salpeter (1979). The authors thank Dr. John E. Heuser for his kind advice on technical details. This study was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology of the Government of Japan to TF (25111510, 15H02500, 15H05902) and TT (15K18954, 17K15544).
References
- Botstein, D. and Fink, G. R. (2011). Yeast: an experimental organism for 21st Century biology. Genetics 189(3): 695-704.
- Cheng, J., Fujita, A., Yamamoto, H., Tatematsu, T., Kakuta, S., Obara, K., Ohsumi, Y. and Fujimoto, T. (2014). Yeast and mammalian autophagosomes exhibit distinct phosphatidylinositol 3-phosphate asymmetries. Nat Commun 5: 3207.
- Giddings, T. H., Jr., O’Toole, E. T., Morphew, M., Mastronarde, D. N., McIntosh, J. R. and Winey, M. (2001). Using rapid freeze and freeze-substitution for the preparation of yeast cells for electron microscopy and three-dimensional analysis. Methods Cell Biol 67: 27-42.
- Heuser, J. E. and Salpeter, S. R. (1979). Organization of acetylcholine receptors in quick-frozen, deep-etched, and rotary-replicated Torpedo postsynaptic membrane. J Cell Biol 82(1): 150-173.
- Tsuji, T., Fujimoto, M., Tatematsu, T., Cheng, J., Orii, M., Takatori, S. and Fujimoto, T. (2017). Niemann-Pick type C proteins promote microautophagy by expanding raft-like membrane domains in the yeast vacuole. Elife 6: e25960.
- Slot, J. W. and Geuze, H. J. (2007). Cryosectioning and immunolabeling. Nat Prot 2: 2480-2491.
Article Information
Copyright
Tsuji and Fujimoto. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Tsuji, T. and Fujimoto, T. (2017). Freeze-fracture-etching Electron Microscopy for Facile Analysis of Yeast Ultrastructure. Bio-protocol 7(18): e2556. DOI: 10.21769/BioProtoc.2556.
- Tsuji, T., Fujimoto, M., Tatematsu, T., Cheng, J., Orii, M., Takatori, S. and Fujimoto, T. (2017). Niemann-Pick type C proteins promote microautophagy by expanding raft-like membrane domains in the yeast vacuole. Elife 6: e25960.
Category
Cell Biology > Cell structure > Cell organelle
Cell Biology > Cell imaging > Electron microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









