- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Intracellular cAMP Levels Using the Cyclic Nucleotide XP Enzymatic Immunoassay Kit in Bacteria
Published: Vol 6, Iss 8, Apr 20, 2016 DOI: 10.21769/BioProtoc.1792 Views: 13098
Reviewed by: Valentine V TrotterDaan C. SwartsAksiniya Asenova

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
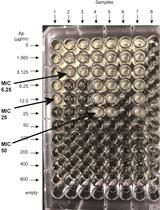

β-lactamase (Bla) Reporter-based System to Study Flagellar Type 3 Secretion in Salmonella
Fabienne F. V. Chevance and Kelly T. Hughes
Jun 20, 2023 1764 Views
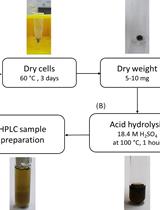
Determination of Poly(3-hydroxybutyrate) Content in Cyanobacterium Synechocystis sp. PCC 6803 Using Acid Hydrolysis Followed by High-performance Liquid Chromatography
Janine Kaewbai-ngam [...] Tanakarn Monshupanee
Aug 20, 2023 1808 Views
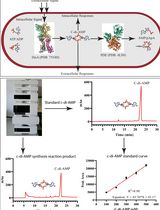
An HPLC-based Assay to Study the Activity of Cyclic Diadenosine Monophosphate (C-di-AMP) Synthase DisA from Mycobacterium smegmatis
Avisek Mahapa [...] Dipankar Chatterji
Dec 20, 2024 1779 Views
Abstract
Cyclic AMP (cAMP) is a ubiquitous secondary signaling molecule, commonly associated with many bacterial processes, including the regulation of virulence factors, such as biofilms, pellicles and motility (Wolfgang, 2003). The quantity of available cAMP is controlled by the interplay between the synthesis of adenosine triphosphate (ATP) to cAMP by adenylyl cyclases, and the degradation of cAMP by phosphodiesterase (McDonough et al., 2012). Adequate quantification of cAMP levels within a bacterial cell is an important step in identifying the impact that secondary signaling molecules play on the regulatory pathway within the cell. The principle of this method is to measure total cAMP levels within a bacterial cell, using crude bacterial whole cell lysate. The Cyclic AMP XPTM Assay kit used in this protocol was originally designed to be used for determining the level of cAMP in eukaryotic cells, however, the antibodies used in coating the wells will react with cAMP from any species and thus can be used for determining levels in bacterial cells. The measurement of cAMP in prokaryotic cells described here is a simple and cost effective method of producing quantifiable results.
Materials and Reagents
- 1 ml cuvettes (SARSTEDT AG & Co., catalog number: 67.742 )
- 15 ml tubes (Falcon Tube) (DKSH, catalog number: LP021015 )
- 50 ml tubes (Falcon Tubes) (DKSH, catalog number: LP21050 )
- 96-well microtitre tray (tissue culture plate) (DKSH, catalog number: LP031096 )
- Aluminum foil
- Bacterial cell culture
- Mueller Hinton (MH) broth (prepared as per manufacturer’s instructions) (Oxoid Australia, catalog number: CM0405 )
- MH agar media (25 ml/Petri dish) (prepared as per manufacturer’s instructions) (Oxoid Australia, catalog number: CM0337 )
- Cyclic AMP XPTM Assay kit (Cell Signaling Technology, catalog number: 4339 )
- Protein assay DCTM reagents A & B (Bio-Rad Laboratories, catalog number: 500-121 )
- Protein assay bovine serum albumin (BSA) standards (0, 0.125, 0.25, 0.5, 1, 2.5, 5 mg/ml of protein)
- Milli Q water
- Sodium Chloride (NaCl) (Chem Supply, catalog number: SA046-5 kg )
- Potassium Chloride (KCl) (Thermo Fisher Scientific, Ajax Finechem, catalog number: A383-500 g )
- Sodium Phosphate (Na2HPO4) (Merck Millipore Corporation, catalog numer: 7558-79-4 )
- Potassium Phosphate (KH2PO4) (Merck Millipore Corporation, catalog number: 7778-77-0 )
- Phosphate buffered saline (PBS) (see Recipes)
Equipment
- VIS spectrophotometer able to read OD600 nm [e.g., DU®640 (Beckman Coulter)]
- ELISA plate reader able to read absorption at 450 nm [e.g., Multiskan EX original (Adelab Scientific)]
- French pressure cell able to operate at 30,000 psi or cell disruptor [e.g., One Shot Head from the TS series bench top disruptor (Constant Systems)]
- Ice bucket
- Sterile inoculating loop
- Shaking incubator (37 °C, at 200 rpm)
- Centrifuge (able to hold 50 ml tubes, spin at 8,000 x g and hold a pre-cooled temperature of 4 °C) [e.g. Benchtop Centrifuge Hermle Z383K (DKSH)]
- Horizontal orbital shaker (at ~30 rpm) [e.g., OM5 (Ratek, Adelab Scientific)]
Software
- Software for use with the ELISA plate reader that can export raw OD600 nm values (e.g. Microsoft Excel)
- GraphPad prism (GraphPad Software Inc)
Procedure
- Grow an overnight culture (~16 h) of the required strain on an agar plate, including any antibiotic selection if required. Prepare MH broth media for culture and sterile PBS.
- Next Day: Under sterile conditions, using a sterile inoculating loop remove a single colony from the agar plate and inoculate 5 ml of MH broth in a 50 ml tube including any antibiotic selection if required. Grow culture overnight (~16 h) at 37 °C at 200 rpm.
- Next Day: Make a 1:100 dilution of the overnight culture into 15 ml of MH broth in a 50 ml tube. Grow culture at 37 °C shaking at 200 rpm until the culture reaches an OD600 = 0.7 (approximately 3-4 h), measure using a spectrophotometer.
- While the bacterial culture is growing, pre-cool the centrifuge to 4 °C, prepare ice bucket, pre-cool PBS on ice.
- Remove bacterial cells from incubator and put on ice, centrifuge sample for 20 min, 4 °C at 8,000 x g, decant supernatant and keep cell pellet on ice; sample can be frozen at -20 °C for up to 6 months.
- Resuspend the pellet in 3 ml of cooled PBS buffer and keep sample on ice.
- Set up cell disruptor with a one shot head (pre-cooled to 4 °C); set to 30,000 psi, and process sample, collect lysed cells into a new 15 ml tube on ice.
- Centrifuge sample at 8,000 x g for 20 min at 4 °C, remove the supernatant and store at -80 °C until required, up to 6 months.
- Remove 10 μl of sample to quantify total protein in sample using the Protein assay DCTM kit following manufacturer’s instructions.
- Using a microtitre tray, add 20 μl of Reagent A across 7 wells of row A; this is to be used for the BSA standard samples. Add 20 μl of Reagent A across 3 wells of row B; this is to be used for your sample.
- To the 20 μl of Reagent A in row A, add 5 μl of each BSA protein standard (0, 0.125, 0.25, 0.5, 1, 2.5, 5 mg/ml) to the 7 individual wells. To the 20 μl of Reagent A in row B, add 5 μl of your sample to each of the three wells.
- Subsequently add 200 μl of Reagent B into all wells used and leave samples to react for 3 min at room temperature.
- Measure the absorbance of the wells using a Multiskan at 600 nm; create a standard curve using the BSA protein standards in an Excel spread sheet. Then using the equation of the line calculate the amount of protein in your sample. See Note 3 for experimental details, as well as Table 1 and Figure 1 as a representative example.
- Using the calculated protein concentration of your samples from step 13 adjust the final protein concentration to 1 mg/ml in PBS for cAMP analysis.
- Put all Cyclic AMP XPTM assay kit solutions and the cAMP XPTM Rabbit mAb coated microwell strips at room temperature.
- Dilute the 20x wash buffer in Milli-Q to 1x buffer.
- Prepare a 1x cell lysis buffer from the 10x stock solution provided in Milli-Q water, then add 1 mM of phenylmethylsulfonyl fluoride provided.
- Make fresh cAMP standards in 1x cell lysis buffer: In 450 μl of 1x cell lysis buffer add 50 μl of cAMP standard (2.4 M) to make 240 nM of cAMP.
- Using the 240 nM cAMP solution make a 1:3 serial dilution in the cell lysis buffer producing the following concentrations: 80 nM, 26.7 nM, 8.9 nM, 3.0 nM, 1 nM, 0.3 nM and 0 nM.
- For the unknown samples add 50 μl of the crude protein extract (at 1 mg/ml) in triplicate into the microwell strips. Subsequently, add 50 μl of provided horseradish peroxidase-linked cAMP solution to all standards and samples. Cover with aluminum foil and incubate at room temperature for 3 h on a horizontal orbital plate shaker (~30 rpm).
- Tip out the solution in the microwell strips and wash 4 times with 200 μl/well of 1x wash buffer; discard all liquid each time but do not allow wells to dry.
- Add 100 μl of 3, 3', 5, 5'-tetramethylbenzidinesubstrate solution provided in the kit and incubate the plate at room temperature for 30 min. Continue to monitor the wells as they will have a transparent color at the start of the reaction which will change to a yellow. Monitor wells for color change and when an intense yellow color is seen the reaction should be stopped; this may be necessary to do prior to the 30 min incubation time-frame described in the kit.
- Add 100 μl of stop solution and then immediately measure the absorbance at 450 nm see representative data 3 for example (Table 2 and Figure 2).
- Using the inverse for the standards derived from a standard curve, and adjusted for the amount of total protein added from the protein assay; calculate the amount of cAMP in nM/1 mg of crude protein extract.
Representative data
- A representative experiment demonstrating quantification of cAMP in the bacterium Acinetobacter baumannii can be found in Giles et al. (2015) Investigation of genes essential for pellicle formation in Acinetobacter baumannii.
- Below is an example of the excel BSA standard curve.
Table 1. BSA protein standards and their relative absorbance at OD600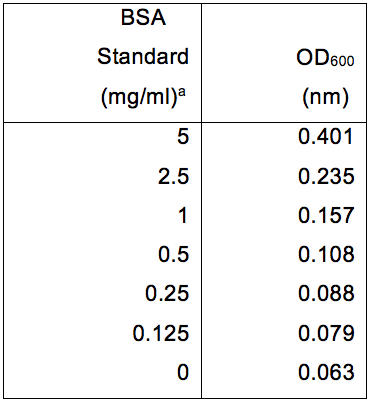
aAbsorbance was determined using a Protein Assay DCTM kit and read using a Multiskan EX. Data was subsequently plotted as seen in Figure 1.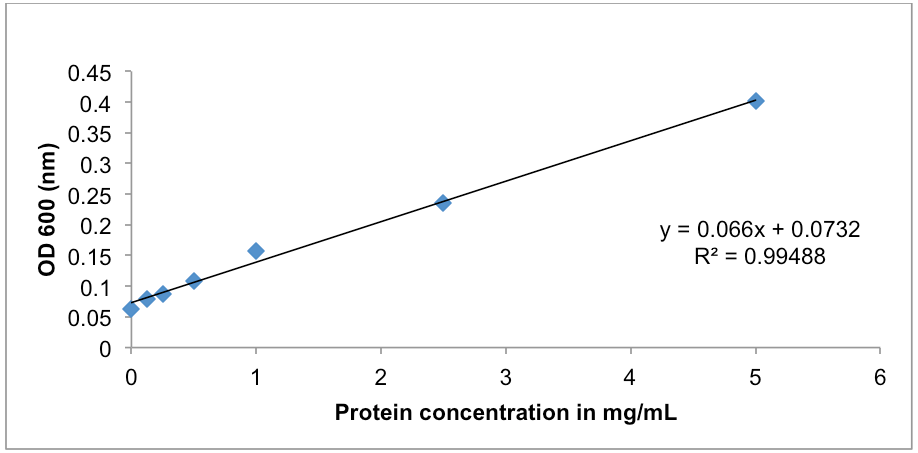
Figure 1. BSA protein standard curve. The absorbance at OD600 (nm) was determined for a range of BSA protein standards from 0-5 mg/ml. Protein concentration was determined using the Protein Assay DCTM kit. The line of best fit was plotted using y=mx+c (Excel 2003, MicrosoftTM). - A representative of the cAMP standards diluted in 1x cell lysis buffer is on page 1 of the Cyclic AMP XPTM assay kit manufacturer’s instruction; an example (Table 2 and Figure 2) is also provided below.
Table 2. cAMP standards and their relative absorbance at OD450 (nm)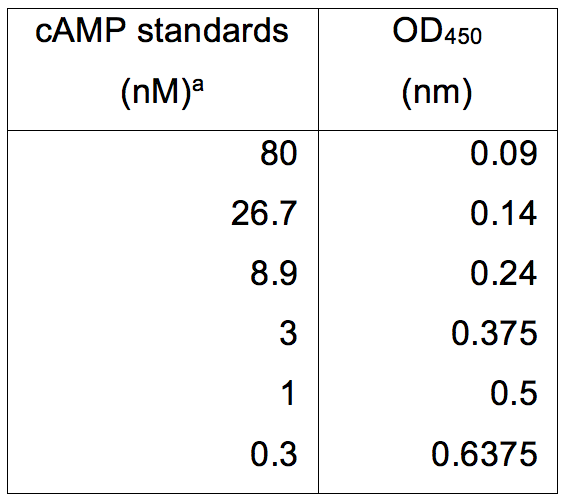
aThe amount of cAMP (nM) was determined using the Cyclic AMP XPTM Assay kit and absorbance read using a Multiskan EX. Data was subsequently plotted as seen in Figure 2.
Figure 2. cAMP concentration standard curve. The absorbance at OD450 (nm) was determined for a range of cAMP standards from 0.3-80 nM. cAMP concentration was determined using the Cyclic AMP XPTM Assay kit. The line of best fit was plotted using y=mln(x)+c (Excel 2003, MicrosoftTM). Using this standard curve, the quantity of cAMP can be determined for cell lysates.
Notes
- This protocol is optimized for measuring intracellular cAMP levels in Acinetobacter baumannii. For other bacteria, use the appropriate growth medium, antibiotics, and growth conditions.
- The method for reading OD600: Under sterile conditions pipette 1 ml of culture into a 1 ml cuvette and measure cellular density at an OD600 length in the spectrophotometer; repeat this every 1 h or until the culture reaches the required OD600 = 0.7.
- Protein assay using the Multiskan at OD600: after waiting 3 min at room temperature measure the samples in the microtitre tray, then output the data to an excel spread sheet for data manipulation and save. Spread sheet 1 should contain your Multiskan raw data. Then in spread sheet 2 column A row 1 type in BSA standard, column B type in OD600; In column A rows 2-8 type in the BSA standards numbers, in column B type in the Multiskan reads for the BSA standards (see Table 1). Then highlight all the data and make a scatter plot with no markers, and then add a trend line, add the equation of the line as well as a title and axis labels (see Figure 1). Your samples are in triplicate, average your samples, then insert your average number into the equation of the line as the Y intersect and find X: example Y = 0.066 X + 0.0732: if Y is 0.2, then 0.2 - 0.0732 = 0.1268/0.066 =1.92 mg/ml of protein in your sample.
- When waiting for the color change to occur from clear to yellow before adding the stop solution, it is imperative that the microwell plates are continually monitored throughout this reaction. After approximately 15 min, depending on the concentration of cAMP within a sample, the development of an intense yellow color will occur. Once the entire well is yellow add the stop solution.
- Cyclic AMP is stable for long periods of time when frozen (approximately 6 months at -20 °C), it can be stored for 2 weeks at 4 °C.
- The use of the Constant System one shot head results in reliable disruption of Acinetobacter baumannii cells. The efficiency of cell disruption using other mechanical of chemical methods or the use of differing species will need to be determined empirically.
Recipes
- Phosphate buffer saline (PBS)
For 500 ml:
4.0 g NaCl
0.1 g KCl
0.7 g Na2HPO4
0.1 g KH2PO4
Add dH2Oup to 500 ml
pH is adjusted with HCl to 7.4
Acknowledgments
This protocol was modified and adapted from the manufacturer’s instructions specifically for eukaryotic cells using the Cyclic AMP XPTM Assay kit. This project was financially supported by the National Health and Medical Research Council Australia Project Grant 535053 and a Flinders Medical Research Foundation Grant. SKG is the recipient of a Flinders University Research Scholarship (FURS).
References
- Giles, S. K., Stroeher, U. H., Eijkelkamp, B. A. and Brown, M. H. (2015). Identification of genes essential for pellicle formation in Acinetobacter baumannii. BMC Microbiol 15: 116.
- McDonough, K. A. and Rodriguez, A. (2012). The myriad roles of cyclic AMP in microbial pathogens: from signal to sword. Nat Rev Microbiol 10(1): 27-38.
- Wolfgang, M. C., Lee, V. T., Gilmore, M. E. and Lory, S. (2003). Coordinate regulation of bacterial virulence genes by a novel adenylate cyclase-dependent signaling pathway. Dev Cell 4(2): 253-263.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Giles, S. K., Stroeher, U. H. and Brown, M. H. (2016). Measurement of Intracellular cAMP Levels Using the Cyclic Nucleotide XP Enzymatic Immunoassay Kit in Bacteria. Bio-protocol 6(8): e1792. DOI: 10.21769/BioProtoc.1792.
Category
Microbiology > Microbial signaling > Secondary messenger
Microbiology > Microbial biochemistry > Other compound
Biochemistry > Other compound > cAMP
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












