- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Mouse Embryonic Stem Cell Differentiation to Hematopoietic Precursors
Published: Vol 2, Iss 7, Apr 5, 2012 DOI: 10.21769/BioProtoc.144 Views: 16027

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
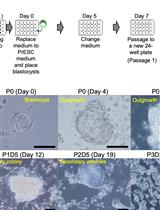
Generation of Mouse Primitive Endoderm Stem Cells
Yasuhide Ohinata [...] Haruhiko Koseki
Nov 20, 2023 2447 Views
Isolation of Human Bone Marrow Non-hematopoietic Cells for Single-cell RNA Sequencing
Hongzhe Li [...] Stefan Scheding
Jun 20, 2024 2556 Views
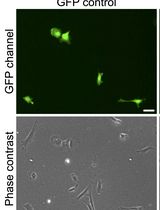
An Efficient Method for Immortalizing Mouse Embryonic Fibroblasts by CRISPR-mediated Deletion of the Tp53 Gene
Srisathya Srinivasan and Hsin-Yi Henry Ho
Jan 20, 2025 2742 Views
Abstract
Embryonic stem cells are derived from inner cell mass of an embryo that can differentiate into every cell type in the body. Clinically, cultured red blood cell supply is of great interest. However, some of the hurdles need to be overcome. This protocol describes a two step protocol to form embryoid body and differentiate them in to hematopoietic lineage cells, especially erythroid cells.
Materials and Reagents
- Mouse embryonic stem cell (ES cells)
- 200 mM L-Glutamine (100 ml) (Life Technologies, Gibco®, catalog number: 320-5030AG )
- Methylcellulose (STEMCELL Technologies, catalog number: 03134 )
- Gelatin (STEMCELL Technologies, catalog number: 07903 )
- Transferrin (Human) (Boehringer-Mannheim) (Roche Diagnostics, catalog number: 652202 )
- MTG (Sigma-Aldrich, catalog number: M6145 )
- Plasma derived serum (PDS) (Animal technologies) special order from company. Typically serum contains some PDGF, which inhibit differentiation (Must call company, animal technologies, Inc)
- Epo (Amgen) (Thermo Fisher Scientific, catalog number: 50948385 )
- Protein Free Hybridoma Media II (PFHM-II) (Life Technologies, Gibco®, catalog number: 12040-077 )
- Ascorbic acid
- Penicillin/Streptomycin (Pen/Strep) (Thermo Fisher Scientific, catalog number: SV30010 )
- Fetal bovine serum (FBS)
- Fetal calf serum (FCS)
- Phosphate buffered saline (PBS)
- Trypsin
- EDTA
- NaHCO3
- Dulbecco's modified eagle medium (DMEM)
- ES-IMDM (see Recipes)
- Differentiation medium (see Recipes)
- Secondary differentiation media (see Recipes)
- Iscove’s modified dulbecco medium (IMDM) (Life Technologies, Gibco®, catalog number: 12200-036 , 1 L/pack) (see Recipes)
- Cellulase (Sigma-Aldrich, catalog number: C-1794 ) (see Recipes)
Equipment
- Standard table-top centrifuges
- T25 flask
- Snap cap tube
- Incubator
- Water bath
- Gauge needle
- 50 ml Falcon tube
- 0.22 µm filter
Procedure
- Primary differentiation of ES cells (making Embryoid Bodies)
- Two days prior to setting up differentiation, split cells into ES-IMDM medium without feeder cells in gelatinized T25 flask (this dilutes out most of the STO cells). Add 4 x 105 ES cells per T25 flask.
- Change medium the next day (ES cells grow faster therefore use up nutrients faster; doubles every 8 h).
- Set up differentiation as follows.
- Aspirate medium from the flask.
- Add 1 ml of trypsin/EDTA, swirl around, and quickly remove.
- Add 1 ml of trypsin/EDTA and wait until cells start to detach. It usually takes about 1-2 min. Do not over-trypsinize cells.
- Deactivate the trypsin by adding 1 ml FCS (differentiation serum) and 4 ml IMDM and pipette up and down to make single cell suspensions. It is important not to have cell clumps. Transfer to 15 ml snap cap tube.
- Centrifuge for 5-10 min at 1,000 rpm.
- Resuspend the pellet in 10 ml IMDM (w/o FCS; this removes LIF and any other factors that might interfere with differentiation). Spin down cells for 5-10 min at 1,000 rpm.
- Resuspend the pellet in 5 ml IMDM (w/ differentiation serum) and count the VIABLE ES cell number using eosin (0.2% eosin in 1× PBS). Make sure to count ES cells only.
- Add 5,000-6,000 ES cells/ml of differentiation media for day 2.75-3 EBs. For ChIP purpose, I prepared 8-10 dishes and prepared enough media for one extra dish. Add 4,000-5,000 cells/ml for day 4-5 EBs. Add 500-2,000 cells/ml for day 6-10 EBs. Add more cells (7-8,000 cells/ml for day 2.75-3 EBs) if they differentiate poorly. Too many cells could result in poor differentiation of EBs. You don’t want the cells to use up the nutrients before they reach the certain days.
- Primary differentiation to derive EBs is typically done in 10 ml/10 cm dish. Use bacterial petri dish (Valmark (good), Fisher (poor) VWR (not recommended)). Do not use tissue culture dishes because all the cells will adhere and will not form EBs. If you have a way to constantly shake your dish in the incubator this improves efficiency, otherwise shake your dish 2-3 times a day until the EBs are ready.
- Aspirate medium from the flask.
- Two days prior to setting up differentiation, split cells into ES-IMDM medium without feeder cells in gelatinized T25 flask (this dilutes out most of the STO cells). Add 4 x 105 ES cells per T25 flask.
- Hematopoietic precursor analysis (secondary differentiation)
- Embryoid body (EB) harvest.
- EBs in liquid: Transfer media containing EBs into 50 ml tubes. Wash the dish with IMDM and pool. Let it sit at room temperature (RT) for about 10-20 min. EBs will settle down to the bottom of the tube.
- EBs in methylcellulose: Add equal volume of cellulase (2 units/ml, final 1 unit/ml) and incubate 20 min at 37 °C. Collect EBs in 50 ml tubes. Wash the dish with IMDM. Let it sit at RT for about 5-10 min. EBs will settle down to the bottom of the tube.
- EBs in liquid: Transfer media containing EBs into 50 ml tubes. Wash the dish with IMDM and pool. Let it sit at room temperature (RT) for about 10-20 min. EBs will settle down to the bottom of the tube.
- While the EBs are settling down prepare replating media enough for one extra plate (if you need 10 plates prepare media for 11 plates) look at the recipe for details.
- Aspirate off as much media as possible minimizing the EB loss, add 3 ml trypsin and incubate for 3 min at 37 °C (use water bath). Very important to keep the time. It is directly related with survival and differentiation potential of ES cells. If you are harvesting for FACS analysis, 7.5 mM EDTA should be used to dissociate the EBs. This is more harsh treatment, however, receptors are kept intact.
- Quickly vortex briefly and add 1 ml FCS (differentiation serum).
- Dissociate by passing 8-10 times through a 20 gauge needle. Transfer to 15 ml tube and spin for 5-10 min at 1,000 rpm. Use collagenase for older EBs (i.e. >day 9). When collagenase is used, incubate cells for 1 h at 37 °C.
- Resuspend the pellet in 0.3-1 ml of IMDM (w/ 10% FCS). Smaller the volume the better. Count the viable cells. At this point, there should be no cell clumps.
- Add EBs to methylcellulose replating media. Shake well and let it sit for few minutes for bubbles to rise up. Using the 16 gauge blunt-end needle dispence the media+EBs on to appropriate dishes (you can use any bacterial dish for this). Always prepare bit more than what you need. Methylcellulose is sticky.
- Small scale differentiation: Add 3-6 x 104 EBs ml-1 of methylcellulose replating media. Add 1.25 ml methylcellulose mix to each 35 mm bacterial dishes for blast colony assay. Otherwise, add 1 ml each. Prepare three replica dishes for each sample, i.e. make 4.5 ml methylcellulose replating media for blast colony assay and make 4 ml methylcellulose replating media for erythroid and other myeloid colony assay.
- For ChIP: Add 1-1.5 x 105 EBs/ml of methylcellulose replating media. Add 10 ml methylcellulose mix to each 10 cm dishes. 10 x 10 cm plates will yield enough cells for 4 IPs (8-10 x 107).
- Small scale differentiation: Add 3-6 x 104 EBs ml-1 of methylcellulose replating media. Add 1.25 ml methylcellulose mix to each 35 mm bacterial dishes for blast colony assay. Otherwise, add 1 ml each. Prepare three replica dishes for each sample, i.e. make 4.5 ml methylcellulose replating media for blast colony assay and make 4 ml methylcellulose replating media for erythroid and other myeloid colony assay.
- Place the plates in 245 mm square dish and a small 35 mm dish with water to maintain the humidity. This prevents the gel from drying up and cracking.
- Embryoid body (EB) harvest.
- Harvest
- Prepare celluase + PBS solution (2 μg/ml)
- Add same volume of cellulase solution to each plate so the final concentration of cellalase is 1 μg/ml.
- Incubate at 37 °C for 30-40 min until methocell is watery.
- Scrape plates and collect cells in 50 ml Falcon tube. Vortex and spin at 1,000 rpm, 5 min.
- Wash with fresh media or PBS and spin at 1,000 rpm, 5 min.
- Prepare celluase + PBS solution (2 μg/ml)
Recipes
- IMDM
For 1 L
1 pack IMDM powder
10 ml/liter Pen/Strep
3.024 g/L NaHCO3
Filter thru 0.22 µm filter
If using premade liquid media -> add 5.05 ml of P/S - ES-IMDM
This media is essentially the same as ES-DMEM but use IMDM (richer media) instead of DMEM. Used for gelatin+ES cell culture. This is used for gelatin+ES cells and for primary differentiation to generate EBs. - Differentiation medium
15% pre-selected FCS in IMDM with the followings
1% ascorbic acid (5 mg/ml): Make ascorbic acid fresh each time you set up a differentiation. Dissolve ascorbic acid (5 mg/ml) in H2O and filter sterilize (0.22 µm).
1% L-glutamine: Once thawed use it for 1 wk.
3 µl/ml MTG (MTG is diluted by adding 26 µl of MTG to 2 ml IMDM). - Cellulase
2 U/ml in 1× PBS filter sterilize with 0.45 µm filter. - Secondary differentiation media (methylcellulose mix)a (all % is v/v)
Erythroid 115 ml for 10 plates Methylcelluloseb 44% 46 ml PDS 10% 11.5 ml Ascorbic acid (5 mg/ml) 0.25% 287 µl L-glutamine 1% 1.15 ml Transferrin 1% 1.15 ml MTG (26 µl per 2 ml)c 3 µl/ml 345 µl Epo 2 unit/ml 5.75 µl PFHM-IId 5% 5.75 ml IMDM To 100% 48.8 ml - If efficiency of EB plating decreases, replace one or more of the following with fresh aliquots or lots: PFHM II, IMDM, MTG, FBS.
- Prewarm methylcellulose to 37 °C to decrease viscosity. Discard after it has been thawed and stored at 4 °C for >1 month. If desired, methylcellulose can be spun down at 6,000 rpm for 10 min to remove debris (transfer to new conical when spin is complete). Methocell you get from stem cell tech. is 2.5%. Calculations are made so that final methocell is 1.1%.
- Close MTG and PFHM II caps immediately after use to prevent oxidation. When opening a new 50 ml aliquots of PFHM II, sub-aliquot into five 10 ml aliquots to prevent oxidation (cover conical tubes with foil to protect from light).
- Various Factors added include bFGF (10 ng/ml), activin A (2-10 ng/ml), BMP-2 (2-10 ng/ml), and BMP-4 (2-10 ng/ml). Add KL (final 1%) and IL-3 (final 1%) for >day 6 EBs. Feed for > day 9 EBs on day 6 (4-5 ml per 10 cm petridish). Make the same methyl cellulose cocktail with 20% methylcellulose instead of 44%.
- If efficiency of EB plating decreases, replace one or more of the following with fresh aliquots or lots: PFHM II, IMDM, MTG, FBS.
Acknowledgments
This protocol was adapted from Im et al., (2003). Funding for this work was obtained from the NIH.
References
- Zhang, W. J., Chung, Y. S., Eades, B. and Choi, K. (2003). Gene targeting strategies for the isolation of hematopoietic and endothelial precursors from differentiated ES cells. Methods Enzymol 365: 186-202.
- Im, H., Park, C., Feng, Q., Johnson, K. D., Kiekhaefer, C. M., Choi, K., Zhang, Y. and Bresnick, E. H. (2003). Dynamic regulation of histone H3 methylated at lysine 79 within a tissue-specific chromatin domain. J Biol Chem 278(20): 18346-18352.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Im, H. (2012). Mouse Embryonic Stem Cell Differentiation to Hematopoietic Precursors. Bio-protocol 2(7): e144. DOI: 10.21769/BioProtoc.144.
- Im, H., Park, C., Feng, Q., Johnson, K. D., Kiekhaefer, C. M., Choi, K., Zhang, Y. and Bresnick, E. H. (2003). Dynamic regulation of histone H3 methylated at lysine 79 within a tissue-specific chromatin domain. J Biol Chem 278(20): 18346-18352.
Category
Stem Cell > Embryonic stem cell > Maintenance and differentiation
Developmental Biology > Cell growth and fate
Stem Cell > Adult stem cell > Hematopoietic stem cell
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








