- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Scanner-based Time-lapse Root Phenotyping
Published: Vol 5, Iss 6, Mar 20, 2015 DOI: 10.21769/BioProtoc.1424 Views: 14102
Reviewed by: Samik BhattacharyaArsalan DaudiRenate Weizbauer

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Closed Systems to Study Plant–Filamentous Fungi Associations: Emphasis on Microscopic Analyses
Vasiliki Skiada and Kalliope K. Papadopoulou
Feb 20, 2025 2855 Views

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1886 Views

Direct Plant Regeneration From Immature Male Inflorescence of Banana (Musa spp.)
Pradeep Chand Deo
Oct 20, 2025 1426 Views
Abstract
Non-destructive phenotyping of root system architecture can facilitate breeding for root traits that optimize resource acquisition. This protocol describes the construction of a low-cost, high-resolution root phenotyping platform, requiring no sophisticated equipment and adaptable to most laboratory and glasshouse environments. The phenotyping platform consists of germination paper abutting scanners and open-source image acquisition software which allows live imaging of root systems. The phenotyping platform is scalable, modular and inexpensive.
Keywords: Root system architectureMaterials and Reagents
- Seeds of the plants to be investigated
Note: We used seeds of Brassica rapa L. - Nutrient solution (see Recipes)
Note: Composition will depend on experimental requirements.
Equipment
- Large (30 x 42 cm) and small (12 x 12 cm) steel blue Anchor seed germination paper (Anchor Paper Company, catalog number: SGB1924B) for phenotyping platform and seed germination, respectively
- Duct tape (office DEPOT)
- Ferrite magnetic strip with adhesive back (0.75 mm thick) (RS Components, catalog number: 297-9116)
- Square bioassay or petri dishes (120 mm x 120 mm x 17 mm) (Sigma-Aldrich, catalog number: Z617679)
Note: This could be improvised with empty compact disc (CD) cases or any kind of transparent box that meets the specified dimensions. - Black Perspex plates (21.5 x 30.0 cm) custom-made from opaque cast acrylic boards (Perspex Cast 9T30 Black Sheet) (plastock)
- Computer
Note: This should be Windows-based and should have a directory with sufficient space to save images. In our platform, a Canon flatbed scanner imaging a root system at 300 dpi produces a single image of 40.2 MB. With 24 scanners in operation and images taken every 12 h, a minimum free space of approximately 30.0 GB is required for an experiment lasting 14 days. We used a Dell computer Intel® CoreTM i3-4150 Processor- Dual Core, 4 GB 1,600 MHz Memory and 500 GB Hard Drive for our experiments but images were saved on 2 TB WD external hard drive. - Canon flatbed scanners (CanoScan, model: 5600F)
Note: Any scanner with a TWAIN driver can be used. A TWAIN driver is software that comes as part of the software package of digital imaging devices such as scanners and cameras. The TWAIN driver provides an interface for communication between the software and the hardware of digital imaging devices (http://www.twain.org/). In our platform, the TWAIN driver translated commands from our ArchiScan image acquisition software (http://www.archiroot.org.uk/) into instructions to control the hardware of the scanners.- Setting up scanners
- Remove the cover of the scanner as shown in Figure 1.
- Attach magnetic strips to the short-edges of the scanner window (Figure 1B).
Note: This is optional as duct tape could be used to hold plates to scanners.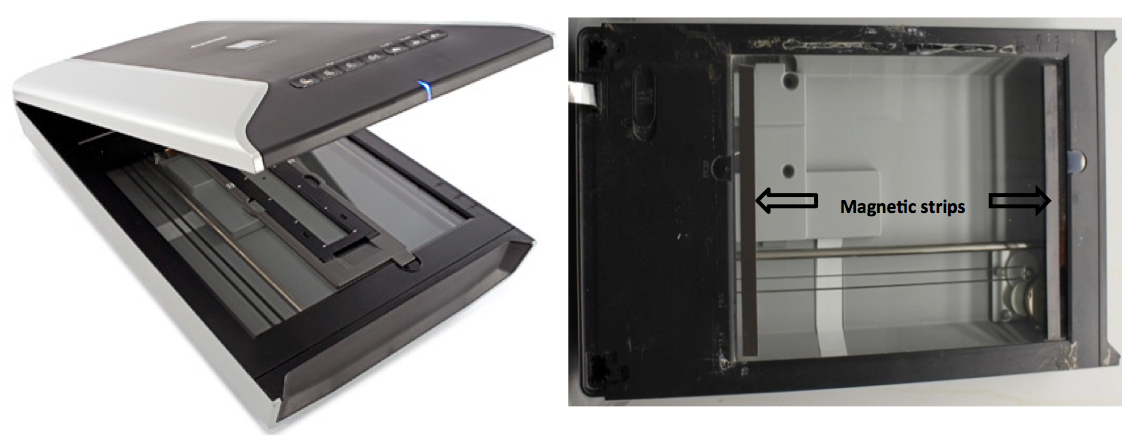
Figure 1. A. Original flatbed scanner. B. Modified flatbed scanner with cover detached.
- Remove the cover of the scanner as shown in Figure 1.
- Setting up scanners
- Support for germination paper
- Glue to the edges of the Perspex plates strips of Perspex (0.3 x 0.3 x 30 cm and 0.3 x 0.3 x 21.5 cm for the long-edges and short-edges of the Perspex plate, respectively).
Note: This is to provide some separation between the plate and the scanners. - Allow two gaps in the Perspex strips, each approximately 3 cm wide, along the long edge of the Perspex plates to allow gas exchange with the surrounding atmosphere and unimpeded shoot growth (Figure 2).
- Fix magnetic strips along the two short-edges of the Perspex plate to aid abutting of the Perspex plate to the scanner window (optional).
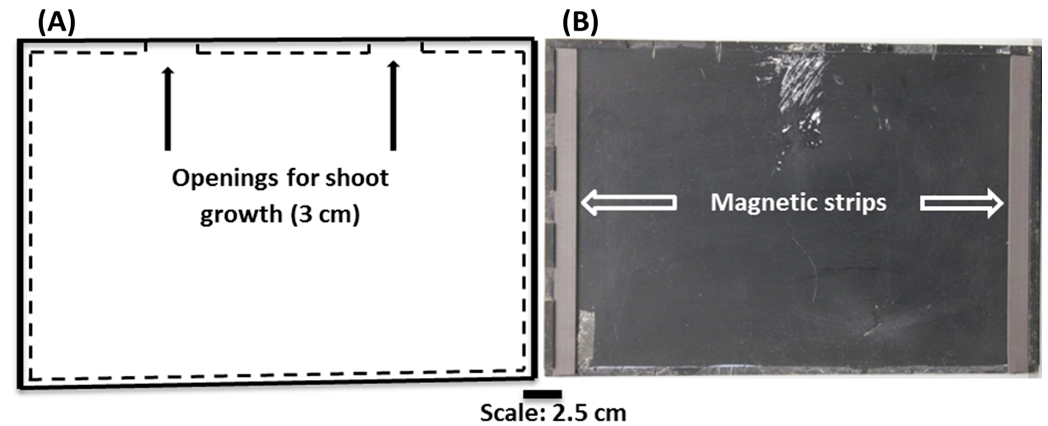
Figure 2. (A) Schematic representation of the Perspex plate used as a scanner cover and also to attach germination paper onto scanners. (B) Black Perspex plates used as scanner covers and also to hold germination paper vertically on scanner surfaces
- Glue to the edges of the Perspex plates strips of Perspex (0.3 x 0.3 x 30 cm and 0.3 x 0.3 x 21.5 cm for the long-edges and short-edges of the Perspex plate, respectively).
Software
- ImageJ (http://imagej.nih.gov/ij/)
- ArchiScan and ImageJ custom macros for convex hull and particle analyses (Adu et al., 2014; http://www.archiroot.org.uk/ )
- SmartRoot (Lobet et al., 2011; http://www.uclouvain.be/en-smartroot)
Procedure
- Seed germination
Note: This procedure was performed in a sterile hood, but the whole process was not necessarily sterile since scanners and tanks were not autoclaved and the experiments were not performed in a sterile room.- Line the bottom of a square petri dish (12 x 12 cm) with paper towel.
Note: Empty CD cases can be used instead of petri dishes. - Cover the paper towel with a small germination sheet (12 x 12 cm).
- Spray the germination papers with deionized water.
- Put seeds on top of the wet seed germination paper. Allow each seed as much space as possible.
Note: Seeds should be surface sterilised prior to germination. Brassica rapa seeds were surface sterilised for 10 min in 10% hydrogen peroxide, followed by 1 min in 70% ethanol, and then rinsed 3 times with deionised water. - Place one layer of paper towels on top of the seeds.
- Spray the top layer of paper towels with deionized water.
- Place the lid on the petri dish and allow excess water to drain off by holding in a tilted position for about 30 sec.
- Cover the petri dish with aluminium foil.
- Place the petri dish in a vertical orientation in an incubator at 20 °C.
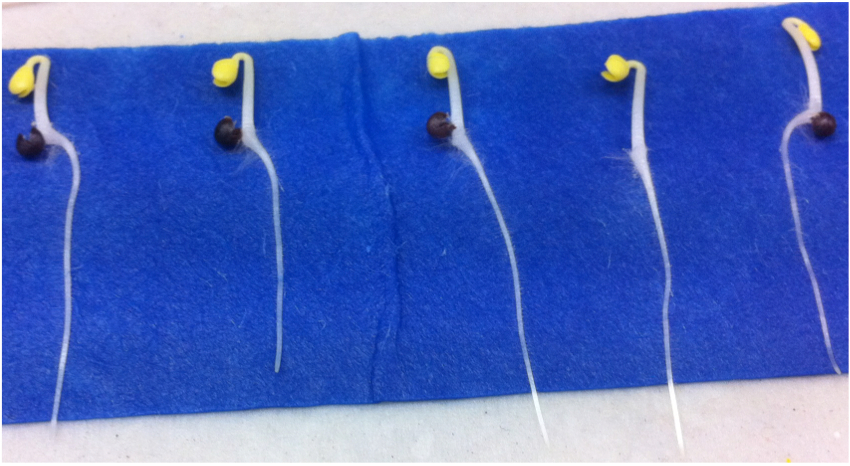
Figure 3. Example of Brassica rapa seeds after 3 days of germination
- Line the bottom of a square petri dish (12 x 12 cm) with paper towel.
- Transferring seedlings onto scanners
- Fix the top half of a large germination paper (30 x 42 cm; www.anchorpaper.com/) onto a Perspex plate.
Note: Scanner surfaces were cleaned with Virkon® disinfectant (http://www2.dupont.com/). Similarly, tanks and Perspex plates were thoroughly washed with Virkon® disinfectant and dried after each experiment. Slightly moistening the paper enables it to be fixed to the plate easily. - Transfer seedlings of similar size with radicles 2-3 cm in length (Figure 3) to the large (30 x 42 cm) sheets of germination paper fixed to Perspex plates.
- Transfer two seedlings per plate.
- Ensure that about 0.5 cm of the small germination paper surrounding each radicle is cut and transferred with the seedling to the large germination papers to minimize disturbance during the transfer process. (Figure 21 shows a video of this).
- Use a moistened tissue or parafilm tape, just below the cotyledons, to stick the seedlings to the large germination papers.
Note: Put the small germination paper that has been cut with the seedling on it at the top of the large germination paper and tape both to the large germination paper by putting the parafilm tape just below the cotyledons of the seedling. Alternatively, in the absence of parafilm tape, a damp, narrow strip of tissue paper (about 4 cm long) can be used to affix the seeds plus small germination paper to the large germination paper.
- Transfer two seedlings per plate.
- Attach the assemblage of Perspex plate, large germination paper and seedlings to the scanners using magnetic strips fixed to the plates and scanner windows.
Note: Duct tape can also be used hold the Perspex plate to the scanner (Figure 4). - Position scanners in near-vertical positions 5 cm above 20 L of nutrient solution contained in opaque polyvinyl plastic tanks, each supplying six scanners (Figure 4).
- The first scanners put on the tank could lean against a wall to provide support and subsequent scanners can be stacked with each leaning against the previous one (Figure 4).
- Allow approximately 10 cm of the large germination paper to be submerged in the nutrient solution (Figure 4).
- Connect all scanners to the computer on which the ArchiScan software is installed using a USB hub with multiple ports. Connect scanners, computers and USB ports to a power supply.
Note: You can use either one computer or multiple computers to control the scanners. The number of scanners per computer depends on the number of plants to be screened, duration of experiment, frequency of scanning and the computers’ storage capacity. In our experiments, there are occasions when up to 24 scanners are managed by one computer but we normally use 2 computers to control 24 scanners. On the latter occasions, each computer managed 12 scanners and scanners were connected to the computer using a D-Link USB Hub; http://www.misco.co.uk/; Misco number: 94282).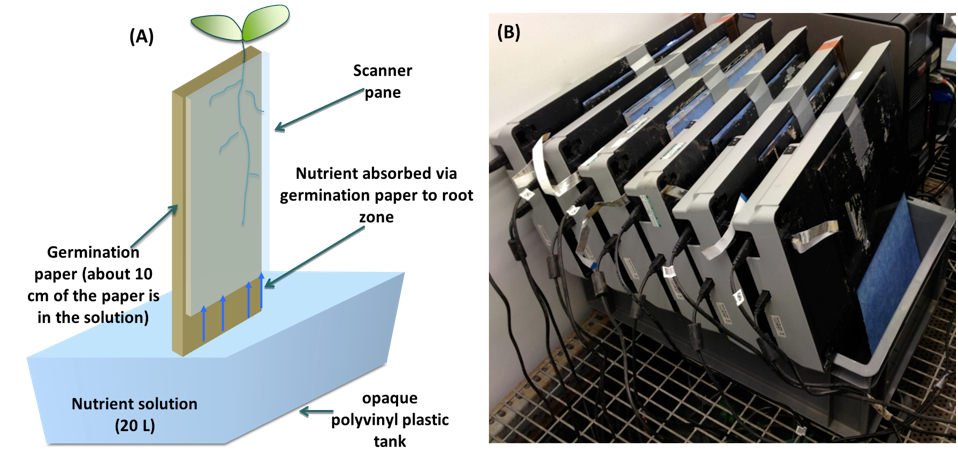
Figure 4. A. Schematic representation of the pouch-and-wick system of a single scanner. Roots grow on the surface of germination paper held between a clear-Perspex plate and the glass window of a scanner. The scanner is fixed in near-vertical position 5 cm above 20 L of nutrient solution contained in opaque polyvinyl plastic tank. Approximately 10 cm of the germination paper is submerged in the nutrient solution (Adu et al., 2014). B. Phenotyping platform composed of a bank of six scanners.
- The first scanners put on the tank could lean against a wall to provide support and subsequent scanners can be stacked with each leaning against the previous one (Figure 4).
- Fix the top half of a large germination paper (30 x 42 cm; www.anchorpaper.com/) onto a Perspex plate.
- Time-lapse imaging of roots
Run the image acquisition software, ArchiScan. ArchiScan is Windows-based software that provides an interface for managing multiple scanners and for simultaneous scanning of images. ArchiScan must be installed on a computer directory on which there is sufficient space to save images (Extended information on ArchiScan is found in the ‘READ ME’ pdf file at http://www.archiroot.org.uk/index.php/tools/ArchiScan).- Input parameters of the new project including:
Initial scanning time
Note: This depends on experimental requirements. In this example, we began scanning at Time=0; Figure 5.
Project duration
Note: This is indicated by Number of Scans (Figure 5) and depends on the frequency of taking images and duration of experiment. e.g.: If images are taken every 24 hours and the experiment was to last 15 d, the number of scans would be 16.
Time period between consecutive scans (Time between Scan; Figure 5)
Note: In most of our experiments images are scanned every 12 h.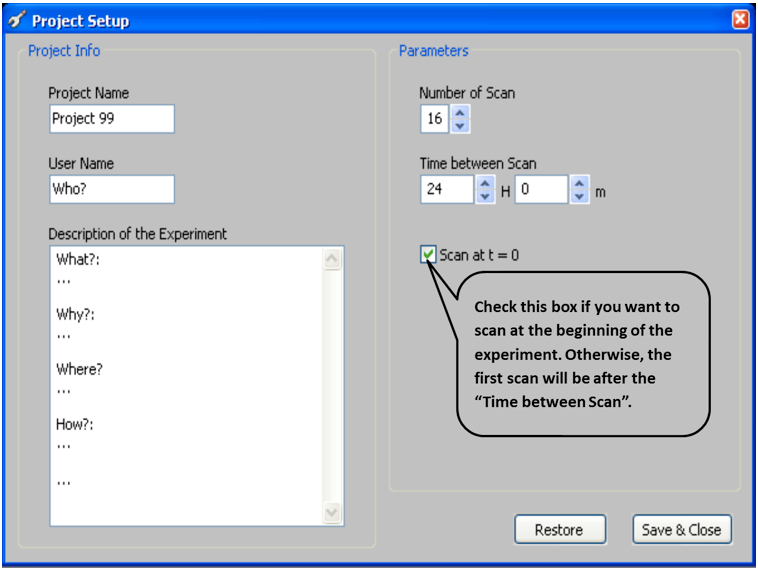
Figure 5. ArchiScan interface for setting experiment parameters - Set desired image parameters including (Figure 6)
Colour (black and white, greyscale or red green blue-RGB channel images)
Resolution (dot per inch-dpi)
Scaling (percent)
Frame size (x10, inch)
Brightness
Contrast
File format (bmp, jpeg etc.)
Note: For our experiments, the following parameters were used: Colour=RGB, Resolution = 300 dpi and default settings (Figure 6) were maintained for the remaining parameters.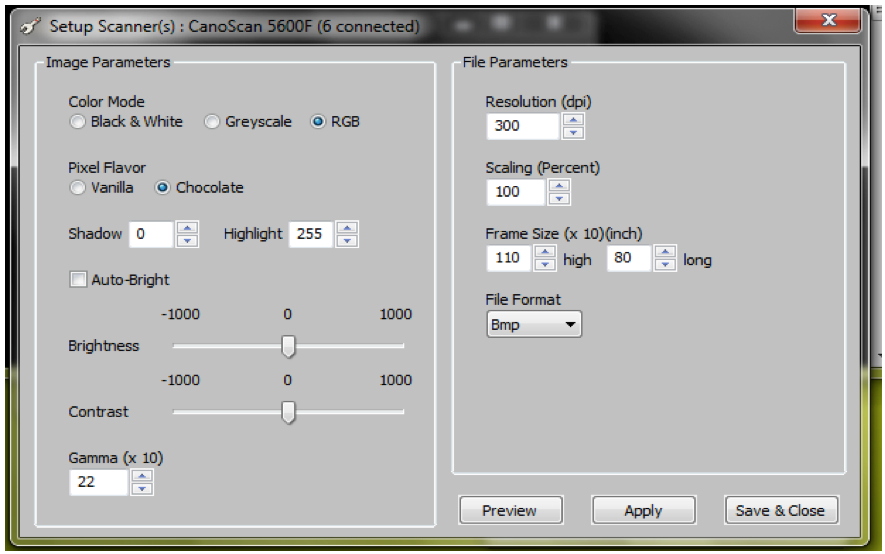
Figure 6. ArchiScan interface for setting image parameters
- Input parameters of the new project including:
- Image segmentation and extraction of root features
Image segmentation and extraction of root features from single root images or from a stack of time-lapse image sequences can be performed with ImageJ (http://imagej.nih.gov/ij/) using custom macros that can be downloaded from http://archiroot.org.uk/index.php/tools/ArchiScan. Extraction of root features can also be done with a variety of software including SmartRoot (Lobet et al., 2011; http://www.uclouvain.be/en-smartroot). - Segmentation and extraction of root features using ImageJ
Total root length:- Total root length of a single image
- To process and analyse a single root image, open the image with ImageJ (File>Open; Figure 7) and proceed from point 3 below.
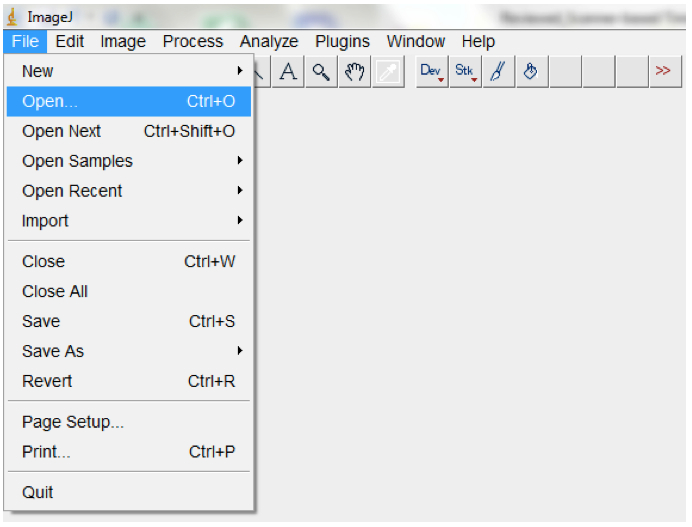
Figure 7. To open a single image, open ImageJ and select File > Open. Navigate to your folder and select the image, then select OK.
- To process and analyse a single root image, open the image with ImageJ (File>Open; Figure 7) and proceed from point 3 below.
- Total root length of a time-lapse stacked images to measure the rate of root length increase per unit time
- For each plant, import the image stack consisting of image sequences from the beginning to the end of an experiment into ImageJ (File>Import>Image Sequences; Figure 8).
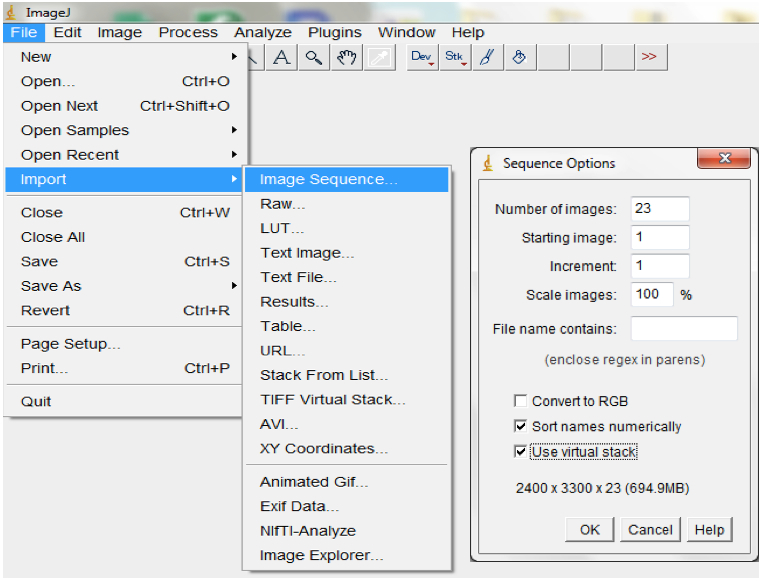
Figure 8. To import a sequence of images, open ImageJ and select File > Import > Image Sequence. Navigate to your folder and select the first image, then select OK. ImageJ will launch a sequence options dialog. The default values are usually appropriate but images could be resized here. We typically maintain the default 100% but checking the ‘Use Virtual Stack’ box opens images as a read-only virtual stack and allows image sequences too big to fit in RAM to be opened. - Set the scale or images by indicating the correct relationship between distance in pixels and the desired unit of measurement before analyses (Analyse > Set Scale; Figure 9).
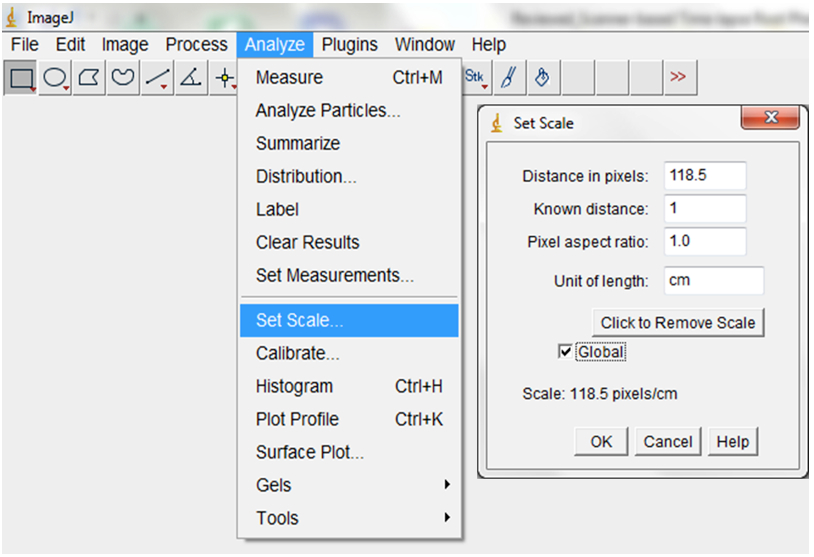
Figure 9. To set the scale for your images, select Analyse > Set Scale (e.g.: for our images, we previously established that 118.5 pixels were equal to 1 cm). Checking the ‘Global’ box ensures that the set scale would apply to all images in the imported images sequences and to all subsequent sequences of images that will be imported as long as ImageJ remains opened. - Convert RGB images to 8-bit greyscale images (depending on the quality of the images, they could also be split into the Red, Green or Blue colour channels and the best colour channel chosen for the analyses; Image>Type>8-bit; Image
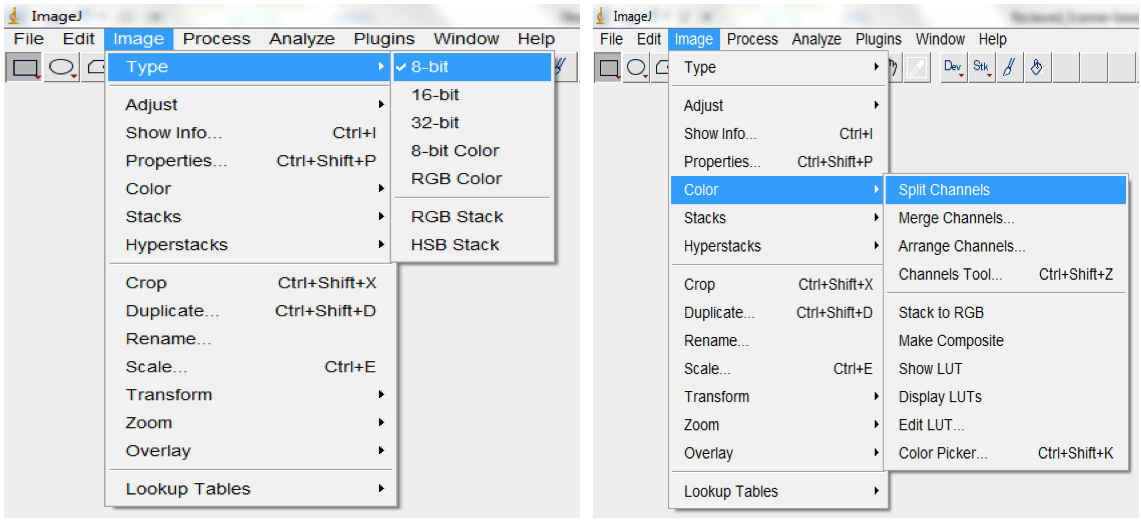
Figure 10. To Convert RGB images to 8-bit greyscale images, select Image>Type>8-bit or select Image<Colour Split Channels to split images into the Red, Green or Blue colour channels - Filter with the median, followed by the Gaussian, filter to remove short range variations on the image (i.e. remove variations introduced by water droplets condensing at the surface of scanners or due to inhomogeneity of the surface of the germination paper; Process>Filter>Median; Process>Filter>Gaussian Blur; Figure 11).
Note: The range of radii for filtering depends on the quality of the image but we typically use a range of 2-4.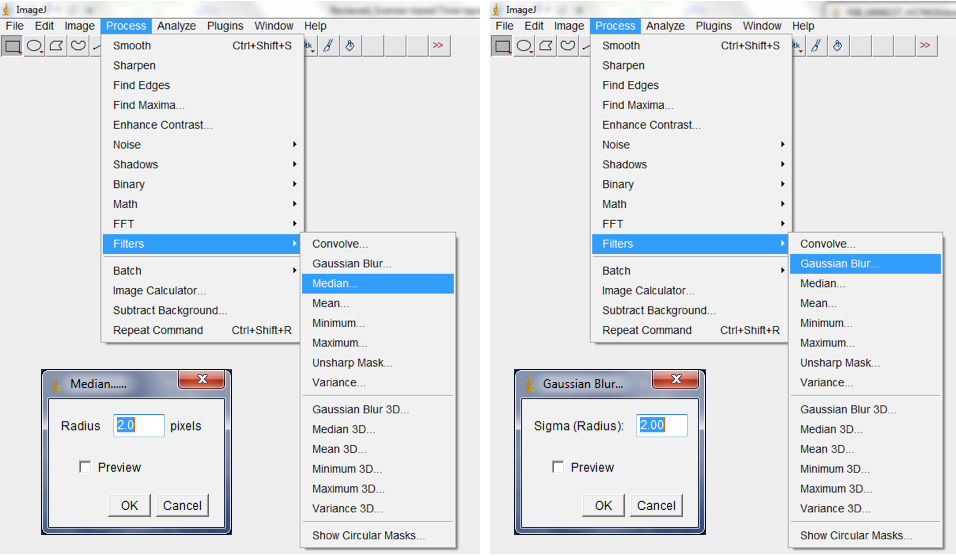
Figure 11. Select Process>Filter>Median, followed by Process>Filter>Gaussian Blur to remove short range artefacts on your image/images. A Median and Gaussian Blur options dialog box would be lunched for the radius and Sigma radius (Gaussian Blur) to be modified subject to the quality of your images. - Use background subtraction to remove variations in pixel intensity over longer distances than the root diameter (long range variations are artefacts or inhomogeneities in the background that remain after short range variations have been removed and may be caused by non-uniform moisture content of the germination paper; Process>Subtract Background; Figure 12).
Note: The range of radii for background subtraction depends on the quality of the image but we typically use a range of 40-50.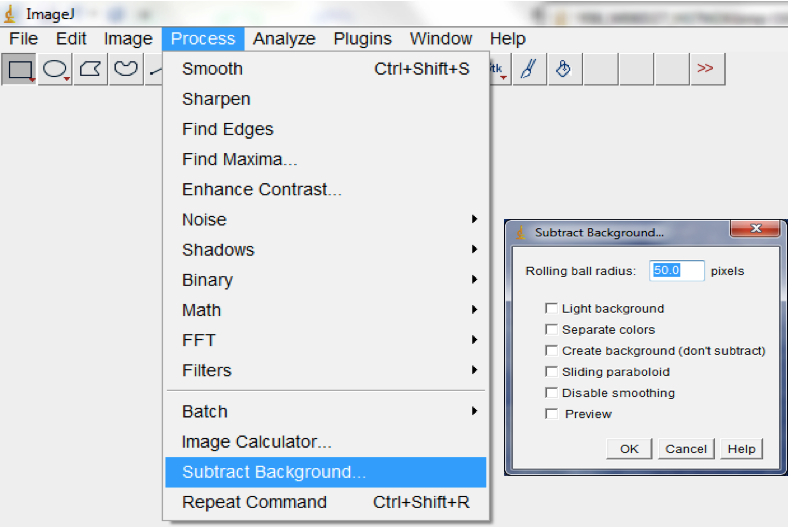
Figure 12. Select Process>Subtract Background to remove long range artefacts on your images. A Subtract Background dialog box would be lunched. We normally modified the ‘Rolling ball’ radius to optimise the background subtraction. - Auto-threshold with the Moment-preserving threshold algorithm.
Note: depending on the quality of the image this algorithm may be suboptimal compared to other auto-threshold algorithms in ImageJ. Other algorithms could be tried or a second algorithm could be added; Image>Adjust>Auto threshold; Figure 13).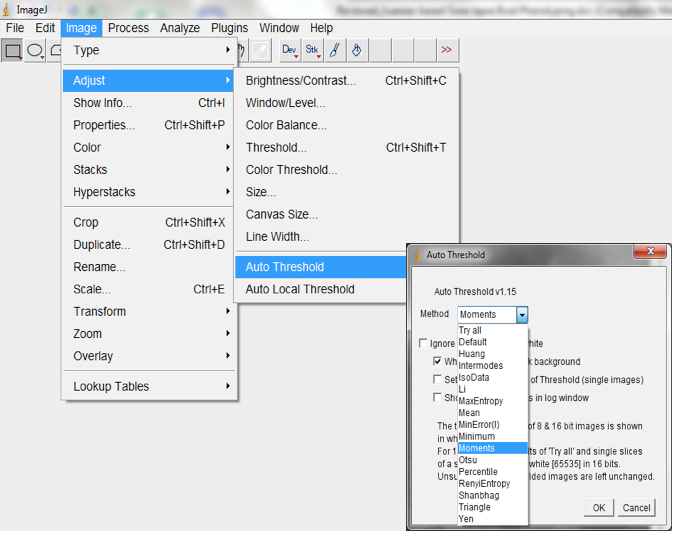
Figure 13. Select Image>Adjust>Auto threshold to threshold images. An Auto Threshold dialog box will be lunched. We typically use the ‘Moments’ threshold method but other algorithms could be tried to select the most optimal for your images. - Convert processed image to mask (Process>Binary>Convert to Mask; Figure 14, Figure 19).
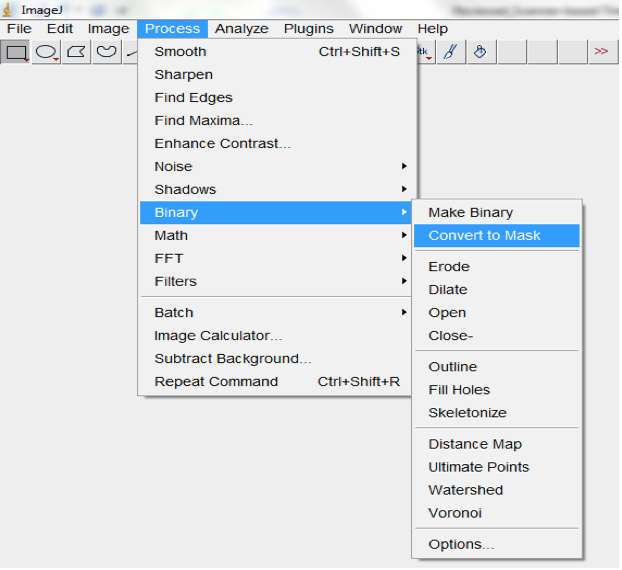
Figure 14. Select Process>Binary>Convert to convert processed image/images to mask - Run particle analyses to measure root features of interest including the root perimeter from which the total root length can be calculated. Note: different size and circularity options might be tried to get the best results. Selecting Show>Outlines in the Particle Analyses pop up box will show the root perimeter and ticking the Display results box will show the measurement of all of the particles; Analyse>Analyse particles; Figure 15).
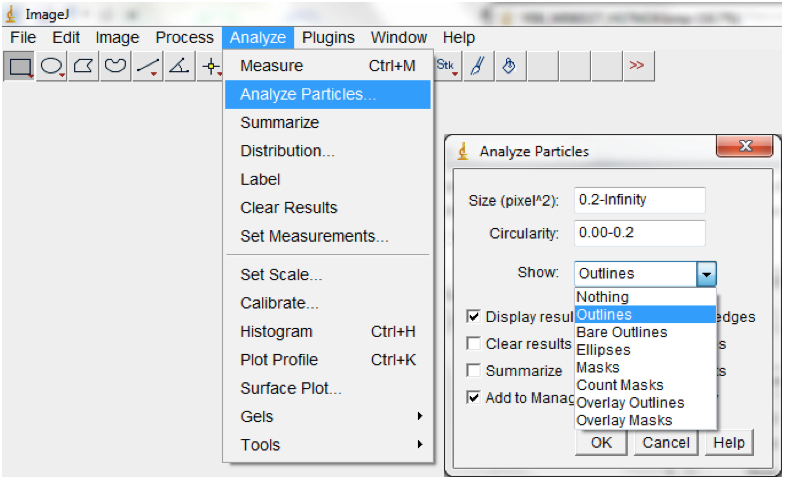
Figure 15. Select Analyse>Analyse particles to measure root traits of interest. An Analyse Particles dialog box will be lunched for the Size and Circularity to be modified Note: circularity is a dimensionless shape descriptor which is a function of the object perimeter and the area and gives a measure of similarity between a given shape and a perfect circle. It ranges between 0-1. We typically used Size = 0.2 and Circularity = 0.00 -0.2. The results will be for a single image (if only one image was opened) or for sequence of images (if time-lapse stacked images were imported into ImageJ) from which rate of change in total root length per unit time can be calculated. 50% of the root perimeter = total root length.
Area of roots: - From #6, run Gaussian Blur filter again (Figure 11).
- Find the local maxima to select root pixels on the image (Process>Find Maxima; Figure 16).
Note: This could be modified to optimise it for your images but we typically used noise=0 output=Point Selection and exclude edges.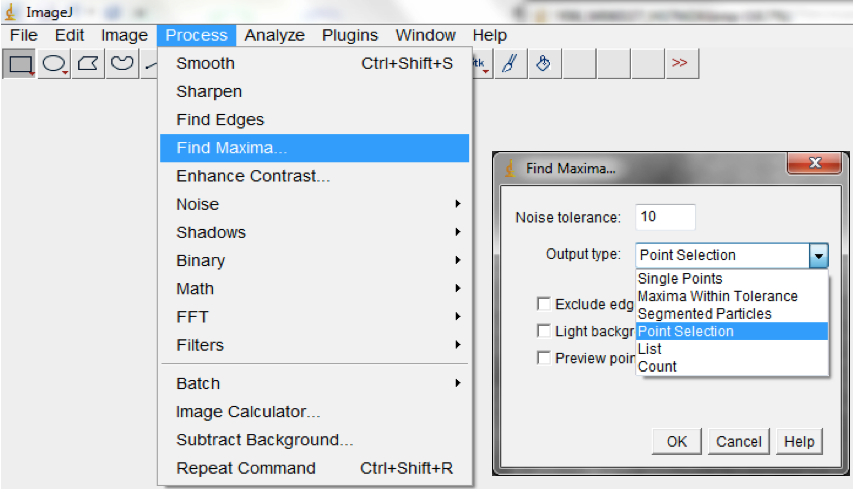
Figure 16. Select Process>Find Maxima to select root pixels on the image. A Find Maxima dialog box will be lunched. The Noise tolerance can be modified and we typically use the Point Selection Option. - Fit Convex Hull (Edit>Selection>Convex Hull; Figure 17; Figure 19).
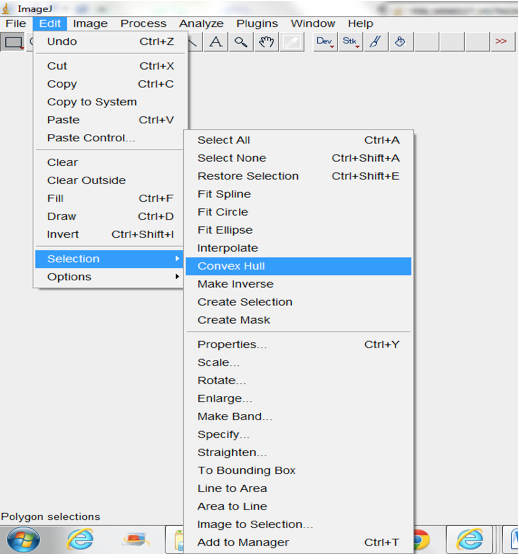
Figure 17. Select Edit>Selection>Convex Hull to fit convex hull around images
Note: Convex hulls are the smallest polygon whose vertices are all points in the image and define a rectangular selection that is the same size as the root system image. - Measure the area of the hull (Analyse>Measure; Figure 18).
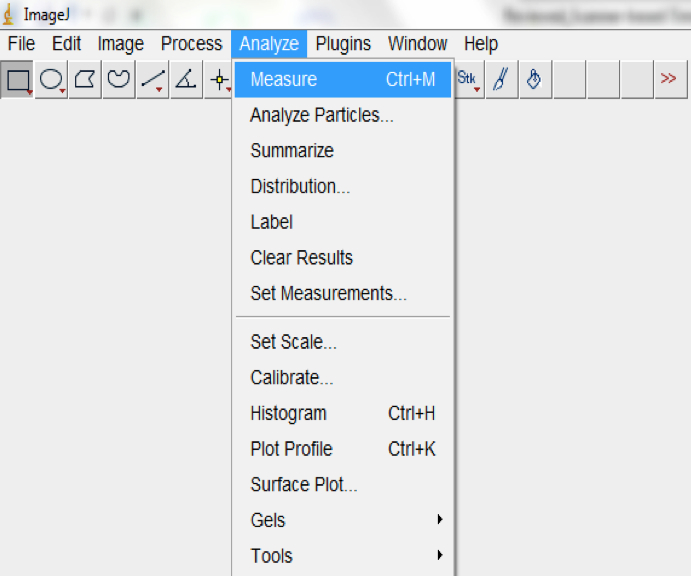
Figure 18. Select Analyse>Measure to measure the area of the hull (Surface area of the root system)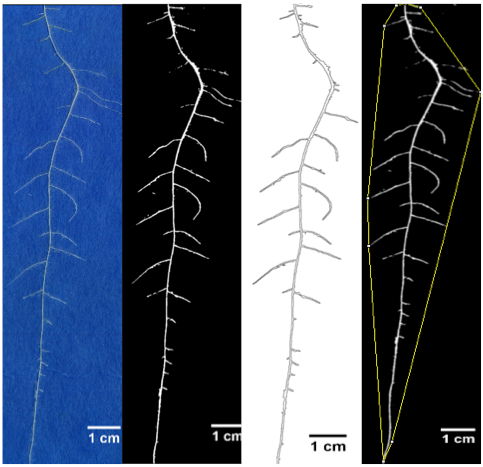
Figure 19. Sample image showing original root RGB image, segmented root image, perimeter or root for measuring root length and convex hull for measuring root area
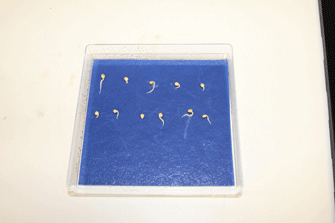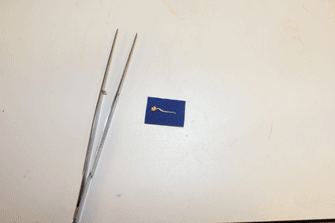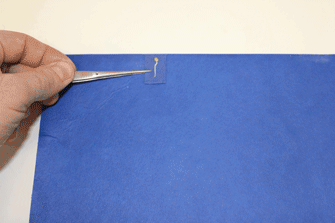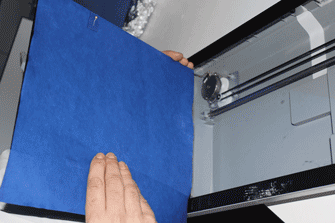
Figure 20. Representative data showing animated video of time-lapse segmented root system
- For each plant, import the image stack consisting of image sequences from the beginning to the end of an experiment into ImageJ (File>Import>Image Sequences; Figure 8).
- Total root length of a single image
Recipes
- Nutrient solution
Reagent Company (Catalogue No.) 1 L Stock Solution 20 L Nutrient Solution Reagent to add (g/L) Conc. (M) Volume of Stock to add (ml) Final Conc. (mM) Ca(NO3)2 VWR International (22388.292) 236.15 1 40 2 NH4NO3 VWR International (21278.295) 80 1 40 2 MgSO4 VWR International (25165.292) 92.43 0.5 30 0.75 KOH VWR International (26668.296) 29.92 0.5333 19 KH2PO4 VWR International (26936.293) 36.3 0.2667 19 0.25 FeNaEDTA VWR International (280233V) 18.35 0.05 40 0.1 CaCl2 VWR International (22317.260) 367.55 2.5 200 25 H3BO3 VWR International (20185.260) 185.49 3 200 30 Na2MoO4 VWR International (27937.236) 120.975 0.5 20 0.5 ZnSO4 VWR International (29253.236) 143.775 0.5 40 1 MnSO4 VWR International (ALFAB22081.22) 223.06 1 200 10 CuSO4 VWR International (23174.233) 124.84 0.5 120 3
Make up nutrient solution in 20 L deionised water and adjust to pH 6 using H2SO4.
Note: Nutrient solution could be replaced or changed depending on experimental requirements. In our experiments we did not replace our solution for experiments that lasted up to 14 day. Nutrient solutions should ideally be changed for experiments of longer duration as there could be depletion of nutrients and/or growth of microorganisms in the solution. We also experienced fungal growth on the germination papers occasionally. - Growth room requirements
Day/night cycle: 16/8 h
Temperature: 15 °C
Light intensity during the day at plant height: 100 μmol/m2/s
Relative humidity: 60%
Acknowledgments
This protocol was adapted from previous work Adu, 2014 and Adu et al., 2014. This work was supported by the Rural and Environment Science and Analytical Services Division (RESAS) of the Scottish Government through Work Package 3.3, ‘The soil, water and air interface and its response to climate and land use change’ (2011-2016). Funding for this study was provided by a BBSRC Professorial Fellowship (to M. J. B.). M.O.A. was supported by the University of Nottingham Vice-Chancellor’s Scholarship for Research Excellence.
References
- Adu, M. O. (2014). Variations in root system architecture and root growth dynamics of Brassica rapa genotypes using a new scanner-based phenotyping system. Ph.D. Thesis. University of Nottingham: U.K.
- Adu, M. O., Chatot, A., Wiesel, L., Bennett, M. J., Broadley, M. R., White, P. J. and Dupuy, L. X. (2014). A scanner system for high-resolution quantification of variation in root growth dynamics of Brassica rapa genotypes. J Exp Bot 65(8): 2039-2048.
- Lobet, G., Pages, L. and Draye, X. (2011). A novel image-analysis toolbox enabling quantitative analysis of root system architecture. Plant Physiol 157(1): 29-39.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Adu, M. O., Wiesel, L., Bennett, M. J., Broadley, M. R., White, P. J. and Dupuy, L. X. (2015). Scanner-based Time-lapse Root Phenotyping. Bio-protocol 5(6): e1424. DOI: 10.21769/BioProtoc.1424.
Category
Plant Science > Plant cell biology > Tissue analysis
Plant Science > Plant developmental biology > Morphogenesis
Plant Science > Plant physiology > Plant growth
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









