- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Enzymatic Activity Assays in Yeast Cell Extracts
Published: Vol 4, Iss 23, Dec 5, 2014 DOI: 10.21769/BioProtoc.1312 Views: 11612
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
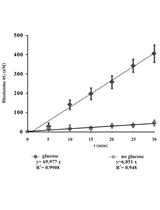

Measurement of Energy-dependent Rhodamine 6G Efflux in Yeast Species
Yvetta Gbelska [...] Alexandra Konecna
Aug 5, 2017 8952 Views
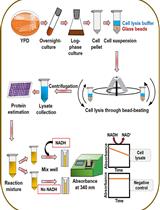
An Assay to Determine NAD(P)H: Quinone Oxidoreductase Activity in Cell Extracts from Candida glabrata
Anamika Battu [...] Rupinder Kaur
Nov 5, 2021 3054 Views
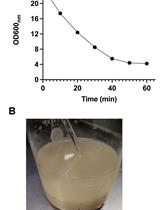
Isolation of Mitochondria from Ustilago maydis Protoplasts
Juan Pablo Pardo [...] Lucero Romero-Aguilar
Jan 5, 2022 3578 Views
Abstract
Saccharomyces cerevisiae (S. cerevisiae) (commonly known as baker’s yeast) is a model organism that has a similar upstream base excision repair (BER) pathway for the repair of methylated bases as that in mammalian cells, and it is very easy to maintain in the laboratory environment. Here, we described a method to prepare cell extracts from yeast to investigate their enzymatic activities. This protocol is a quick and efficient way to make yeast cell extracts without using commercial kits.
Keywords: DNA damageMaterials and Reagents
- EDTA-free protease inhibitor cocktail tablet (Roche Diagnostics, catalog number: 11836170001 )
- Bio-Rad protein assay dye reagent concentrate (Bio-Rad Laboratories, catalog number: 500-0006 )
- Dextrose (Sigma-Aldrich, catalog number: D9434 )
- Bacto-peptone (Sigma-Aldrich, catalog number: P5905 )
- Bacto-yeast extract (Sigma-Aldrich, catalog number: Y1625 )
- Adenine sulfate (Sigma-Aldrich, catalog number: A3159 )
- Trizma base (Sigma-Aldrich, catalog number: T1503 )
- EDTA (Sigma-Aldrich, catalog number: 93283 )
- NaCl (Sigma-Aldrich, catalog number: S7653 )
- β-mercaptoethanol (Sigma-Aldrich, catalog number: B0126 )
- Formamide (Sigma-Aldrich, catalog number: F9037 )
- Bromophenol blue (Sigma-Aldrich, catalog number: B0126)
- Xylene cyanol (Sigma-Aldrich, catalog number: X4126 )
- Dithiothreitol-DTT (Sigma-Aldrich, catalog number: D0632 )
- Sodium borohydride-NaBH4 (Sigma-Aldrich, catalog number: 247677 )
- Sterile water
- Yeast peptone dextrose adenine (YPDA) medium (see Recipes)
- Potassium phosphate buffer (see Recipes)
- Lysis buffer (see Recipes)
- 10x reaction buffer (see Recipes)
- Gel loading buffer (see Recipes)
Equipment
- 0.5 mm diameter glass beads (Bio Spec Products, catalog number: 11079105 )
- Eppendorf tubes
- Table-top centrifuge
- Table-top heat block
Procedure
- Preparation of yeast cell extracts
- The cell extracts from yeast Saccharomyces cerevisiae were prepared as described (Wang et al., 1995; Jazwinski, 1990) with modifications as below.
- Grow yeast culture in yeast peptone dextrose adenine (YPDA) medium at 30 °C overnight (O/N) with shaking.
- Prepare a fresh cell culture by diluting 10-fold from O/N culture in 50 ml YPDA medium.
- Grow the cells to stationary phase at 30 °C with vigorous shaking.
- When the culture reached an absorbance of 0.5-1.0 at 600 nm, centrifuge cells at 2,000 rpm (300 rcf) for 5 min at 4 °C.
- Wash the cell pellet with water and then with potassium phosphate buffer.
- Centrifuge cells again at 2,000 rpm (300 rcf) for 5 min at 4 °C.
- Resuspend the cell pellet in 500 μl of ice-cold lysis buffer.
- Transfer the suspension to an Eppendorf tube prefilled with pre-chilled beads (1:1 ratio of beads to lysate), and keep on ice.
- Disrupt the cells at 4 °C by vortexing the bead/lysate mixture: 10 cycles of 2 min vortex at maximum speed alternating with cycles of 1 min cooling on ice.
- Centrifuge the bead/lysate mixture at full speed at 4 °C for 15 min to remove cell debris.
- Transfer the supernatant fraction to a fresh Eppendorf tube. We recommend not collecting the entire supernatant fraction, instead leaving part of it to prevent touching pellet.
- Determine the protein concentration of the extract prepared by Bradford assay using dye reagent.
- The cell extracts from yeast Saccharomyces cerevisiae were prepared as described (Wang et al., 1995; Jazwinski, 1990) with modifications as below.
- Enzymatic activity assays in yeast cell extracts
- Prepare 10 μl of reaction mixture including 1x reaction buffer and 140 nM DNA substrate. DNA substrate used includes an adenylated uracil base at the 5′ end of the 3′-FAM–labeled oligonucleotide (Caglayan et al., 2014).
- Start the reaction by addition of yeast cell extract prepared as above (to a final amount of 50 μg) to the reaction mixture.
- Incubate the reaction mixture at 30 °C for 5, 15, and 30 min.
- Stabilize the reaction products by addition of 1 M freshly prepared and ice-cold NaBH4 to the final concentration of 100 mM.
- Incubate the reaction samples on ice for 30 min.
- Mix the reaction products with 10 μl of gel loading buffer.
- The reaction products in the yeast cell extracts were separated on a 15% polyacrylamide gel, the gel was scanned and the data were analyzed as reported (Caglayan et al., 2014). The result image for dRP lyase, FEN1 excision and DNA deadenylation enzymatic activities in yeast extracts has been reported (Caglayan et al., 2014).
- Prepare 10 μl of reaction mixture including 1x reaction buffer and 140 nM DNA substrate. DNA substrate used includes an adenylated uracil base at the 5′ end of the 3′-FAM–labeled oligonucleotide (Caglayan et al., 2014).
Representative data
The result image that shows dRP lyase, FEN1 excision and DNA deadenylation enzymatic activities in yeast extracts was published as Supplementary Figure 5 in Caglayan et al. (2014).
Notes
Enzymatic activity in yeast cell extracts may be lost after continuous vortexing if samples warm up. It might be necessary to extend the alternating cooling steps depending on the sample.
Recipes
- Yeast peptone dextrose adenine (YPDA) medium (1 L)
20 g dextrose
20 g Bacto-peptone
10 g Bacto-yeast extract
2 ml 0.5% adenine sulfate - Potassium phosphate buffer
25 mM KPO4 (pH 8.0) - Lysis buffer
25 mM Tris-HCl (pH 7.5)
1 mM EDTA
100 mM NaCl
10 mM β-mercaptoethanol
A protease inhibitor tablet - 10x reaction buffer
500 mM HEPES (pH 7.5)
100 mM MgCl2
200 mM KCl
5 mM EDTA
20 mM DTT - Gel loading buffer
95% formamide
20 mM EDTA
0.02% bromophenol blue
0.02% xylene cyanol
Acknowledgments
This work was supported by the Intramural Research Program of the US National Institutes of Health, National Institute of Environmental Health Sciences (grants Z01 ES050158 and ES050159).
References
- Caglayan, M., Batra, V. K., Sassa, A., Prasad, R. and Wilson, S. H. (2014). Role of polymerase beta in complementing aprataxin deficiency during abasic-site base excision repair. Nat Struct Mol Biol 21(5): 497-499.
- Jazwinski, S. M. (1990). Preparation of extracts from yeast. Methods Enzymol 182: 154-174.
- Wang, Z., Wu, X. and Friedberg, E. C. (1995). The detection and measurement of base and nucleotide excision repair in cell-free extracts of the yeast Saccharomyces cerevisiae. Methods 7(2): 177-186.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Çağlayan, M. and Wilson, S. H. (2014). Enzymatic Activity Assays in Yeast Cell Extracts. Bio-protocol 4(23): e1312. DOI: 10.21769/BioProtoc.1312.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Molecular Biology > DNA > DNA damage and repair
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










