- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Rifampicin-resistance Mutation Frequency and Analysis of Mutation Spectra in Mycobacteria
Published: Vol 4, Iss 13, Jul 5, 2014 DOI: 10.21769/BioProtoc.1168 Views: 12387
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
A Novel Method of Inducible Directed Evolution to Evolve Complex Phenotypes
Ibrahim S. Al’Abri [...] Nathan Crook
Oct 20, 2022 3062 Views

Comprehensive Mapping of EZ-Tn5 Transposon Insertion Sites in Pseudomonas argentinensis SA190 Using RATE-PCR
Büsra Elkatmis [...] Maged M. Saad
Jul 20, 2025 1751 Views
Editing the Serratia proteamaculans Genome Using the Allelic Exchange Method
Ksenia Chukhontseva [...] Ilya Demidyuk
Sep 20, 2025 1387 Views
Abstract
Understanding the genetic safeguarding mechanism of Mycobacterium tuberculosis (Mtb) may help us to explain i), how Mtb survive the genetic assaults elicited by both reactive oxygen species (ROS) and reactive nitrogen species (RNS) produced by host macrophages and ii), why some strains of Mtb, e.g., Mtb strains from East Asian lineage and Beijing sublineage, exhibit high mutation rate and are more likely to acquire drug resistant mutations (e.g., rifampicin-resistance mutation) during infection. Mutation frequency analysis is a basic methods to study the genetic safeguarding mechanism. Moreover, to study the molecular mechanism of mutation, it is necessary to analyse the mutation spectrum (For example, oxidized cytosine may induce CG to TA mutation.). This protocol describes a method to determine the mutation frequency and understand the mutation spectrum in both Mycobacterium smegmatis (Msm) and Mtb.
Materials and Reagents
- Mycobacterium smegmatis mc2 155 or Mycobacterium tuberculosis H37Rv
- Middlebrook 7H9 broth (BD Biosciences, catalog number: 271310 )
- Middlebrook 7H11 agar (BD Biosciences, catalog number: 283810 )
- Middlebrook OADC (BD Biosciences, catalog number: 212351 )
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- Tween 80 (Sigma-Aldrich, catalog number: P1754 )
- Rifampicin (Sigma-Aldrich, catalog number: R3501 )
- 7H9 medium (see Recipes)
- 7H9OADC medium (see Recipes)
- 7H11 agar (see Recipes)
- 7H11OADC agar (see Recipes)
- LBG agar (see Recipes)
- PBST (see Recipes)
- TE buffer (see Recipes)
Equipment
- Incubation shaker
- Centrifuge
- 37 °C incubator
- 100-ml flask
- Sterile glass beads
- 96-well flat bottom plate
- Adhesive film
- 96-well PCR plate
- PCR instrument
- Petri dish (70 mm diameter)
Software
- BLAST or ClustalW
Procedure
- Determination of rifampicin-resistance mutation frequency
- Streak Mycobacterium smegmatis culture on 7H11 agar plate (does not contain rifampicin) and culture at 37 °C for 3 days until colonies are visible.
For Mtb, streak it on 7H11OADC agar plate and culture at 37 °C for 4 weeks.
- Inoculate a single colony in 5 ml 7H9 medium (for Mtb, use 7H9OADC medium) and grow at 37 °C with shaking (150 rpm) until the culture is saturated (OD600~2.0).
- The culture is sub-cultured (1% v/v inoculation) into 20 ml 7H9 medium (for Mtb, use 7H9OADC medium) in a 100-ml flask and grown at 37 °C with shaking to exponential phase (OD600=0.4~0.5).
Note: Culture at this growth stage can be used for stress induction, e.g., oxidative stress (H2O2 treatment) or starvation.
- For CFU determination, the culture is serially diluted 1: 100,000 in PBST. 50 μl of the 10-5 dilution is plated on LBG agar plate (for Mtb, use 7H11OADC agar plate). Incubate the plates at 37 °C for 4 days. CFU/ml = (the number of colonies on each plate) x 2 x 106.
Note: At least three independent dilutions are required for each sample.
- 3 ml of the cell culture from step 3 is centrifuged at 3,000 x g for 5 min and the cell pellet is resuspended in 100 μl PBST (The bacilli may form clumps in liquids without Tween 80.). To isolate rifampicin-resistant mutants, the resuspended pellet is spread with sterile glass beads on LBG agar plates containing 250 μg/ml rifampicin and incubated at 37 °C for 4 days.
For Mtb, the resuspended cell pellet is spread on 7H11OADC agar plate containing 10 μg/ml rifampicin and incubate at 37 °C for 4 weeks.
Note: Glass or metal spreader is not recommended in this step because the mycobacterial cell may adsorb and clump onto the spreader during plating, which may affect the accuracy of the result.
- The spontaneous rifampicin-resistant mutation frequency is calculated by dividing the number of rifampicin-resistant colonies on each plate by the total viable cell count per plate (Figure 1).
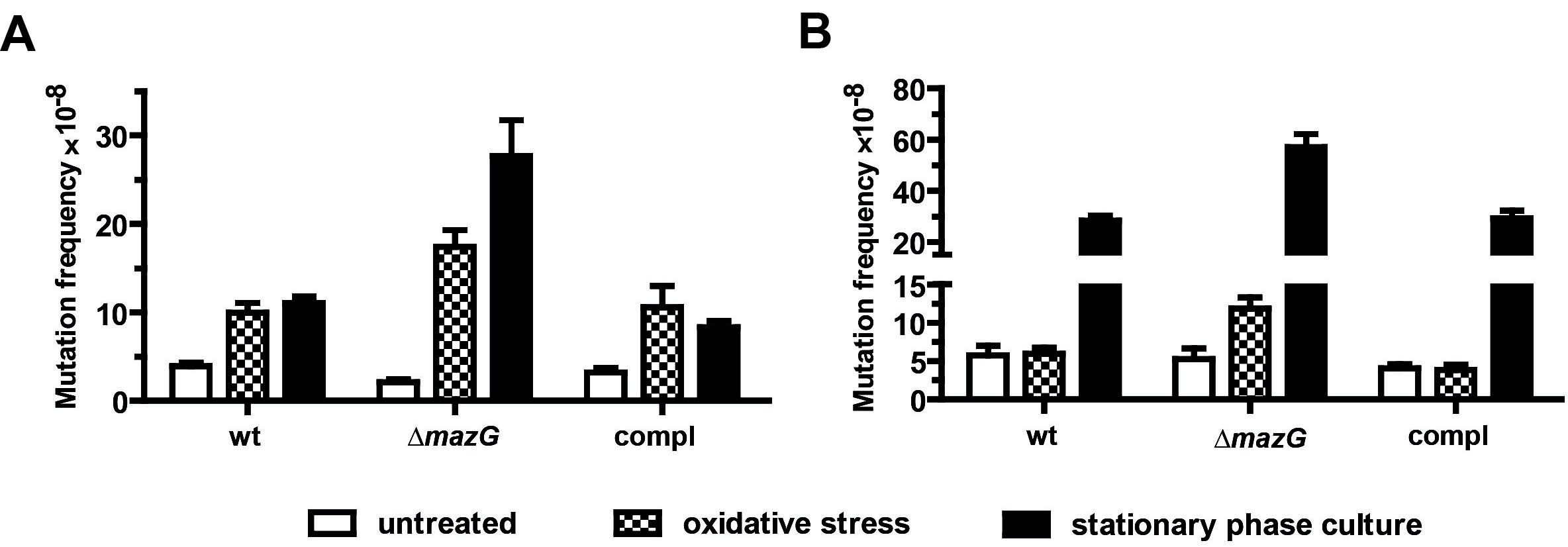
Figure 1. The antimutator role of MazG in Msm (A) and Mtb (B). The frequencies conferring resistance to rifampicin in wild-type (wt), mazG-null (ΔmazG) and the complemented mutant (compl) strains were determined in exponential phase (OD600~0.5) with or without oxidative stress and in the stationary growth phase. Oxidative stress was induced by treating exponential phase cultures with 10 mM H2O2 for 5 h (Msm) or 24 h (Mtb). Stationary phase was at the 5th-day or 28th-day of culture for Msm or Mtb, respectively. The numbers shown are mean ± S.E. of 3 independent experiments totalling 15 cultures of Msm and 6 of Mtb.
- Streak Mycobacterium smegmatis culture on 7H11 agar plate (does not contain rifampicin) and culture at 37 °C for 3 days until colonies are visible.
- Mutation spectrum analysis
- A single rifampicin-resistant Msm colony is inoculated into 1 ml 7H9 media containing 100 μg/ml rifampicin in a 96-well flat bottom plate. The plate is then sealed with an adhesive film and incubated at 37 °C for 7 days. Note that at least 30 colonies are required per bacterial sample for statistically significant results.
To inoculate rifampicin-resistant Mtb colony, use 7H9OADC with 4 μg/ml rifampicin, incubate at 37 °C for 3 weeks.
- Cells are then pelleted (4,000 x g, 5 min) and suspended in 50 μl TE buffer and the cell suspension is transferred to a 96-well PCR plate.
- The plate is heated in a PCR instrument at 95 °C for 20 min.
- The PCR plate is then centrifuged at 12,000 x g for 5 min and the supernatant is transferred to a new 96-well PCR plate.
- A region of the genome sequence containing the cluster I region of rpoB is amplified using primers Rpo-forward (5’-CGACCACTTCGGCAACCG-3’) and Rpo-reverse (5’-CGATCAGACCGATGTTGG-3’). For Mtb, using primer RpoTB-forward (5’-ATCACACCGCAGACGTTG-3’) and RpoTB-reverse (5’-TGCATCACAGTGATGTAGTCG-3’).
PCR mix: 2 μl DNA template, 0.4 ul of each primer (10 μM), 1 U Pfu DNA polymerase, 0.6 μl DMSO, 0.4 μl dNTPs (10 mM each), H2O to 20 μl.
PCR cycling: 95 °C 60 s, 30 cycles of 95 °C 20 s, 58 °C 10 s and 72 °C 40 s.
Note: Most of the rifampicin-resistance mutations are occurred in the cluster I region of rpoB. If no mutation is detected in this region, it may need to amplify the whole rpoB gene (~3.5 Kbp) for sequencing.
- The PCR products are then bi-directionally sequenced and the mutation spectrum of the sequenced region is identified by BLAST or ClustalW.
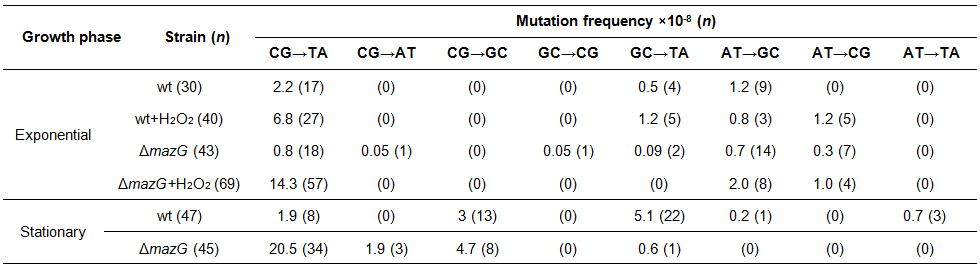
- Calculate the mutation frequency of each specific mutation spectrum (Table 1).
Mutation frequency of a given mutation spectrum = (the proportion of the mutation spectrum in strain A) x (the mutation frequency of strain A). For example, the mutation frequency of Msm is 10 x 10-8, the proportion of CG-TA mutation in Msm derived rifampicin-resistant mutants is 80%, therefore, the CG-TA mutation frequency will be 0.8 x 10 x 10-8=8 x 10-8.
Table 1. mazG-null Msm exhibited elevated CG to TA mutation under oxidative stress conditions and in stationary phase. Spontaneous rifampicin-resistant colonies were collected from 3 independent experiments . Cluster I region of rpoB were PCR-amplified using Pfu DNA polymerase and sequenced bi-directionally. All of the sequenced colonies contain single non-synonymous mutations. wt, wild-type Msm; ΔmazG, mazG-null Msm.
- A single rifampicin-resistant Msm colony is inoculated into 1 ml 7H9 media containing 100 μg/ml rifampicin in a 96-well flat bottom plate. The plate is then sealed with an adhesive film and incubated at 37 °C for 7 days. Note that at least 30 colonies are required per bacterial sample for statistically significant results.
Recipes
- 7H9 medium (100 ml)
Dissolve 0.47 g of 7H9 broth in 80 ml dH2O
Add 0.4 ml 50% glycerol and 0.5 ml 10% Tween 80
dH2O to 100 ml
Sterilized by autoclaving
Stored at 4 °C
- 7H9OADC medium (100 ml)
Dissolve 0.47 g of 7H9 broth in 80 ml dH2O
Add 0.4 ml 50% glycerol and 0.5 ml 10% Tween 80
dH2O to 90 ml
Sterilized by autoclaving
Cool to 50-55 °C and add 10 ml OADC
Stored at 4 °C
- 7H11 agar (100 ml)
Dissolve 2.1 g of 7H11 agar with 99 ml dH2O
Add 1 ml 50% glycerol
Sterilized by autoclaving
Stored at 4 °C
- 7H11OADC agar (100 ml)
Dissolve 2.1 g of 7H11 agar with 89 ml dH2O
Add 1 ml 50% glycerol
Sterilize by autoclaving
Cool to 50-55 °C and add 10 ml OADC
Stored at 4 °C
- LBG agar (100 ml)
Dissolve 1 g tryptone, 0.5 g yeast extract, 1 g NaCl and 1.5 g agar with 99 ml dH2O
Add 1 ml 50% glycerol
Sterilized by autoclaving
Stored at 4 °C
- PBST
Dissolve the following in 800 ml of dH2O
8 g of NaCl
0.2 g of KCl
1.44 g of Na2HPO4
0.24 g of KH2PO4
5 ml of 10% Tween 80
Adjust pH to 7.2
Adjust volume to 1 L with dH2O
Sterilized by autoclaving
- TE buffer
10 mM Tris-HCl (pH 8.0)
1 mM EDTA
Acknowledgments
This protocol was adapted from a previously published paper Lyu et al. (2013). This work was supported by grants from the National Natural Science Foundation of China (No. 30970077, 31121001, 31300126), the Research Unit Fund of Li Ka Shing Institute of Health Sciences (No. 7103506), the Hong Kong Health and Medical Research Fund (No. 12110622), the China Postdoctoral Science Foundation (No. 20110490754), the SIBS Postdoctoral Research Fund (No. 2011KIP509).
References
- Lyu, L. D., Tang, B. K., Fan, X. Y., Ma, H. and Zhao, G. P. (2013). Mycobacterial MazG safeguards genetic stability via housecleaning of 5-OH-dCTP. PLoS Pathog 9(12): e1003814.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Lyu, L. and Zhao, G. (2014). Determination of Rifampicin-resistance Mutation Frequency and Analysis of Mutation Spectra in Mycobacteria. Bio-protocol 4(13): e1168. DOI: 10.21769/BioProtoc.1168.
- Lyu, L. D., Tang, B. K., Fan, X. Y., Ma, H. and Zhao, G. P. (2013). Mycobacterial MazG safeguards genetic stability via housecleaning of 5-OH-dCTP. PLoS Pathog 9(12): e1003814.
Category
Microbiology > Microbial genetics > Mutagenesis
Microbiology > Microbial cell biology > Cell isolation and culture
Molecular Biology > DNA > Mutagenesis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link