Advanced Search
Overcoming Autofluorescence to Assess GFP Expression During Normal Physiology and Aging in Caenorhabditis elegans
Published: Jul 20, 2018 DOI: 10.21769/BioProtoc.2940 Views: 11613
Abstract
Green fluorescent protein (GFP) is widely used as a molecular tool to assess protein expression and localization. In C. elegans, the signal from weakly expressed GFP fusion proteins is masked by autofluorescence emitted from the intestinal lysosome-related gut granules. For instance, the GFP fluorescence from SKN-1 transcription factor fused to GFP is barely visible with common GFP (FITC) filter setups. Furthermore, this intestinal autofluorescence increases upon heat stress, oxidative stress (sodium azide), and during aging, thereby masking GFP expression even from proximal tissues. Here, we describe a triple band GFP filter setup that separates the GFP signal from autofluorescence, displaying GFP in green and autofluorescence in yellow. In addition, yellow fluorescent protein (YFP) remains distinguishable from both the yellowish autofluorescence and GFP with this triple band filter setup. Although some GFP intensity might be lost with the triple band GFP filter setup, the advantage is that no modification of currently used transgenic GFP lines is needed and these GFP filters are easy to install. Hence, by using this triple band GFP filter setup, the investigators can easily distinguish autofluorescence from GFP and YFP in their favorite transgenic C. elegans lines.
Keywords: MicroscopyBackground
Major sources of autofluorescence include intracellular lysosome-derived granules, mitochondria (i.e., autofluorescent molecules such as NAD(P)H and flavins), or extracellular collagen (Hermann et al., 2005; Monici, 2005). During aging, autofluorescent materials such as lipofuscin and advanced glycation end-products (AGE) accumulate. In the nematode C. elegans, the autofluorescence of gut granules starts already during embryogenesis, reflecting the biogenesis of lysosome-related organelles (Hermann et al., 2005). This prominent autofluorescence of these lysosome-related gut granules in the intestine continues throughout development and adulthood. The source of the autofluorescence, whether it is lipofuscin, AGE, tryptophan metabolites, or something else, is still unclear. However, this autofluorescence increases during aging and the two main tissues that show the highest autofluorescence are the intestine and the uterus in C. elegans (Pincus et al., 2016). With current fluorescent filter sets (TRITC, DAPI, FITC), three different autofluorescent wavelengths have been characterized in C. elegans. The red autofluorescence (visualized by TRITC) progressively increases with age, the blue autofluorescence (visualized by DAPI) peaks right before death, and the green autofluorescence (visualized by FITC) is a mixture from the red and blue autofluorescence (Pincus et al., 2016).
The multicellular model organism C. elegans is transparent, allowing GFP fluorescence to be assessed in vivo non-invasively (Chalfie et al., 1994). With commonly used GFP filter sets, for instance, FITC with an excitation center wavelength of 470 nm and a full bandwidth of 40 nm (470/40 nm), and emission range of 525/50 nm, the intestinal autofluorescence overlaps with the GFP signal. Previously, knockdowns by RNA interference (RNAi; e.g., tdo-2 RNAi) or gut granule-loss (glo) mutations (Hermann et al., 2005; Coburn et al., 2013), which either diminish or eliminate the intestinal autofluorescence, have been applied to the desired GFP transgenic C. elegans lines to overcome this problem. However, RNAi knockdowns or mutations that help to diminish autofluorescence alter gene function and might cause artifacts. In addition, there are transgenic GFP fusions of several stress response-regulating transcription factors (DAF-16::GFP, HSF-1::GFP, HLH-30::GFP, SKN-1::GFP) that are routinely used to assess cytoplasmic to nuclear translocation in intestinal cells as a proxy for their activation (Henderson and Johnson, 2001; Libina et al., 2003; Kwon et al., 2010; Lapierre et al., 2013; Morton and Lamitina, 2013; Ewald et al., 2015 and 2017b). Particularly, the transgenic SKN-1::GFP fusion is barely visible and is masked by intestinal autofluorescence even in larval C. elegans (Havermann et al., 2014; Wang et al., 2016; Hu et al., 2017).
To overcome the problem of autofluorescence masking intestinal GFP, Oliver Hobert (http://www.bio.net/mm/celegans/1998-November/001769.html) and several other investigators in the C. elegans community had proposed the principle of this combination of GFP filter sets. Optimization of these GFP filter sets by the Blackwell lab made it possible to assess the subcellular localization of SKN-1 and other proteins that were difficult to visualize (An and Blackwell, 2003). Unfortunately, these previous filter sets are not on sale anymore. Here, we describe the currently and commercially available filters that can be used to rebuild these GFP-filter settings. In contrast to the single band FITC GFP filter set, the proposed triple band GFP filter set has a very narrow excitation bandwidth of 10 nm, which is right by the maximum peak for the S65C mutant GFP excitation (488 nm) that is commonly used in C. elegans (Boulin et al., 2006; Heppert et al., 2016). More importantly, the emission filter used here has a first pass-through (520/20 nm) for the light emitted close to the GFP emission peak (509 nm) and a second pass-through (595/40 nm) from the light around the autofluorescence emission, allowing the separation of GFP (visible in green) and autofluorescence (visible in yellow).
Materials and Reagents
- 250 ml glass Erlenmeyer flask
- 1.5 ml centrifuge tubes
- Microscope slides (size: 76 mm x 26 mm, 1 mm thick; VWR, Thermo Fisher Scientific, catalog number: 631-1303 )
- Cover slip (size: 18 mm x 18 mm; VWR, catalog number: 631-1567 )
- Tape (MILIAN, catalog numbers: 140255B , BA-5419-07 )
- C. elegans strains (available at Caenorhabditis Genetics Center [CGC] https://cbs.umn.edu/cgc/home) or if not available at CGC, can be requested directly from the research labs that generated them: N2 C. elegans wild-type Bristol, LD1 Is007 [Pskn-1::skn-1b/c::gfp; pRF4 rol-6 (su1006)], LSD2022 spe-9(hc88); jgIs5 [Prol-6::rol-6::gfp, Pttx-3::gfp], EQ87 iqIs28 [pAH71 Phsf-1::hsf-1::gfp; pRF4 rol-6 (su1006)], BT24 rhIs23 [gfp::him-4] III, NL5901 pkIs2386 [Punc-54::alpha-synuclein::YFP + unc-119(+)])
Note: For culturing and handling C. elegans, please see Stiernagle (2006). - Agarose (Conda, catalog number: 8010 )
- KH2PO4 (Merck, catalog number: 1048731000 )
- Na2HPO4 (Sigma-Aldrich, catalog number: S5136 )
- NaCl (Sigma-Aldrich, catalog number: S3014 )
- MgSO4 (Fisher Scientific, catalog number: 10316240 )
- Levamisole hydrochloride (Sigma-Aldrich, catalog number: L0380000 ) (2 mM) solved in M9 buffer
Note: Used here to paralyze worms; it is not recommended to use sodium azide (NaN3) (Sigma-Aldrich, catalog number: S2002-100G ) (20 mM, solved in M9 buffer). - Agarose pads (see Recipes)
- M9 buffer (see Recipes) (Stiernagle, 2006; He, 2011)
Equipment
- Autoclave
- Microwave to heat up agarose
- Heated water bath or heat block to keep agarose molten
- For loading C. elegans onto agarose pads:
Stereomicroscope, worm pick, pipettes (Figure 1A) (Ewald et al., 2017a) - Upright bright field fluorescence microscope (Tritech Research, model: BX-51-F , Figure 1B)
- Camera (The Imaging Source, model: DFK 23UX236 , with IC Capture 2.4 software)
It is important to use a color camera, since a monochrome camera is unable to distinguish between colors, so the filter set would be ineffective. - Triple band filter sets
Note: The triple band filter sets used here are from Chroma Technology Corp., but similar filter sets can be acquired from other manufacturers.
The triple band filter sets consist of the 69000 ET-DAPI/FITC/TRITC (69000x, 69000m, 69000bs, EX/EM 25 mm, ringed, DC 25.5 x 36 x 1 mm) (Chroma Technology, catalog number: 69000 ). However, the excitation filter 69000x is exchanged with an ET485/10x narrow band excitation filter (25 mm, ringed; Chroma Technology, catalog number: ET485/10x). The interpretation of the filter nomenclature for example for ET485/10x is: “ET” stands for magnetron sputtered exciter, “485” indicates the center wavelength of 485 nm and the “/10” indicates the full bandwidth of 10 nm (i.e., +/- 5 nm from the center), and the “x” stands for excitation. Hence, the triple band filter set consists of the ET485/10x excitation filter, the 69000bs dichroic beam splitter, and the 69000m emission filter. The triple band filters are then assembled in a microscope filter cube. A schematic of this setup is shown in Figures 1C and 1D. A comparison between the filter set properties and the resulting images are shown in Figure 2. In brief, the 69000m emission filter allows light coming through from 520/20 nm (green) and from 595/40 (yellow to orange/red) but blocks the greenish to yellow light (535-572 nm), which is the key feature of the triple band filter set that allows distinguishing GFP from autofluorescence (Figure 2). This is in contrast to a GFP long-pass emission filter (ET500lp, > 500 nm), which allows all the light from green to red to pass through.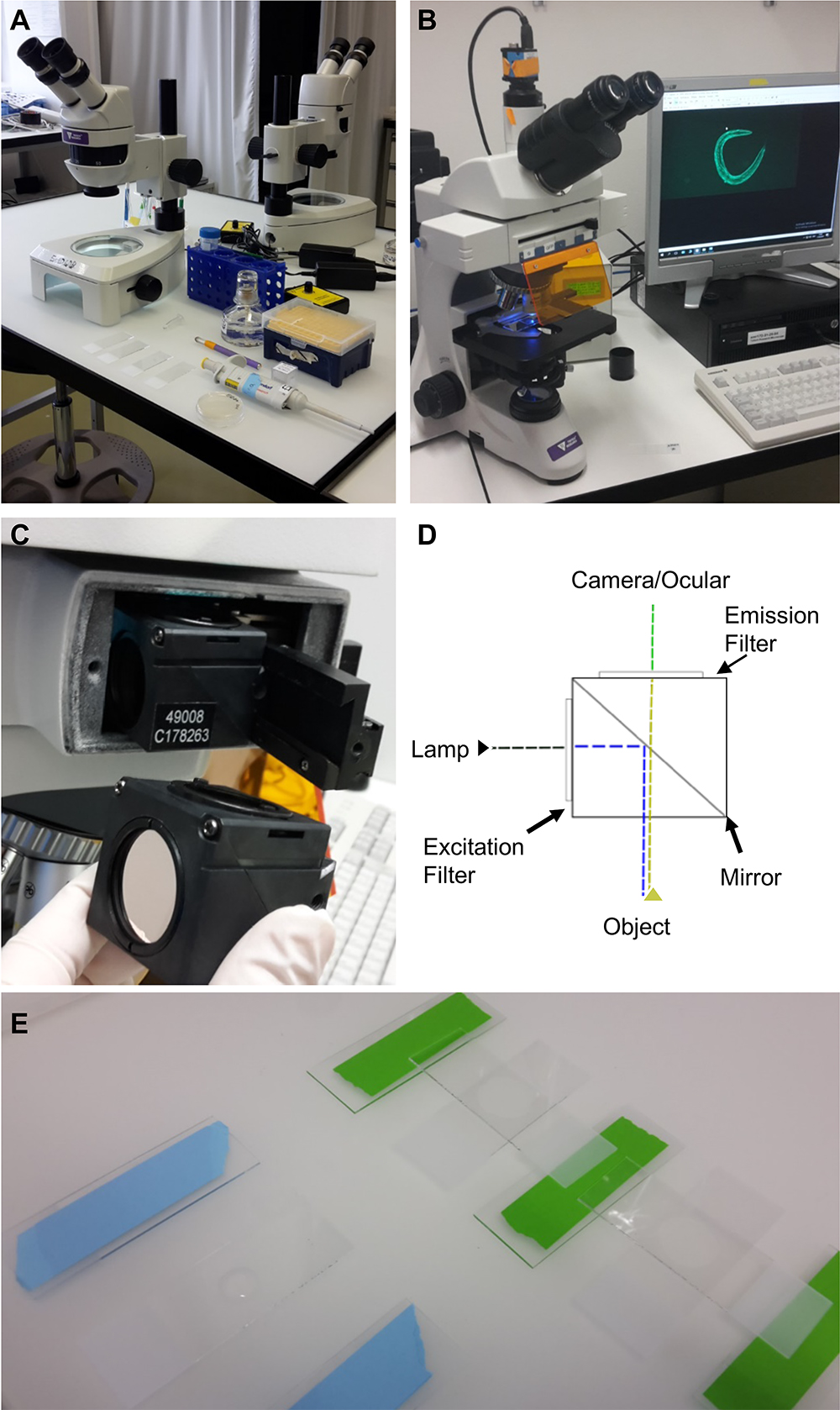
Figure 1. Equipment and experimental setup. A. Equipment and utilities for mounting C. elegans on microscope slides. Shown from the top left: a stereoscope, M9 buffer in a falcon tube, levamisole in a 1.5 ml centrifuge tube, a pipette with tips, a worm-pick to mount C. elegans in the liquid droplet, microscope slides with 2% agarose pad, and C. elegans on culturing plate. B. Upright bright field fluorescence microscope setup; C. The filter cube and its position in the microscope; D. A schematic representation of the filter cube. The dashed lines indicate the pathway of the light through the filter cube, while the arrows indicate the filters and the mirror. E. Preparation of the 2% agarose pad slides for microscopy. On the left side is a slide with a drop of 2% agarose dropped between two blue taped slides, while on the right side the agarose drop was already covered by another slide perpendicular to the green taped slides.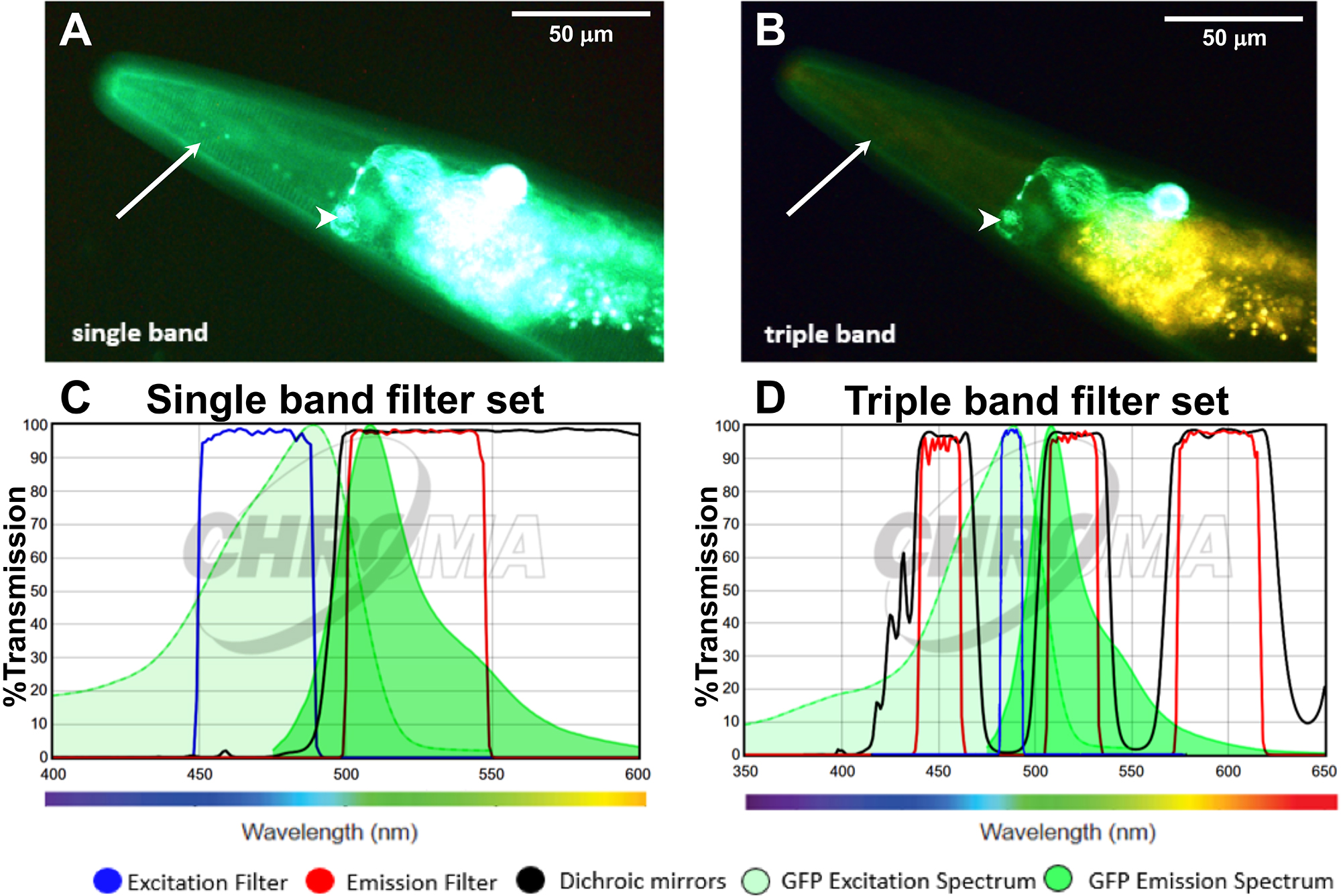
Figure 2. Applying the triple band filter set to distinguish between GFP signal and C. elegans autofluorescence. A-B. The LSD2022 C. elegans strain expresses an integrated collagen::GFP transgene (ROL-6::GFP), which is visible in the cuticle (white arrow). In addition, the strain LSD2022 expresses GFP driven by the ttx-3 promoter in the AIY interneuron pair (white arrowhead) (Kim et al., 2010). A C. elegans worm (LSD2022) imaged with a commonly used single band filter set (A). The same animal imaged with the triple band filter set. Green is GFP and yellow is autofluorescence (B). C. Transmission graph of the single band filter set we used in (A) [49002-ET-EGFP (FITC/Cy2) by Chroma]; D. Transmission graph of the triple band filter setup used for (B). The triple band filter setup consists of a narrow band ET485/10x excitation filter, a 69000bs dichroic mirror filter, and a 69000m emission filter, which allows light coming through from 520/20 nm (green) and from 595/40 (yellow to orange/red). However, the 69000m emission filter blocks the greenish to yellow light (535-572 nm), which is the key feature that helps to distinguish GFP from autofluorescence. (C and D) The graphs are both adapted from www.chroma.com.
Software
- IC Capture 2.4 software (https://www.theimagingsource.com/support/downloads-for-windows/end-user-software/iccapture/)
- ImageJ (https://imagej.net/Image_Stitching)
Procedure
Category
Developmental Biology > Cell signaling > Stress response
Cell Biology > Cell imaging > Fluorescence
Molecular Biology > Protein > Detection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link


