- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Expression, Purification and Crystallization of Recombinant Arabidopsis Monogalactosyldiacylglycerol Synthase (MGD1)
Published: Vol 6, Iss 24, Dec 20, 2016 DOI: 10.21769/BioProtoc.2064 Views: 8813
Reviewed by: Arsalan DaudiAdam IdoineAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
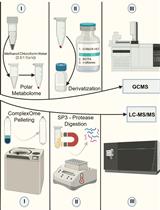
Streamlining Protein Fractional Synthesis Rates Using SP3 Beads and Stable Isotope Mass Spectrometry: A Case Study on the Plant Ribosome
Dione Gentry-Torfer [...] Federico Martinez-Seidel
May 5, 2024 2861 Views
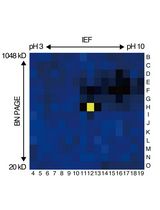

An Activity-Based Proteomics with Two-Dimensional Polyacrylamide Gel Electrophoresis (2D-PAGE) for Identifying Target Proteases in Arabidopsis Apoplastic Fluid
Sayaka Matsui and Yoshikatsu Matsubayashi
Mar 5, 2025 1983 Views
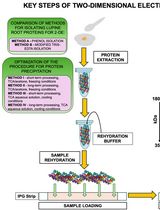
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1768 Views
Abstract
In plant cells, galactolipids are predominant, representing up to 50% of the lipid content in photosynthetic tissues. Galactolipid synthesis is initiated by MGDG synthases (MGDs), which use UDP-galactose as a donor sugar and diacylglycerol (DAG) as acceptor, to form monogalactosyldiacylglycerol (MGDG). This protocol is used to produce a recombinant form of Arabidopsis thaliana (A. thaliana) monogalactosyldiacylglycerol synthase 1 (MGD1) protein, in Escherichia coli (E. coli), using a two-step chromatographic purification procedure. The protein is easily expressed and purified to milligram quantities, suitable for biochemical and structural studies. The crystallization of MGD1 is also described.
Keywords: Photosynthetic tissuesBackground
Previous attempts to express plant MGDs in E. coli showed that approximately 99% of the recombinant protein accumulated in inclusion bodies (Miège et al., 1999). Solubilization of bacterial membranes using detergents, or in vitro inclusion bodies refolding protocols were developed and yielded pure and active fractions, sufficient to monitor the activity of the enzyme, but not to pursue its structural study (Nishiyama et al., 2003; Botté et al., 2005). Using a combination of different biochemical and biophysical techniques, and investigating the effects of various buffers and additives on the biochemical behavior of the enzyme, a simple, efficient and fast protocol was developed for the expression and purification of recombinant MGD1, addressing the problems frequently encountered with the purification of glycosyltransferases, particularly protein aggregation (Rocha et al., 2013). Conditions detailed here allowed the unprecedented production of a pure, soluble and active form of MGD1 and comply with both structural and functional dissections of this enzyme (Rocha et al., 2016). The protocol here described can also serve as a starting strategy to purify similar proteins.
Materials and Reagents
- 50 ml Corning tubes (Corning, catalog number: 430829 )
- 1.5 ml Eppendorf tubes (Treff, catalog number: 96.07246.9.01 )
- 30 ml centrifuge tubes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 3138-0030 )
- 0.2 µm PES-membrane filters (Dominique Dutscher, catalog number: 051732 )
- Vivaspin 20, MWCO 30,000 Da concentrators (Sartorius, catalog number: VS2022 )
- HisTrap FF 1 ml column (GE Healthcare, catalog number: 17-5319-01 )
- Superdex S200 10/300 GL column (GE Healthcare, catalog number: 17-5175-01 )
- 24-well VDX plate with sealant (HAMPTON RESEARCH, catalog number: HR3-170 )
- 22 mm cover slips (HAMPTON RESEARCH, catalog number: HR3-233 )
- E. coli BL21(DE3) strain (New England Biolabs, catalog number: C2527I )
- Luria Broth (LB) media (Thermo Fisher Scientific, InvitrogenTM, catalog number: 12780-052 )
- Kanamycin (Euromedex, catalog number: EU0420 )
- Isopropyl-β-D-thiogalactopyranoside (IPTG) (Euromedex, catalog number: EU0008 )
- Antifoam 204 (Sigma-Aldrich, catalog number: A6426 )
- Benzonase® nuclease (Sigma-Aldrich, catalog number: E1014 )
- Imidazole (Sigma-Aldrich, catalog number: 56749 )
- Ethanol absolute anhydrous (CARLO ERBA Reagents, catalog number: 308607 )
- Magnesium chloride hexahydrate (MgCl2·6H2O) (Sigma-Aldrich, catalog number: M9272 )
- PEG 3350 (Sigma-Aldrich, catalog number: 88276 )
- Liquid nitrogen (LN2)
- Glycerol (Sigma-Aldrich, catalog number: G7757 )
- HEPES (Thermo Fisher Scientific, Fisher Scientific, catalog number: BP310 )
- Sodium chloride (NaCl) (Thermo Fisher Scientific, Fisher Scientific, catalog number: 10553515 )
- Urea (Sigma-Aldrich, catalog number: U1250 )
- Dithiothreitol (DTT) (Euromedex, catalog number: EU0006 )
- cOmpleteTM protease inhibitor cocktail (Roche Diagnostics, catalog number: 11873580001 )
- Bis-tris propane (Sigma-Aldrich, catalog number: B6755 )
- Tris(2-carboxyethyl)phosphine (TCEP) (Sigma-Aldrich, catalog number: C4706 )
- Tris base (Thermo Fisher Scientific, Fisher Scientific, catalog number: 10376743 )
- Sodium dodecyl sulfate (SDS) 10% (Euromedex, catalog number: EU0760 )
- Bromophenol blue (EMD Millipore, catalog number: 108122 )
- 2-mercaptoethanol (Sigma-Aldrich, catalog number: M3148 )
- Hydrochloric acid 37% (CARLO ERBA Reagents, catalog number: 403871 )
- Sodium hydroxide (EMD Millipore, catalog number: 1064621000 )
- Lysis buffer (see Recipes)
- Washing/binding buffer (see Recipes)
- Elution buffer (see Recipes)
- Size exclusion buffer (see Recipes)
- SDS-PAGE deposition buffer (5x DB) (see Recipes)
- 20% (v/v) ethanol (see Recipes)
- 6 N hydrochloric acid solution (for pH adjustment) (see Recipes)
- 10 M sodium hydroxide solution (for pH adjustment) (see Recipes)
- Precipitant solution (or mother liquor) (see Recipes)
Equipment
- Culture flasks (IDEA Conception, catalog number: ART0004104 )
Note: These are custom made 3 L culture flasks. - Orbital shakers at 30 °C and 37 °C
- Incubator at 20 °C (for crystallization plates)
- Sonication bath (room temperature)
- Lab balances
- Cooled centrifuges
- Filtration system or syringes
- Cell Disruptor CSL ‘One-shot’ (Constant Systems, Ltd)
- ÄKTAFPLC instrument (GE Healthcare, catalog number: 18190026 ) or equivalent
- SDS-PAGE equipment
- Zetasizer Nano ZSTM for Dynamic Light Scattering Analysis (Malvern Instruments, model: ZEN3600 ) or equivalent
- BioPhotometer 6131 (Eppendorf, Germany)
- NanoDropTM ND-2000 spectrophotometer (Thermo Fisher Scientific, Thermo ScientificTM, model: ND-2000 )
- Olympus Zoom Stereo microscope (Olympus, model: SZ1145 TR CTV ) or equivalent
Note: This product has been discontinued. - Olympus Transmitted Light Illumination Base (Olympus, model: SZX-ILLK200 ) or equivalent
Note: This product has been discontinued.
Note: Cryocrystallography material for crystals manipulation in liquid nitrogen (LN2)
- Litholoops several sizes (Molecular Dimensions)
- CryoCaps (Plain) (Molecular Dimensions, catalog number: MD7-400 )
- Magnetic CryoVials (Molecular Dimensions, catalog number: MD7-402 )
- EMBL/ESRF Sample changer starter kit (Molecular Dimensions, catalog number: MD7-500 )
- Magnetic Cryo Wand (Molecular Dimensions, catalog number: MD7-411 )
- LN2 foam dewars (Molecular Dimensions, catalog number: MD7-35 )
- Dry shipper (CX100) dewar (Molecular Dimensions, catalog number: MD7-21 )
Procedure
- Expression of MGD1△137-6His protein
The protein construction here described is a truncated form of MGD1 (MGD1△137-6His), deprived of its N-terminal 136 amino-acid residues and with a non-cleavable 6-histidine tag at the protein C-terminus. The deleted segment comprises the canonical N-terminal chloroplast signal peptide (res. 1-106; cleaved upon import to the organelle), and a 30-mer region predicted to be disordered, and not necessary for activity. The mgd1△137 DNA fragment was cloned at the NdeI/NotI sites of the pET29b vector (Novagen, 69872), generating the pET29b-MGD1△137-6His construct. E. coli BL21 (DE3) cells were used as host cells for protein production.- Prepare 1 L of sterile LB medium supplemented with kanamycin (30 µg/ml). Inoculate media with E. coli BL21 (DE3) cells harboring the pET29b-MGD1△137-6His construct. Incubate culture at 37 °C and 180 rpm.
- Monitor OD600nm of culture on a UV/VIS spectrophotometer.
- At OD600nm ~0.6-0.8, take 1 ml sample of culture for SDS-PAGE analysis (before induction - T0). Centrifuge (13,000 x g, 1 min), discard supernatant and resuspend pellet in 40 µl of distilled water and 10 µl of 5x DB. Boil samples for 5 min and sonicate for 2 min prior to gel loading.
- Induce cells with IPTG to a final concentration of 0.25 mM. Incubate 4 h at 30 °C, 180 rpm.
- After induction, take 1 ml sample of culture for SDS-PAGE analysis (after induction - T4). Centrifuge (13,000 x g, 1 min), discard supernatant and resuspend pellet in 80 µl of distilled water and 20 µl of 5x DB. Boil samples for 5 min and sonicate for 2 min prior to gel loading.
- Harvest cells by centrifugation (7,000 x g for 20 min at 4 °C).
- Discard supernatant and store the pellet in a previously weighed 50 ml Corning tube.
- Weigh wet cell pellet and store at -20 °C if not immediately used (typically ~4 g/L culture).
- Prepare 1 L of sterile LB medium supplemented with kanamycin (30 µg/ml). Inoculate media with E. coli BL21 (DE3) cells harboring the pET29b-MGD1△137-6His construct. Incubate culture at 37 °C and 180 rpm.
- IMAC purification of MGD1△137-6His protein
- Resuspend cell pellet in the lysis buffer (typically 5 ml/g of cells). If cell resuspension is difficult, up to 10 ml/g of cells of lysis buffer can be used.
- Add Antifoam 204 (1-2 drops for ~20 ml).
- Disrupt cells on a Cell Disruption system at 1.5-1.8 kbar and incubate lysate for 20 min on ice. If lysate is too viscous upon disruption, add more Benzonase nuclease and/or increase incubation time on ice.
- Spin lysate at 45,000 x g for 30 min at 4 °C. Recover supernatant, dilute 2x in binding/washing buffer and further clarify it by filtration using a 0.2 µm filter.
- Equilibrate the Histrap FF 1 ml (or 5 ml for scale-up) with 5-10 CV (column volumes) of binding/washing buffer at 1 ml/min (2 ml/min on the 5 ml column).
- Inject clarified lysate in the column at 1 ml/min. Collect the column flow-through.
- Wash column with the same buffer until Abs280nm is stable.
- Wash column at 1 ml/min (2 ml/min on the 5 ml column) with 5-10 CV of 8% elution buffer (40 mM imidazole) followed by 5-10 CV of 16% (80 mM imidazole) of the same solution.
- Elute proteins at 1 ml/min (2 ml/min for the 5 ml column) using a linear gradient from 16-100% of the same buffer (80-500 mM imidazole). Typically, gradient is performed for 10 min (15 min on the 5 ml column).
- Collect fractions (1 ml) throughout all purification steps.
- Evaluate protein purity of MGD1△137-6His containing fractions by SDS-PAGE analysis - take 16 µl of sample and add 4 µl of 5x DB. Boil samples for 5 min and sonicate for 2 min prior to gel loading.
- Evaluate protein homogeneity of MGD1△137-6His containing fractions by dynamic light scattering (DLS) analysis (see Note 7 for details).
- Pool appropriate fractions and concentrate up to 20 mg/ml using a Vivaspin 20, MWCO 30,000 Da concentrator (see Note 8 for details).
- Monitor protein concentration by UV absorbance at 280 nm on a NanoDropTM 2000 spectrophotometer, using a sequence-derived extinction coefficient of 47,900 M-1 cm-1. Use 1:1 washing/elution buffer mix [(imidazole)final ~250 mM] as blank.
- Resuspend cell pellet in the lysis buffer (typically 5 ml/g of cells). If cell resuspension is difficult, up to 10 ml/g of cells of lysis buffer can be used.
- Size exclusion purification of MGD1△137-6His protein
- Equilibrate the Superdex S200 10/300 GL column with 2 CV (48 ml) of size exclusion buffer at 0.5-0.8 ml/min (control that pressure does not exceed the maximum limit of 1.5 MPa, otherwise decrease the flow).
- Inject a maximum of 500 µl of a 20 mg/ml protein sample in the column.
- Wash the column with the same buffer to elute proteins.
- Collect 0.5 ml fractions and perform SDS-PAGE and DLS analysis of the samples. For the SDS-PAGE, take 16 µl of sample and add 4 µl of 5x DB. Boil samples for 5 min and sonicate for 2 min prior to gel loading. If samples are too viscous for gel loading, increase boiling and/or sonication times.
- Wash the column with 2 CV of Mili-Q (MQ) water, followed by 2 CV of 20% ethanol.
- Pool fractions containing pure and homogeneous MGD1△137-6His protein.
- Concentrate the protein to ~10 mg/ml using a Vivaspin 20, MWCO 30,000 concentrator.
- Monitor protein concentration by UV absorbance at 280 nm, using the size exclusion buffer as blank.
- Prepare 40 µl aliquots and store at -20 °C, until further use.
- Equilibrate the Superdex S200 10/300 GL column with 2 CV (48 ml) of size exclusion buffer at 0.5-0.8 ml/min (control that pressure does not exceed the maximum limit of 1.5 MPa, otherwise decrease the flow).
- Crystallization of MGD1△137-6His protein
- Prepare a protein stock solution at 6 mg/ml (use the size exclusion buffer for dilution).
- Prepare the precipitant solution containing 200 mM MgCl2·6H2O and 22% (w/v) PEG 3350.
- Pipet 400 µl of precipitant solution into the reservoir(s) of the crystallization plate.
- Prepare the crystallization drop(s) on the cover slip with 1 µl of MGD1△137-6His at 6 mg/ml and 2 µl of precipitant solution.
- Place the cover slip over the well (drop facing reservoir) and seal.
- Incubate plate at 20 °C. Inspect the plate frequently, under a microscope (crystals appear in one week, but often grow bigger within 10-12 days). Avoid temperature changes and keep the plate at 20 °C as much as possible.
- Crystals cryoprotection is achieved in a two-step procedure (i) by transferring into a stabilizing solution of mother liquor (precipitant solution) containing 27% (w/v) PEG 3350 precipitant, (ii) followed by a quick soak in the same cryoprotective solution supplemented with 30% (w/v) of the same precipitant. Use the cryocrystallography material to mount and handle crystals.
- Flash-cool and store crystals in LN2.
- Prepare a protein stock solution at 6 mg/ml (use the size exclusion buffer for dilution).
Notes
- A more detailed explanation on the role of MGD1 in galactolipid biosynthesis is described in Rocha et al. (2013; 2016).
- All solutions must be filtered prior to use. Solutions to be used in size exclusion chromatography have also to be degassed.
- To accelerate cell growth, a pre-inoculum can be used. Prepare a small culture using the procedure described, and incubate overnight at 37 °C, 180 rpm. Inoculate the new LB media with 2-3% of the overnight culture (typically 20-30 ml/L of media).
- Length of column washing/equilibration steps can be adjusted (shortened or extended), by monitoring the stabilization of the absorbance at 280 nm, as given by the AKTA FPLC system.
- Because MGD1△137-6His has a molecular weight of 45 kDa, 10% acrylamide SDS-PAGE gels can be used.
- DTT can be used during the procedure, but for long-term storage TCEP is more stable and efficient.
- DLS is an ideal technique for characterizing proteins in a variety of conditions and obtain information on their state in solution. It can be used to identify conditions that provide biologically relevant oligomeric structures, ensure monodispersity and optimal stability, minimize aggregation and ultimately, are ideal for crystallization. The default standard operation protocol (SOP) for protein size analysis of the Zetasizer Nano ZSTM was used, with modifications to temperature (4 °C or 20 °C) and type of solvent used (e.g., water + 5% glycerol) to take into account the viscosity of the solution. Triplicate measurements are performed automatically. If the sample is monodisperse, only one population is seen (single peak), whereas polydispersity is represented by multiple populations (several peaks); also, the size distribution (peak width) around the protein diameter should be as narrow as possible; and finally the particle diameter should be compatible with the protein species (very large diameters indicate protein aggregation). A typical DLS experiment for MGD1, together with its expected results, can be found in Rocha et al. (2013), where the protein production optimized protocol was first published.
- Throughout the purification, the appropriate fractions are determined according to their purity (SDS-PAGE) and homogeneity (DLS). MGD1△137-6His fractions that are not very pure or not homogeneous are discarded from the pooling and not used in the following steps. Typically, fractions from the beginning and end of the eluted chromatographic peak are not included in the pooling.
- Several tutorials are available showing how to crystallize a protein in vapour diffusion techniques. Some useful links to tutorials are given here:
- xray.bmc.uu.se/terese/tutorials.html – A tutorial by Terese Bergfors on protein crystallization
- https://www.youtube.com/watch?v=gLsC4wlrR2A – Understanding Crystallography, Part 1: From proteins to crystals
- https://www.youtube.com/watch?v=Grh8-pHpAJ4 – Protein crystallization hanging drop vapour diffusion
Recipes
- Lysis buffer
25 mM HEPES buffer
500 mM NaCl
1 M urea
10 mM imidazole
1 mM DTT
Adjust pH to 7.5
Add Benzonase® nuclease (1 µl/10 ml of solution) and cOmpleteTM protease inhibitor cocktail (1 tablet/50 ml solution) - Washing/binding buffer
25 mM HEPES
500 mM NaCl
1 M urea
10 mM imidazole
1 mM DTT
Adjust pH to 7.5 - Elution buffer
25 mM HEPES
500 mM NaCl
1 M urea
500 mM imidazole
1 mM DTT
Adjust pH to 7.5 - Size exclusion buffer
25 mM Bis-Tris propane
150 mM NaCl
5% (v/v) glycerol
1 mM TCEP
Adjust to pH 6.5 - SDS-PAGE deposition buffer (5x DB)
250 mM Tris, pH 6.8
50 % (v/v) glycerol
10 % (v/v) SDS
0.05% (w/v) bromophenol blue
250 mM DTT - 20% (v/v) ethanol
- 6 N hydrochloric acid solution (for pH adjustment)
- 10 M sodium hydroxide solution (for pH adjustment)
- Precipitant solution (or mother liquor)
200 mM MgCl2·6H2O
22% (w/v) PEG 3350
Acknowledgments
This protocol is an extension of that described in Rocha et al., 2013 and Rocha et al., 2016. This work was supported by Agence Nationale de la Recherche (grant ANR- 10-BLAN-1524, ReGal) and by the Rhône-Alpes region (France).
References
- Botté, C., Jeanneau, C., Snajdrova, L., Bastien, O., Imberty, A., Breton, C. and Marechal, E. (2005). Molecular modeling and site-directed mutagenesis of plant chloroplast monogalactosyldiacylglycerol synthase reveal critical residues for activity. J Biol Chem 280(41): 34691-34701.
- Miège, C., Maréchal, E., Shimojima, M., Awai, K., Block, M. A., Ohta, H., Takamiya, K., Douce, R. and Joyard, J. (1999). Biochemical and topological properties of type A MGDG synthase, a spinach chloroplast envelope enzyme catalyzing the synthesis of both prokaryotic and eukaryotic MGDG. Eur J Biochem 265(3): 990-1001.
- Nishiyama, Y., Hardré-Liénard, H., Miras, S., Miège, C., Block, M. A., Revah, F., Joyard, J. and Marechal, E. (2003). Refolding from denatured inclusion bodies, purification to homogeneity and simplified assay of MGDG synthases from land plants. Protein Expr Purif 31(1): 79-87.
- Rocha, J., Audry, M., Pesce, G., Chazalet, V., Block, M. A., Maréchal, E. and Breton, C. (2013). Revisiting the expression and purification of MGD1, the major galactolipid synthase in Arabidopsis to establish a novel standard for biochemical and structural studies. Biochimie 95(4): 700-708.
- Rocha, J., Sarkis, J., Thomas, A., Pitou, L., Radzimanowski, J., Audry, M., Chazalet, V., de Sanctis, D., Palcic, M. M., Block, M. A., Girard-Egrot, A., Marechal, E. and Breton, C. (2016). Structural insights and membrane binding properties of MGD1, the major galactolipid synthase in plants. Plant J 85(5): 622-633.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Rocha, J., Chazalet, V. and Breton, C. (2016). Expression, Purification and Crystallization of Recombinant Arabidopsis Monogalactosyldiacylglycerol Synthase (MGD1). Bio-protocol 6(24): e2064. DOI: 10.21769/BioProtoc.2064.
Category
Plant Science > Plant biochemistry > Protein > Isolation and purification
Plant Science > Plant biochemistry > Protein > Structure
Biochemistry > Protein > Expression
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












